Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.116121
Revised: December 10, 2025
Accepted: January 13, 2026
Published online: April 21, 2026
Processing time: 163 Days and 16.3 Hours
While the pioneering study by Liu et al published in World Journal of Gastroenterology masterfully addresses the critical challenge of predicting chemotherapy-induced myelosuppression, it also illuminates the necessary path for future research: The development of integrated models that concurrently predict a spec
Core Tip: We propose expanding Liu et al’s interpretable machine learning-based nomogram beyond myelosuppression to an integrated efficacy-toxicity platform that simultaneously forecasts tumor response and common multi-organ adverse events. By embedding these dual predictions into a real-time, utility-based clinical decision support system, designed for practical clinical use, clinicians can instantly identify the regimen offering each patient the maximal therapeutic index, thereby optimizing personalized treatment decisions.
- Citation: Qin YW, Li PW, Liang XY, Mo Y, Chen DW. Letter to the Editor: Balancing efficacy and toxicity: The critical role of predictive models in colorectal cancer chemotherapy. World J Gastroenterol 2026; 32(15): 116121
- URL: https://www.wjgnet.com/1007-9327/full/v32/i15/116121.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i15.116121
Liu et al’s study[1] published in World Journal of Gastroenterology for adeptly predicting the risk of chemotherapy-induced myelosuppression (CIM) in colorectal cancer patients is a scientific and pioneering work. By integrating multiple machine learning algorithms via a feature mapping approach, they have successfully bridged the critical gap between the high predictive accuracy of complex models and the clinical necessity for interpretability. With an impressive area under the receiver operating characteristic curve of 0.95 in the testing set, this model empowers clinicians to proactively manage one of the most common and debilitating dose-limiting toxicities in colorectal cancer treatment.
The successful prediction of CIM illuminates the next frontier in personalized chemotherapy: Developing integrated models that predict a spectrum of toxicities while crucially linking them to efficacy outcomes. CIM is but one facet of a patient’s tolerance to chemotherapy. Other frequent and equally impactful adverse events include severe gastrointestinal distress (diarrhea, nausea/vomiting), debilitating neurotoxicity, hand-foot syndrome, and hematological toxicities[2,3]. The clinical decision-making process weighs all these potential toxicities simultaneously (Table 1).
| Toxicity type | Occurrence of risk | Clinical impact | Associated regimens | Impact on decision-making |
| Chemotherapy-induced myelosuppression | High (FOLFOX); low (CAPOX) | Neutropenia, anemia, thrombocytopenia; risk of fever, infection, fatigue, bleeding | FOLFOX, FOLFIRI, CAPOX | Common reasons for dose delay, reduction, or use of growth factor support |
| Severe diarrhea | High (FOLFOX); medium (CAPOX) | Watery stools, dehydration, and electrolyte imbalance | FOLFOX, FOLFIRI, CAPOX | Frequent cause of dose modification or treatment interruption; requires proactive management |
| Peripheral neuropathy | Low (FOLFOX); high (CAPOX) | Numbness, tingling in the limbs, can progress to functional impairment (e.g., difficulty with fine motor tasks) | FOLFOX, CAPOX | Cumulative toxicity often leads to dose reduction or discontinuation of oxaliplatin |
| Hand-foot syndrome | Medium (FOLFOX); high (CAPOX) | Erythema, swelling, pain, blistering on palms and soles | FOLFOX, CAPOX | May necessitate dose reduction or interruption of capecitabine/fluorouracil |
This field is already witnessing the emergence of predictive models for individual toxicities. For instance, pharmacogenetic testing for uridine diphosphate glucuronosyltransferase family 1 member A1 is routinely used to identify patients at high risk for irinotecan-induced severe neutropenia and diarrhea[4,5], while another research area is identifying genetic markers for oxaliplatin-induced neurotoxicity[6,7]. Similarly, models exist to predict the efficacy of chemotherapy. Studies have explored radiographic features, transcriptomic signatures, and clinical factors to forecast responses to neoadjuvant therapy or outcomes in metastatic settings[8-12]. Yet, these models often operate in isolation, creating a fragmented decision-making landscape. A clinician may be confronted with one model suggesting high efficacy for fluorouracil, leucovorin, and irinotecan (FOLFIRI) and the other predicting severe diarrhea from the same regimen. The CIM prediction model of Liu et al[1] indicates a moderate risk of myelosuppression. Synthesizing this multidimensional prediction into a single, coherent treatment plan that optimally balances efficacy against toxicity remains a profound clinical challenge. The value of a unified model lies precisely in its ability to illuminate this critical trade-off, rather than presenting it as a conflict.
Therefore, we propose that the ultimate goal is not merely to avoid a single toxicity, but to identify the treatment regimen with the most favorable therapeutic index for each individual patient. A unified model that could predict, for instance, both a patient’s high risk for severe diarrhea from irinotecan and their high probability of achieving a profound pathological response to an FOLFIRI regimen would provide an invaluable, holistic framework for clinical decision-making. This framework could be practically integrated into the clinical workflow, such as during pre-chemotherapy multidisciplinary team meetings or patient consultations, to visually compare the predicted efficacy-toxicity profiles of different regimen options. Such a model empowers clinicians by explicitly quantifying the competing dimensions of benefit and risk. In this scenario, a clinician might still justifiably choose FOLFIRI but intensify prophylactic anti-diarrheal measures and schedule more frequent follow-ups. Conversely, if the predicted efficacy is low and the toxicity risk is high, an alternative regimen would be strongly favored.
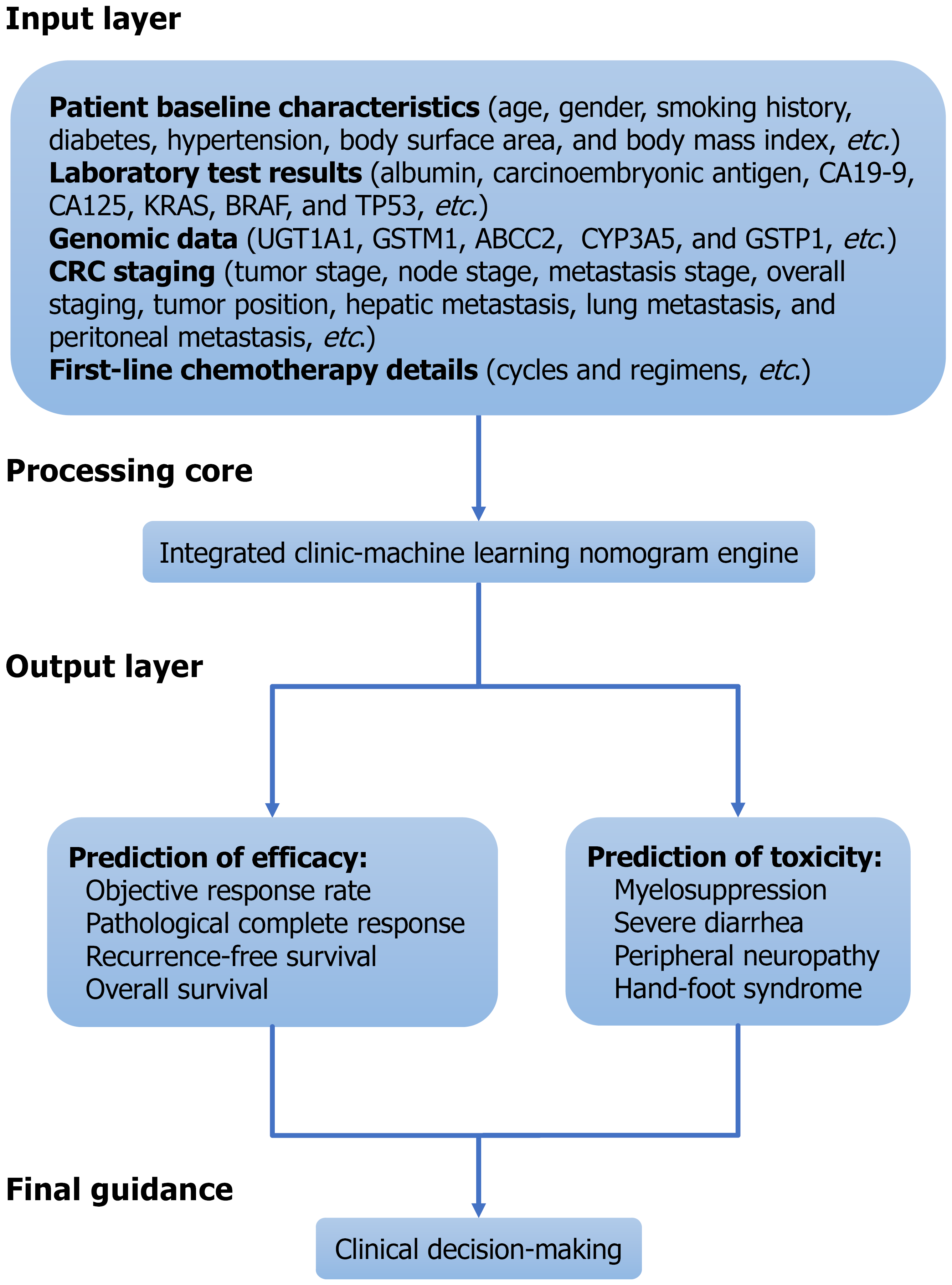
We envision the development of an integrated clinical decision-support platform that synthesizes these elements. Future research should build upon the robust methodology established by Liu et al[1]. The next step is to integrate a wider array of inputs to create a unified, multi-output predictive framework. This proposed platform would not be a simple aggregation of independent sub-models, but an integrated system where the interactions between features for different outcomes can be captured (Figure 1). Multi-task learning (MTL) architectures offer a powerful paradigm for developing such an integrated prediction platform. MTL models, particularly those based on neural networks, are designed to learn multiple related tasks (e.g., predicting various toxicities and efficacy) concurrently by sharing representations across tasks[13]. This approach can improve generalization for individual tasks by leveraging common underlying factors and inherently models the correlations between different clinical outcomes, moving beyond isolated predictions. MTL frameworks have been widely applied in oncology, providing a methodological precedent for our proposal[14-16].
However, the development of this platform faces substantial challenges. Data requirements are paramount, necessitating large, curated, multi-modal datasets that harmonize clinical variables, genomic data, biomedical images, and longitudinal toxicity reports. The most pivotal challenge, however, lies in quantifying the “therapeutic index” for clinical decision support. This requires defining a clinically valid utility or net benefit function, where weights reflecting the relative clinical importance of different efficacy levels and toxicity severities are formally incorporated. Determining these weights is a non-trivial research endeavor in itself, likely requiring methods like discrete choice experiments involving oncologists, patients, and health economists to establish consensus[17,18]. Therefore, advancing this field demands not only robust predictive algorithms but also a parallel interdisciplinary effort to bridge predictive analytics with actionable clinical choice.
We again congratulate Liu et al[1] on their valuable contribution. Their work provides a solid methodological foundation and a powerful tool for managing CIM. By developing integrated models that concurrently predict toxicity and efficacy, we can move from isolated risk assessment to truly optimizing the balance between treatment safety and anti-tumor activity.
| 1. | Liu YM, Du YY, Song Y, Xiong HT, Yu HB, Li BH, Cai L, Ma SS, Gao J, Zhang HY, Fang RY, Cai R, Zheng HG. Predicting chemotherapy-induced myelosuppression in colorectal cancer: An interpretable, machine learning-based nomogram. World J Gastroenterol. 2025;31:112180. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 2. | Aoullay Z, Slaoui M, Razine R, Er-Raki A, Meddah B, Cherrah Y. Therapeutic Characteristics, Chemotherapy-Related Toxicities and Survivorship in Colorectal Cancer Patients. Ethiop J Health Sci. 2020;30:65-74. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 13] [Article Influence: 2.2] [Reference Citation Analysis (1)] |
| 3. | Tong L, Ahn C, Symanski E, Lai D, Du XL. Effects of newly developed chemotherapy regimens, comorbidities, chemotherapy-related toxicities on the changing patterns of the leading causes of death in elderly patients with colorectal cancer. Ann Oncol. 2014;25:1234-1242. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 30] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 4. | Innocenti F, Undevia SD, Iyer L, Chen PX, Das S, Kocherginsky M, Karrison T, Janisch L, Ramírez J, Rudin CM, Vokes EE, Ratain MJ. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J Clin Oncol. 2004;22:1382-1388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 821] [Cited by in RCA: 697] [Article Influence: 31.7] [Reference Citation Analysis (5)] |
| 5. | Ichikawa W, Uehara K, Minamimura K, Tanaka C, Takii Y, Miyauchi H, Sadahiro S, Fujita K, Moriwaki T, Nakamura M, Takahashi T, Tsuji A, Shinozaki K, Morita S, Ando Y, Okutani Y, Sugihara M, Sugiyama T, Ohashi Y, Sakata Y. An internally and externally validated nomogram for predicting the risk of irinotecan-induced severe neutropenia in advanced colorectal cancer patients. Br J Cancer. 2015;112:1709-1716. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 29] [Article Influence: 2.6] [Reference Citation Analysis (1)] |
| 6. | Terrazzino S, Argyriou AA, Cargnin S, Antonacopoulou AG, Briani C, Bruna J, Velasco R, Alberti P, Campagnolo M, Lonardi S, Cortinovis D, Cazzaniga M, Santos C, Kalofonos HP, Canonico PL, Genazzani AA, Cavaletti G. Genetic determinants of chronic oxaliplatin-induced peripheral neurotoxicity: a genome-wide study replication and meta-analysis. J Peripher Nerv Syst. 2015;20:15-23. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 34] [Article Influence: 3.1] [Reference Citation Analysis (1)] |
| 7. | Lecomte T, Landi B, Beaune P, Laurent-Puig P, Loriot MA. Glutathione S-transferase P1 polymorphism (Ile105Val) predicts cumulative neuropathy in patients receiving oxaliplatin-based chemotherapy. Clin Cancer Res. 2006;12:3050-3056. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 167] [Cited by in RCA: 160] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 8. | Tsuji A, Sunakawa Y, Ichikawa W, Nakamura M, Kochi M, Denda T, Yamaguchi T, Shimada K, Takagane A, Tani S, Kotaka M, Kuramochi H, Furushima K, Koike J, Yonemura Y, Takeuchi M, Fujii M, Nakajima T. Early Tumor Shrinkage and Depth of Response as Predictors of Favorable Treatment Outcomes in Patients with Metastatic Colorectal Cancer Treated with FOLFOX Plus Cetuximab (JACCRO CC-05). Target Oncol. 2016;11:799-806. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 3.2] [Reference Citation Analysis (2)] |
| 9. | Cremolini C, Loupakis F, Antoniotti C, Lonardi S, Masi G, Salvatore L, Cortesi E, Tomasello G, Spadi R, Zaniboni A, Tonini G, Barone C, Vitello S, Longarini R, Bonetti A, D'Amico M, Di Donato S, Granetto C, Boni L, Falcone A. Early tumor shrinkage and depth of response predict long-term outcome in metastatic colorectal cancer patients treated with first-line chemotherapy plus bevacizumab: results from phase III TRIBE trial by the Gruppo Oncologico del Nord Ovest. Ann Oncol. 2015;26:1188-1194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 165] [Cited by in RCA: 161] [Article Influence: 14.6] [Reference Citation Analysis (1)] |
| 10. | Yen LC, Yeh YS, Chen CW, Wang HM, Tsai HL, Lu CY, Chang YT, Chu KS, Lin SR, Wang JY. Detection of KRAS oncogene in peripheral blood as a predictor of the response to cetuximab plus chemotherapy in patients with metastatic colorectal cancer. Clin Cancer Res. 2009;15:4508-4513. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 72] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 11. | Sakamaki K, Kito Y, Yamazaki K, Izawa N, Tsuda T, Morita S, Boku N. Exploration of time points and cut-off values for early tumour shrinkage to predict survival outcomes of patients with metastatic colorectal cancer treated with first-line chemotherapy using a biexponential model for change in tumour size. ESMO Open. 2017;2:e000275. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 8] [Article Influence: 0.9] [Reference Citation Analysis (1)] |
| 12. | Sveen A, Bruun J, Eide PW, Eilertsen IA, Ramirez L, Murumägi A, Arjama M, Danielsen SA, Kryeziu K, Elez E, Tabernero J, Guinney J, Palmer HG, Nesbakken A, Kallioniemi O, Dienstmann R, Lothe RA. Colorectal Cancer Consensus Molecular Subtypes Translated to Preclinical Models Uncover Potentially Targetable Cancer Cell Dependencies. Clin Cancer Res. 2018;24:794-806. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 192] [Article Influence: 24.0] [Reference Citation Analysis (4)] |
| 13. | Lin B, Jiang W, Ye F, Zhang Y, Chen P, Chen YC, Liu S, Tsang IW, Kwok JT. Dual-balancing for multi-task learning. Neural Netw. 2025;195:108317. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 14. | van der Voort SR, Incekara F, Wijnenga MMJ, Kapsas G, Gahrmann R, Schouten JW, Nandoe Tewarie R, Lycklama GJ, De Witt Hamer PC, Eijgelaar RS, French PJ, Dubbink HJ, Vincent AJPE, Niessen WJ, van den Bent MJ, Smits M, Klein S. Combined molecular subtyping, grading, and segmentation of glioma using multi-task deep learning. Neuro Oncol. 2023;25:279-289. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 108] [Article Influence: 27.0] [Reference Citation Analysis (1)] |
| 15. | Chen K, Wang M, Song Z. Multi-task learning-based histologic subtype classification of non-small cell lung cancer. Radiol Med. 2023;128:537-543. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (1)] |
| 16. | Zou Y, Cai L, Chen C, Shao Q, Fu X, Yu J, Wang L, Chen Z, Yang X, Yuan B, Liu P, Lu Q. Multi-task deep learning based on T2-Weighted Images for predicting Muscular-Invasive Bladder Cancer. Comput Biol Med. 2022;151:106219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 19] [Article Influence: 4.8] [Reference Citation Analysis (1)] |
| 17. | Bahrampour M, Byrnes J, Norman R, Scuffham PA, Downes M. Discrete choice experiments to generate utility values for multi-attribute utility instruments: a systematic review of methods. Eur J Health Econ. 2020;21:983-992. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 18. | Saad ED, Zalcberg JR, Péron J, Coart E, Burzykowski T, Buyse M. Understanding and Communicating Measures of Treatment Effect on Survival: Can We Do Better? J Natl Cancer Inst. 2018;110:232-240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 57] [Article Influence: 7.1] [Reference Citation Analysis (1)] |