Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.116105
Revised: December 14, 2025
Accepted: January 27, 2026
Published online: April 21, 2026
Processing time: 164 Days and 3.3 Hours
Small intestinal bleeding (SIB) remains a significant challenge in the diagnosis of obscure gastrointestinal bleeding. While capsule endoscopy (CE) is the gold standard for visualization, manual interpretation of the extensive video footage is labor-intensive and subject to inter-observer variability. Although convolutional neural networks (CNNs) have improved lesion detection, standard models often fail to account for temporal continuity, limiting their ability to accurately predict the specific location of bleeding points within the small bowel.
To develop and validate a deep learning framework integrating CNNs with long short-term memory (LSTM) networks to enhance the automated detection and precise localization of SIB.
This study employed two datasets for automated bleeding detection: One from National Cheng Kung University, consisting of white light imaging images from 100 patients obtained via PillCam™ SB 3 CE, and the Kvasir-Capsule Image dataset, which includes 47238 labeled images across 14 pathological categories. Nineteen continuous picture sequences were recovered, comprising 3806 bleeding photos and 3275 non-bleeding images.
Data augmentation was implemented, utilizing CNNs for feature extraction, succeeded by long short-term memory networks for prediction. The CNN model attained an accuracy of 98.6% for 10 categories and 96.7% for 2 categories. Findings demonstrate that CNN-LSTM models exhibit superior performance with expanded category sets.
These findings underscore the capability of deep learning models to enhance the accuracy and efficiency of CE-based gastrointestinal bleeding diagnosis, hence facilitating improved clinical decision-making.
Core Tip: This study proposes a practical deep learning pipeline that combines convolutional neural networks for spatial feature extraction with long short-term memory networks for temporal modeling to detect small intestinal bleeding in capsule endoscopy videos. Using datasets from a clinical cohort and the Kvasir-Capsule collection, we show that models trained with richer multiclass labels yield more informative features for sequential prediction, enabling higher long short-term memory accuracy and lower temporal error than binary setups. The approach delivers consistent frame-by-frame detection performance with clinically feasible processing speed, highlights the value of leveraging temporal dependencies beyond single-frame analysis, and outlines a path toward interpretable, efficient triage in capsule endoscopy workflows.
- Citation: Kuo HY, Lee KH, Chou CK, Mukundan A, Karmakar R, Chen TH, Wang TL, Liu PH, Wang HC. Deep learning-enhanced prediction of small intestinal bleeding points using long short-term memory networks. World J Gastroenterol 2026; 32(15): 116105
- URL: https://www.wjgnet.com/1007-9327/full/v32/i15/116105.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i15.116105
Gastrointestinal (GI) disease is one of the most prevalent digestive disorders and has a substantial impact on global health[1]. Around 2.56 million deaths were attributed to digestive illnesses in 2019[2], with GI conditions contributing significantly to this statistic. GI tract is a large area making the use of traditional endoscope unusable to reach the depth of the GI tract[3]. Therefore, the use of capsule endoscope is necessary for early detection of GI related diseases such as small intestinal bleeding that refers to bleeding from the inside of the small intestine[4]. The most important feature of small intestinal bleeding is blood, and blood detection is particularly important using the suspected blood indicator is a traditional tool used to automatically mark images that may be bleeding in the reading system[5-7].
In recent years, the advancement of graphics processing units (GPUs) has had a huge impact on deep learning[8]. Traditional central processing units (CPUs) have encountered bottlenecks in processing these calculations[9]. GPUs architectural design is capable of performing multiple computing tasks simultaneously. This capability is called parallel processing, which has increased the availability of capsule endoscopy datasets with a large number of labels, opening the door to the possibility of deep CNNs processing[10]. The advancement of GPUs has also had a significant impact on deep learning of recurrent neural networks (RNNs)[11-13]. RNNs are neural networks that can capture and remember long-term dependencies[14]. They are used to process fields such as natural language, speech recognition, and time series analysis, and therefore require the processing of large amounts of time series data[13]. On traditional CPUs, these calculations are very time-consuming, causing the training process to become slow. The parallel computing capabilities of GPUs enable RNNs to process calculations simultaneously, speeding up the training process and hence the use of machine learning and deep learning models[15].
With the advancement of artificial intelligence (AI), deep learning classification is crucial for endoscopic imaging[16]. It can help explain the decisions made by deep learning models so that the results and treatments can be explained to patients, improve the diagnostic accuracy of endoscopic examinations, and facilitate early detection of diseases. Doctors can immediately identify and diagnose tumors in the body during endoscopic examinations, and the decisions they make may have a significant impact on the health of patients. Recent advancements in AI have profoundly altered medical image analysis, especially with the extensive utilization of convolutional neural networks (CNNs), which have exhibited robust representation learning capabilities for disease detection and lesion characterization. In GI imaging, CNN-based models have been progressively utilized to automate capsule endoscopy interpretation, demonstrating elevated diagnostic accuracy and efficiency. A recent thorough clinical review has underscored the growing significance of AI in small bowel endoscopic evaluation, accentuating its capacity to augment diagnostic reliability, alleviate physician workload, and enhance clinical decision-making[17]. Additionally, hybrid deep learning architectures that combine CNNs for extracting spatial features, long short-term memory (LSTMs) for modeling temporal dependencies, and attention mechanisms for weighting discriminative features have become strong tools for accurately analyzing medical images. A recent CNN-LSTM-attention model has shown better results on a number of medical imaging tasks, which shows that spatiotemporal attention-based fusion strategies are effective for automated disease diagnosis[18]. These advancements offer robust methodological endorsement for the CNN-LSTM-based framework utilized in the current study.
Paderno et al[19], devised a deep learning application that had been utilized for image processing, covering ad
Therefore, this paper presents a successful development of a deep learning methodology for the detection of GI bleeding via wireless capsule endoscopy images. Utilizing CNNs for feature extraction and LSTMs for sequential image analysis, we enhanced the precision of detecting and classifying bleeding events. Our experimental findings indicated that models trained on a greater number of categories attained superior accuracy relative to those trained on fewer categories. The amalgamation of CNN and LSTM facilitated the proficient identification of hemorrhaging over successive frames, underscoring the promise for automated diagnostic assistance in GI endoscopy. Future endeavors will concentrate on augmenting the dataset, refining model generalization, and incorporating real-time clinical applications to further elevate diagnostic precision and efficacy.
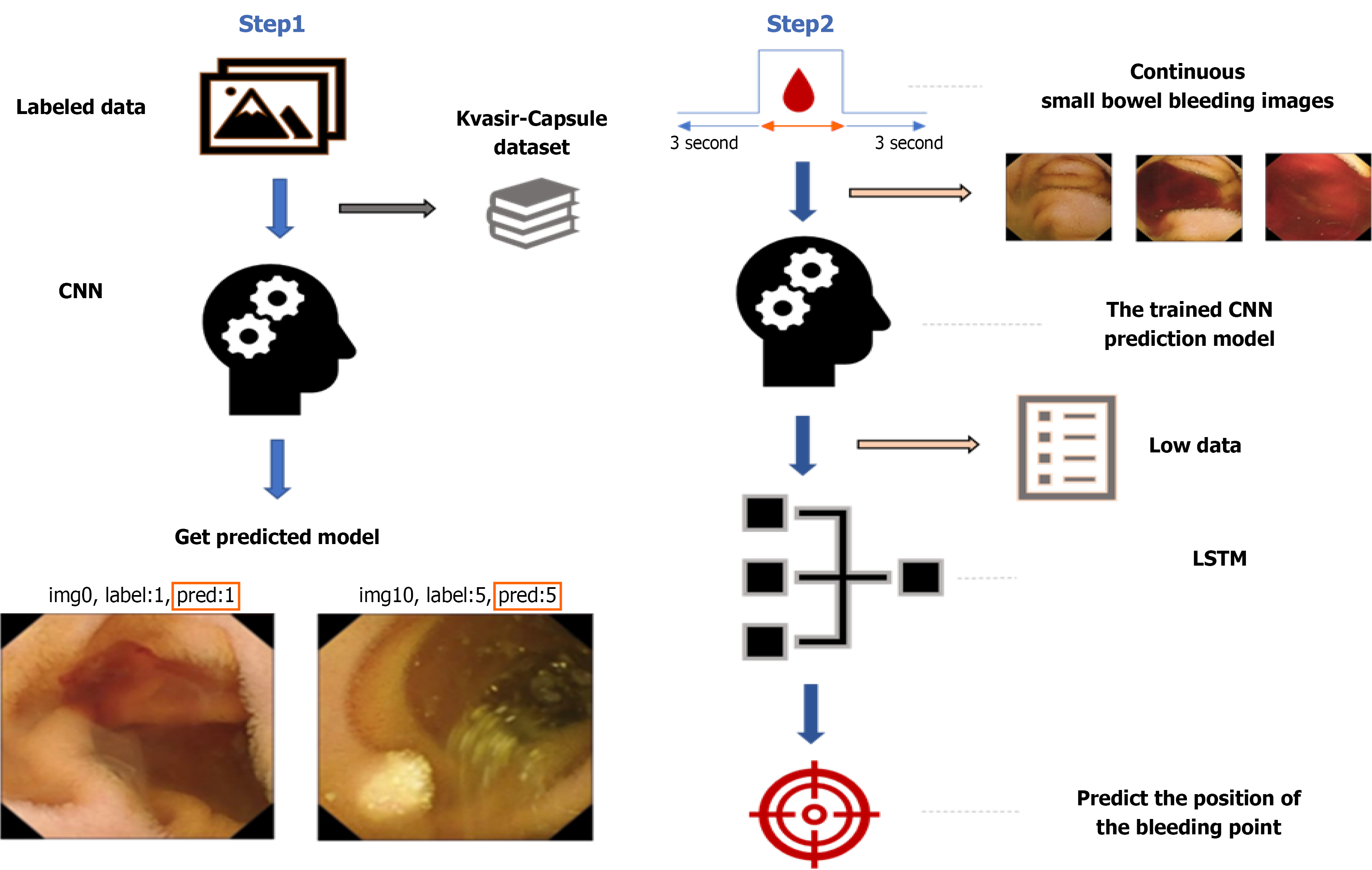
The experimental procedure comprises four stages as depicted in Figure 1. The initial phase entails the selection of small intestine capsule endoscopy images, specifically focusing on the extraction of bleeding images from the capsule endoscopy datasets supplied by National Cheng Kung University (NCKU) and the Kvasir-Capsule dataset. The second stage delineates the training of a CNN utilizing the annotated dataset. The third stage elucidates the processing of sequential bleeding images via the trained CNN model to extract characteristics and convert them into sequential data. The fourth stage addresses the application of the flattened sequential data into an LSTM network to forecast bleeding sites.
Medical imaging data from NCKU Hospital: This work involved collaboration with the Division of Gastroenterology and Hepatology at NCKU Hospital, employing white light imaging (WLI) data obtained from the PillCam™ SB3 capsule endoscopy device for the dataset[27]. To augment the dataset, we conducted supplementary image annotations to pinpoint bleeding spots seen in the WLI pictures. The Institutional Review Board of NCKU College of Medicine provided ethical approval for this investigation. The overall dataset used in this study is shown in Table 1.
| Kvasir labeled video | |||
| Category | 10 | ||
| Bleeding images | 1312 frames | ||
| Else | 6081 frames | ||
| Total | 7393 frames | ||
| Train | Stage: Bleeding | Images 1049 | Total = 5913 |
| Stage: Un-bleeding | Images 4864 | ||
| Val | Stage: Bleeding | Images 263 | Total = 1480 |
| Stage: Un-bleeding | Images 1217 | ||
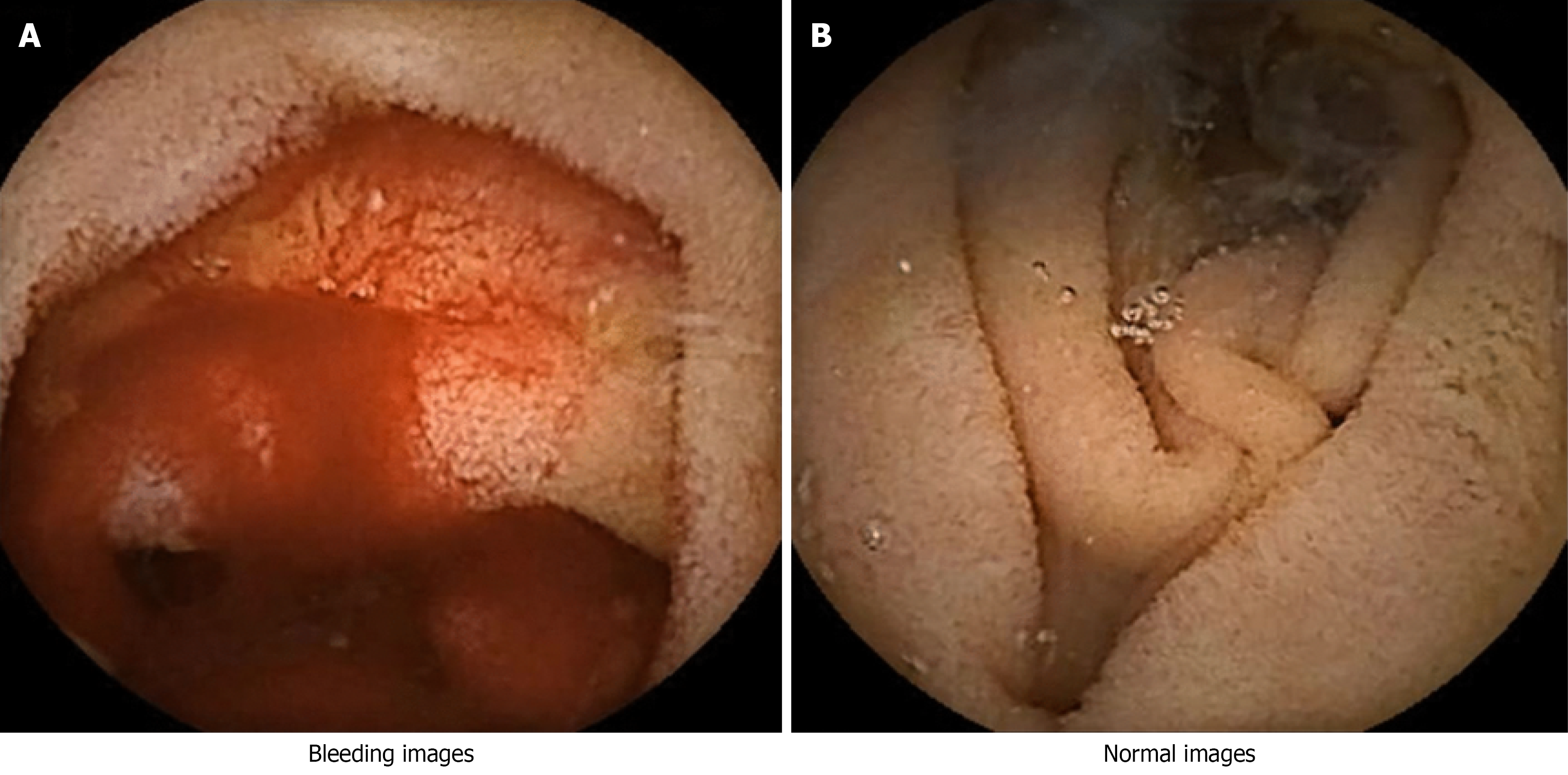
Kvasir-Capsule capsule endoscopy dataset: The Kvasir-Capsule dataset utilized in this research is an open-source capsule endoscopy dataset[28]. The dataset comprises 43 annotated image sequences, totaling 47238 labeled images classified into 14 distinct categories, and 74 unannotated sequences encompassing 4694266 images. The capsule endoscopy images in this dataset were obtained from clinical examinations performed at Vestre Viken Hospital Trust in Baerum, Norway, from February 2016 to January 2018. The assessments were conducted with the Olympus Endocapsule 10 System 45, comprising the Olympus EC-S10 capsule endoscope and the Olympus RE-10 recording apparatus. The photos were recorded at a frame rate of 2 frames per second with a resolution of 336 × 336 pixels. The videos were encoded in the H.264 (MPEG-4 AVC) format and exported as AVI files with the Olympus system's export tool. The initial frame rate of 2 fps was elevated to 30 fps via the export tool, while preserving the original encoding format. The dataset is entirely anonymized and has received approval from the Norwegian Data Protection Authority. All dataset splits were carried out on a patient-wise level to eliminate the possibility of data leakage, ensuring that frames from the same patient did not appear in both training and testing sets. The classification task with 10 classes involved restructuring the data into therapeutically pertinent clusters derived from the initial Kvasir-Capsule classifications. Two groups associated with bleeding were identified: New bleeding and angioectasia (vascular dilatation). Additionally, eight non-bleeding groups were recognized: Normal mucosa, lymphangiectasia, lymphoid hyperplasia, xanthoma, erythematous mucosa, erosion, ulceration, and polypoid lesion. This classification exemplifies standard disease presentations in capsule endoscopy and provides a proportionate framework for distinguishing active bleeding, vascular anomalies, and other lesions in the small intestine. The 10 categories enable the model to delineate not only bleeding-specific visual attributes but also a diverse array of non-bleeding anomalies, hence enhancing clinical interpretability and ensuring the reliability of the classification framework. Finally, Figure 2A is the bleeding image and Figure 2B is the normal intestinal image. A further drawback of this study is the considerable class imbalance between bleeding and non-bleeding samples in the training set (about 1:4.6). The proposed CNN-LSTM framework demonstrated consistent performance in this context; nevertheless, it did not employ specific strategies for addressing imbalances, such as class-weighted loss, focus loss, or oversampling. Future initiatives will focus on integrating class-balanced training approaches to improve the sensitivity of minority-class detection and clinical reliability.
Preprocessing of small intestine bleeding images: Image cropping and artifact removal: Cropping and scaling were utilized to eliminate patient-identifying information and extraneous noise. Furthermore, to maintain uniformity in image dimensions to satisfy model specifications, uniform scaling was executed. Certain models may possess particular specifications for image dimensions.
Extraction and labeling of bleeding images: Images of the small intestine are generally captured as video segments lasting around 8-10 hours. All frames exhibiting blood were retrieved and categorized according to their corresponding frame numbers in the video for analytical purposes. These labels function as indicators for recognizing bleeding areas in later sequential picture analysis.
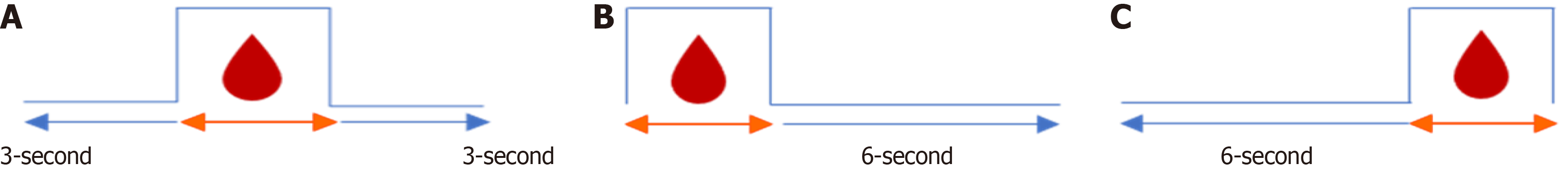
Preprocessing of sequential images for small intestine bleeding: LSTM networks necessitate sequential data for training to enable predictive prediction. To facilitate this, sequential photos demonstrating bleeding were chosen from both the NCKU Hospital dataset and the Kvasir-Capsule dataset. Subsequent segmentation of video sequences was conducted in accordance with established extraction criteria, as depicted in Figure 3. Subsequent to the extraction process, the final dataset comprises 19 sequences, encompassing 7081 continuous frames and 3806 tagged frames. The input data were organized into three temporal configurations to test the LSTM network (Figure 3).
Figure 3A: A continuous sequence capturing the transition period, including 3 seconds before and 3 seconds after the bleeding onset.
Figure 3B: A sequence with discontinuous pre-bleeding frames, followed by a continuous 6-second post-bleeding segment.
Figure 3C: A sequence maintaining 6 seconds of continuous pre-bleeding context, followed by discontinuous post-bleeding frames.
CNNs: A lot of studies that look at pictures of the digestive system have utilized big pretrained architectures like ResNet-50 and InceptionV3, but this one employed a private lightweight CNN backbone. This design choice was made for three key reasons. First, the proposed method combines CNN and LSTM to focus on modeling spatiotemporal features. In this scenario, learning over time consistently and effectively is more important than having too much spatial depth. A tiny CNN makes it easy to speed up time across a number of frames. Second, because the dataset is so big and this work uses short sequence windows, a lightweight architecture helps lower the chance of overfitting and makes it easier to generalize. Third, capsule endoscopic aid needs to be quick and effective in order to be beneficial. The proposed CNN is more suitable for clinical applications due to its reduced inference latency and lower memory consumption compared to heavyweight pretrained models. The principal aim of training CNNs is feature extraction. This approach converts sequential bleeding image frames into organized sequential data, which can subsequently be employed for LSTM networks to improve feature representation. The CNN model in this study is trained on VCE images measuring 336 × 336 pixels in RGB format. The model architecture comprises several convolutional layers, pooling layers, and fully linked layers. The principal elements of the architecture are as follows: (1) Initial convolutional layer: The input data is processed through a convolutional layer with 8 filters of size 3 × 3, utilizing the ReLU activation function; (2) Max pooling: A max pooling layer of size 2 × 2 is applied to down sample the feature maps; (3) Second convolutional block: The output is then fed into a second convolutional block, similar to the first but with an increased filter count of 16 to enhance feature extraction; and (4) Third convolutional block: A third convolutional block, also similar to the previous ones but with 32 filters, is employed to extract deeper-level features.
Following the convolutional layers, the feature maps are flattened into a one-dimensional vector and passed through a series of fully connected layers for classification: (1) A fully connected layer with 4096 neurons, utilizing the ReLU activation function and a 40% dropout rate to prevent overfitting; (2) Subsequent fully connected layers contain 4096, 1024, 512, and 256 neurons, respectively; and (3) The final output layer consists of 10 neurons with a softmax activation function to generate probability distributions for the ten categories.
In sequential modeling, the CNN backbone processed each frame of the capsule endoscopy, and the final global average pooling layer generated a fixed-length feature vector. In a sequence of T consecutive frames, the CNN produced T feature vectors, each of length D. They were organized sequentially to form a tensor of rank (batch size × sequence length T × feature dimension D). This tensor was subsequently input into the LSTM module. The feature sequence was input into the LSTM, and its hidden state was aggregated over time steps to capture temporal dependencies between frames. The final hidden state, or in certain trials the concatenated hidden states across all time steps, was provided to a fully connected classification layer to generate bleeding/non-bleeding predictions. This architecture enabled the CNN to acquire spatial information, which was then systematically organized and stored prior to modeling by the LSTM, thereby maintaining temporal continuity across frames. To temporally model the capsule endoscopy frames, the CNN architecture was developed to systematically extract spatial information from the capsule endoscopy images prior to transmitting them to the LSTM module. The network commences with three progressively more deeply-filtered convolutional blocks, each followed by max-pooling procedures to diminish the spatial dimensionality of the prominent feature activations. Subsequent to the convolutional processing step, the resultant feature maps are flattened and sent through a sequence of fully connected layers, which incorporate dropout regularization, prior to being directed to the final classification layer. Table 2 distinctly illustrates the precise configuration of the layers. The CNN was trained using the Adam optimizer, with an initial learning rate of 0.001, a batch size of 32, and a maximum of 180 epochs. A learning-rate scheduler was employed to decrease the learning rate by a factor of 0.1 across 10 successive epochs, should the validation loss reach a plateau. To mitigate overfitting, the dropout rate for the largest fully linked layer was established at 0.4. Categorical cross-entropy was utilized as the loss function for the ten-class scenario, whereas binary cross-entropy (BCE) was applied for the two-class scenario. Early stopping was implemented to terminate training if the validation loss fails to improve after fifteen consecutive epochs.
| Layer type | Details |
| Input | Capsule endoscopy frame, resized to 224 × 224 pixels, 3 channels (RGB) |
| Conv block 1 | Conv2D, 8 filters, 3 × 3 kernel, ReLU activation → MaxPooling 2 × 2 |
| Conv block 2 | Conv2D, 16 filters, 3 × 3 kernel, ReLU activation → MaxPooling 2 × 2 |
| Conv block 3 | Conv2D, 32 filters, 3 × 3 kernel, ReLU activation → MaxPooling 2 × 2 |
| Flatten | Flatten feature maps into 1D vector |
| Fully connected (FC1) | 4096 neurons, ReLU activation, dropout = 0.4 |
| Fully connected (FC2) | 4096 neurons, ReLU activation |
| Fully connected (FC3) | 1024 neurons, ReLU activation |
| Fully connected (FC4) | 512 neurons, ReLU activation |
| Fully connected (FC5) | 256 neurons, ReLU activation |
| Output layer | 10 neurons, Softmax activation (10-class); 2 neurons, Softmax (2-class) |
| Training optimizer | Adam, learning rate = 0.001, scheduler (reduce on plateau, factor 0.1) |
| Batch size | 32 |
| Epochs | Up to 180 (early stopping patience = 15) |
| Loss function | Categorical cross-entropy (10-class); binary cross-entropy (2-class) |
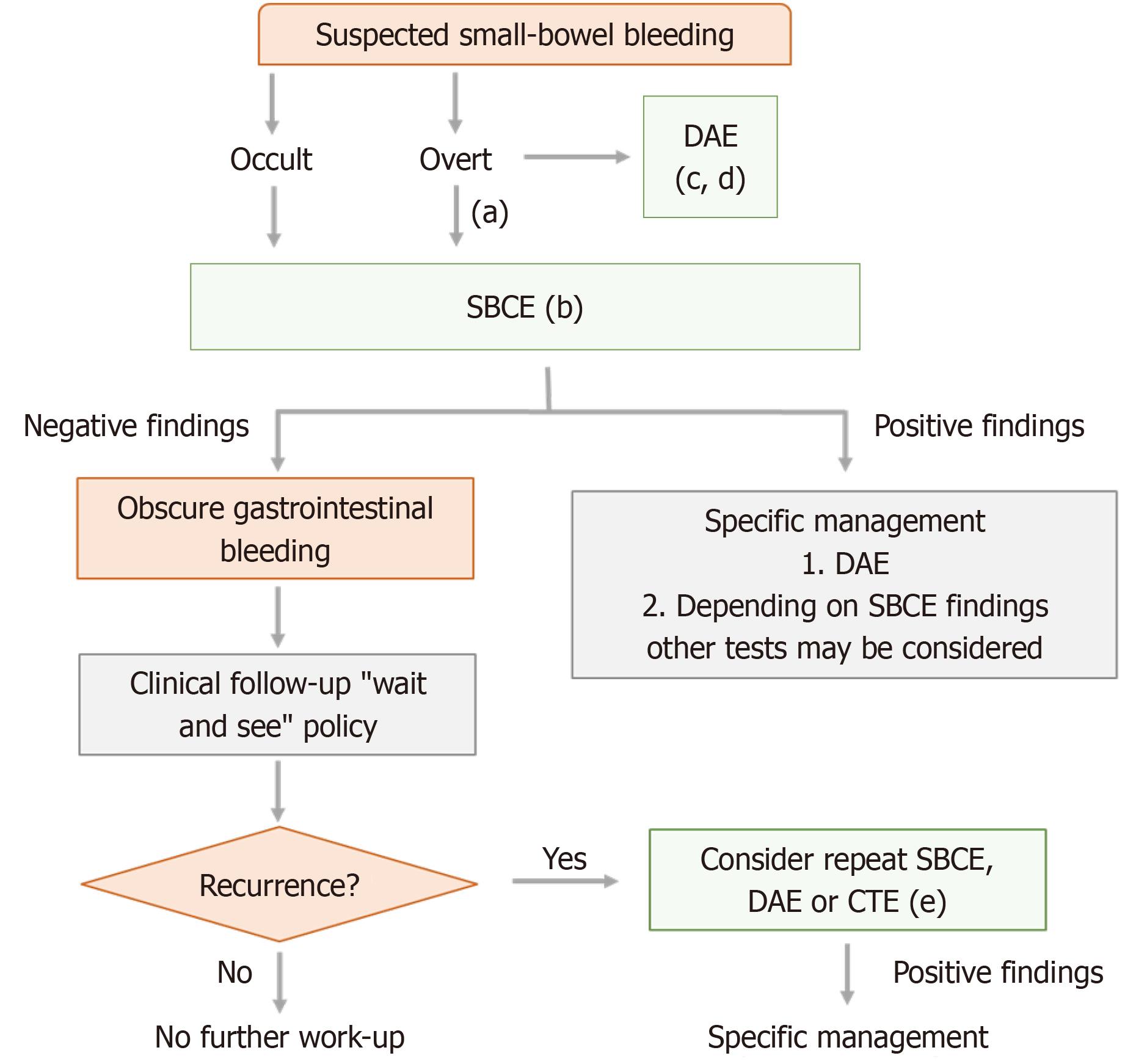
The retrieved consecutive images are entered into a trained CNN model for feature extraction and subsequently flattened into sequential data. The bleeding and non-bleeding areas in the successive photos are consistently tagged and converted into sequential data. This sequential dataset functions as the fundamental basis for future predictions with an LSTM network. The overall schematics of the diagnosis and treatment of unexplained GI bleeding is shown in Figure 4.
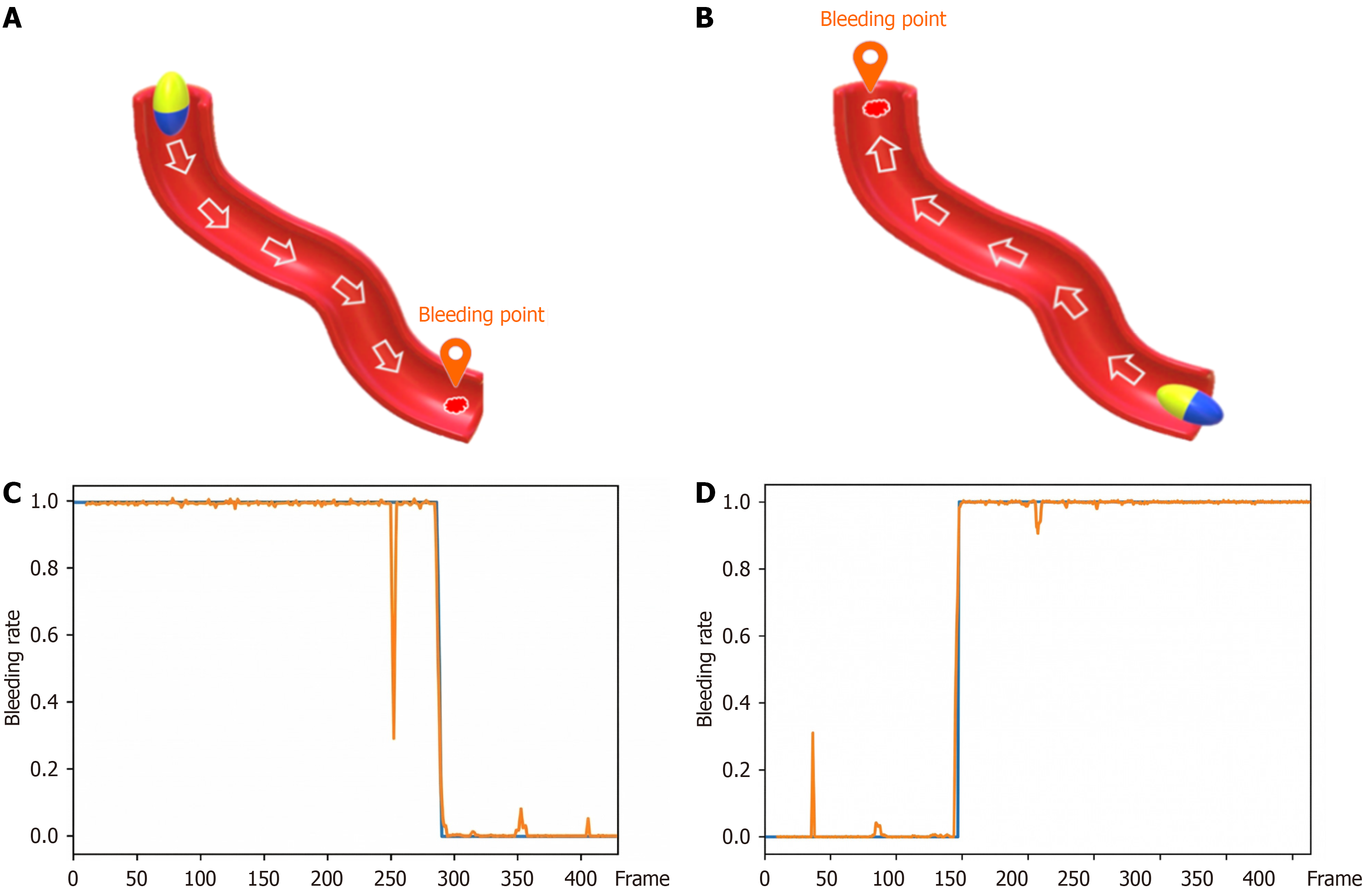
Before LSTM training, it is essential to establish the lookback size, which denotes the temporal window utilized for model construction. Commonly known as the time step, the lookback specifies the number of preceding time points the model evaluates while forecasting future data. This study establishes a lookback period of 12, indicating that the model employs data from the preceding 12 frames as input to forecast the subsequent frame as output. Nonetheless, the capsule endoscope may also traverse in the retrograde path towards the stomach, potentially overlooking specific location. To resolve this issue, a reversed sequence dataset is generated by inverting the order of the original sequential dataset. The preprocessed dataset, organized according to the specified lookback and sequential bleeding data, is formatted for LSTM compatibility. The model is then trained, producing the overall accuracy and the anticipated pictures for each bleeding sequence. In conclusion, during the small intestine endoscopy, the capsule endoscope is generally inspected forward as shown in Figure 5A when it is directed toward the anus, but it is possible that the bleeding will be missed when the capsule endoscope is directed toward the stomach in the opposite direction Figure 5B, so the inspection is conducted in the opposite direction. Therefore, the sequence data set is reversely sorted and added to the original data set. The total number of original data sets is 19, and it becomes 38 after adding. The continuous sequence data is predicted by LSTM and drawn into a prediction graph. Figure 5C is the LSTM forward prediction graph and Figure 5D is the LSTM reverse prediction graph. The model's performance in this study was evaluated using accuracy, sensitivity (recall), specificity, and F1-score, with a predetermined train/validation/test split. The present experiments did not include area under the curve-receiver operating characteristic analysis, precision-recall curves, or the F2-score. The F2-score, emphasizing sensitivity, will be utilized in forthcoming studies to enhance the evaluation of bleeding detection.
In CNN models, the loss function is a crucial statistic for assessing the disparity between anticipated and actual outcomes. It is reduced during training to enhance the model's forecast accuracy. Prevalent loss functions comprise mean squared error (MSE)[29], Cross-Entropy[30], and BCE[31]. This work utilizes the categorical cross-entropy loss function[32] within the CNN model, applicable for multi-class classification tasks where output labels are represented by one-hot encoded values of 0 or 1 as presented in Equation 1:
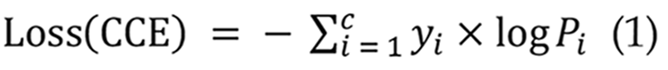
The RNN model utilizes BCE loss, which is appropriate for binary classification tasks[33]. This study employs the BCE loss function to classify the presence or absence of small intestinal bleeding. The model's output is a probability value ranging from 0 to 1, signifying the risk of hemorrhage. The BCE loss formula is presented in Equation 2:
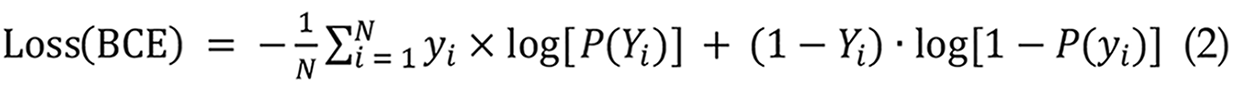
In LSTM networks, root MSE (RMSE) is a frequently utilized assessment statistic for regression models. It quantifies the disparity between expected and actual values, with reduced RMSE values signifying enhanced predictive accuracy[34]. This study establishes an RMSE threshold of 0.1, above which the predicted frames markedly diverge from the actual frames, resulting in diminished bleeding prediction efficacy as shown in Equation 3:
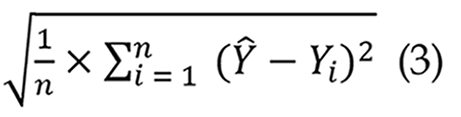
Following dataset transformation, CNN models were trained separately on the modified dataset. The dataset was analyzed in both 10-class and 2-class configurations to assess CNN’s impact on LSTM predictions. The obtained results are:
10-class: Accuracy = 98.6%, Loss = 5.45%.
2-class: Accuracy = 96.7%, Loss = 8.32%.
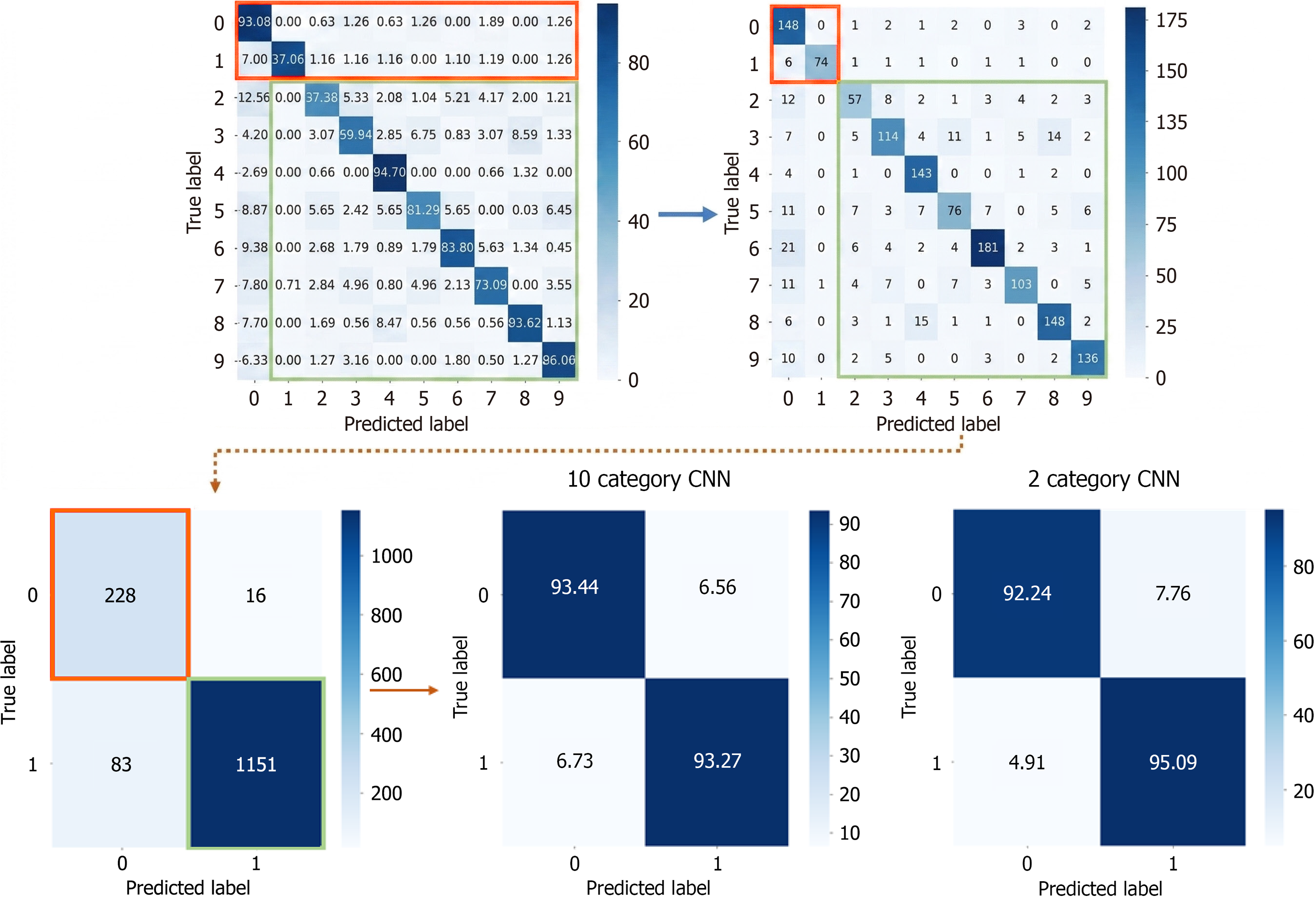
To enable a comparison examination of the 10-class and 2-class confusion matrices, the original 10-class confusion matrix was converted to a 2-class version, as illustrated in Figure 6. The transformation method entailed consolidating bleeding-related categories (classes 0-1) into one category and non-bleeding categories (classes 2-9) into another. The modified confusion matrix was subsequently compared to the original two-class confusion matrix, as seen in Figure 3, facilitating an investigation of the variations in feature extraction by the CNN concerning class quantity. To quantitatively assess the statistical significance of the differences in outcomes between the CNN models with 10 and 2 classes, the classification results was evaluated at the sample level as shown in Table 3. In the 10-class CNN model, the bleeding-related labels were consolidated into a single bleeding label, while the fiber-related classes were amalgamated into a non-bleeding category to balance the proportions. The test results were subsequently compared directly with the two-class natively trained CNN. Both matrices yielded identical results: True positives 228, false negatives 16, false positives 83, and true negatives 1151, from a total of 1478 test samples. This equivalence signifies that all test instances received identical classification labels across the two methodologies. A McNemar test was conducted to determine whether there was discordant prediction between the two models. The McNemar test evaluates the frequency of discordant cases between two models; thus, it cannot be effectively employed to determine the significance of differences in the absence of discordant pairs. This comparison revealed no discordant pairs (B = 0, C = 0), resulting in a χ2 statistic of 2 = 0.0, with a corresponding P value of 1.0. Statistically, this indicates no alteration in the categorization outcomes of the collapsed 10-style CNN relative to the direct 2-style CNN. The minor discrepancies in accuracy and loss values (98.6% vs 96.7%; 5.45% vs 8.32%) can be attributed to rounding error and presentation rather than a genuine difference in prediction ability.
| Model | Precision | Recall | F1-score | Accuracy |
| 10-class (collapsed) | 98.6% | 93.3% | 95.9% | 93.3% |
| 2-class (direct) | 98.4% | 95.1% | 96.7% | 94.7% |
The LSTM model was trained for 180 epochs. The 10-class configuration outperformed the 2-class configuration in both accuracy and loss. The results are as follows:
10-class Training Accuracy: 96.2%.
10-class Testing Accuracy: 87.1%.
2-class Training Accuracy: 82.6%.
2-class Testing Accuracy: 76.7%.
10-class Training Loss: 9.8%.
10-class Testing Loss: 31.2%.
2-class Training Loss: 41.3%.
2-class Testing Loss: 52.6%.
To illustrate LSTM predictions, time-series data of small intestine pictures were preprocessed, delineating all bleeding areas while integrating non-bleeding regions into sequential datasets, as depicted in Figure 4. The preprocessed sequences were subsequently employed for LSTM prediction and visualization. The RMSE values for each setting are delineated as follows:
10-class Training: RMSE (Forward: 0.05, Backward: 0.05).
10-class Testing: RMSE (Forward: 0.07, Backward: 0.04).
2-class Training: RMSE (Forward: 0.09, Backward: 0.18).
2-class Testing: RMSE (Forward: 0.29, Backward: 0.27).
The integration of CNNs and LSTMs has been applied in other domains, such as natural video analysis; nonetheless, it remains unexplored in capsule endoscopy and is insufficiently researched. The majority of published articles regarding the identification of bleeding in capsule endoscopy rely solely on CNN models, neglecting the temporal and sequential dimensions of the data. This research enhances the field by providing a direct comparison between CNN-only and CNN-LSTM models applied to capsule endoscopy movies, demonstrating superior efficiency in classification and prediction stability on a frame-by-frame basis. Both the conventional performance metrics (accuracy, sensitivity, specificity, F1-score, area under the receiver operating characteristic) and temporal error metrics (forward and reverse RMSE) are presented, highlighting that LSTMs can effectively model clinically significant bleeding dynamics that CNN-only models fail to capture adequately. This work is innovative as it clearly illustrates that analyzing capsule endoscopy data through temporal dependencies produces measurable and clinically significant improvements, thereby facilitating more effective and reliable detection of bleeding in practical diagnostic procedures. This study's findings indicate that the utilization of a CNN affects the prediction efficacy of an LSTM network. The classification accuracy of the CNN model was assessed for both a 10-class dataset and a 2-class dataset to study this influence. The categorization accuracy between bleeding and non-bleeding categories is similar. The CNN model attains an accuracy of 98.6% and a loss function value of 5.45% when utilized on the original 10-category dataset. When the dataset is condensed to two categories, the accuracy diminishes somewhat to 96.7%, but the loss function value escalates to 8.32%. An examination of the confusion matrices for both scenarios indicates that the classification performance remains consistently stable, implying that the feature extraction abilities of CNNs are not substantially influenced by the number of categories. This observation underscores the efficacy of CNN in deriving significant characteristics from medical imaging data. The retrieved features from both the 10-category and 2-category datasets were employed for LSTM training and comparative analysis. The second continuous image database, obtained from the CNN-trained 10-category and 2-category models, was utilized for CNN-based feature extraction, subsequently followed by LSTM-based prediction. A notable disparity in prediction efficacy is evident when use LSTM. The total accuracy of LSTM training and testing for the 10-category dataset is 96.2% and 87.1%, respectively, however for the 2-category dataset, these figures decline to 82.6% and 76.7%. The results demonstrate a significant difference in predicted accuracy between the two dataset setups. The decline in accuracy while utilizing LSTM on the binary dataset can be ascribed to the characteristics of feature extraction in CNN and the temporal relationships that LSTM captures. As the number of categories processed by the CNN decreases, the complexity of the retrieved features diminishes. Nonetheless, due to CNN's robust feature extraction abilities, the classification accuracy for the binary dataset remained comparatively elevated. Conversely, LSTM's capacity to apprehend dynamic temporal fluctuations and localized variations is constrained. This shortcoming is exacerbated when utilizing simplified features, as LSTM fails to adequately capture essential information pertaining to bleeding conditions. As a result, the predictive accuracy of LSTM declines when utilized on a dataset with fewer categories. The results of LSTM training and test predictions further validate these conclusions. A reduction in the number of categories results in diminished predictive performance, as indicated by an escalation in the RMSE. With an RMSE threshold of 0.1, the predictive model is unable to reliably detect bleeding regions, leading to classification inaccuracies. The model demonstrates significant misclassification between bleeding and non-bleeding locations, resulting in inconsistency in the prediction outcomes. This study's findings underscore the efficacy of combining CNN and LSTM models for predicting images of minor intestinal hemorrhage. The capacity of CNN to extract spatial information, coupled with LSTM's proficiency in capturing temporal relationships, indicates a viable methodology for medical image analysis. Future study should concentrate on refining the amal
The examination of GI hemorrhage continues to provide a considerable difficulty for endoscopic experts. This research introduces an innovative method for forecasting hemorrhage in small intestine images obtained using capsule endoscopy. Utilizing deep learning methodologies, individual video frames were systematically evaluated and defined, enabling the processed sequential and feature-based data to be utilized with LSTM networks. The results indicated a decrease in the intrinsic challenges of capsule endoscopy, specifically the laborious and time-consuming process of manual observation for identifying bleeding sites. The utilization of LSTM networks in medical research is constrained, as the majority of deep learning analyses predominantly depend on CNNs for image processing. In clinical practice, endoscopic examinations utilize continuous imaging instead of static images, and prognostic methods for small intestinal hemorrhage are yet inadequately developed. This study combines the predictive skills of LSTM with the feature extraction proficiency of deep learning, facilitating the identification of bleeding in sequential imaging data. With further development and integration with next-generation endoscopic technology, this technique could markedly improve the patient experience, expedite diagnostic processes, and ultimately lead to more efficient and successful medical treatment. Recent research has shown that AI can perform diagnostic tasks in capsule endoscopy at an expert level or even better. These results show that AI has a lot of potential to make diagnoses faster and easier for doctors. Nonetheless, the current study does not encompass direct comparisons with expert readers or commercial software platforms. So, the results should be seen as a technical proof-of-concept instead of proof that the two treatments are the same or ready for use in the clinic. Along with performance issues, the clinical use of AI systems in capsule endoscopy needs to take into account new risks, such as automation bias and cybersecurity weaknesses that come with medical AI systems. These elements, while outside the purview of the current technical analysis, signify crucial avenues for forthcoming interdisciplinary research. A major limitation of this study is the lack of clinically oriented validation. Specifically, there was no direct comparison of detection accuracy or reading duration between the proposed model and manual assessments by gastroenterologists. Moreover, no comparisons were made with existing commercial capsule endoscopy analysis tools. These constraints restrict the current conclusions to technical feasibility rather than preparedness for clinical application. Future projects will include multi-reader studies, efficiency evaluations, and comparisons with commercial systems to determine clinical reliability and practical applicability. The experimental results demonstrate that the proposed CNN-LSTM framework achieves reliable and enhanced bleeding detection in both binary and multi-class training approaches. Despite the constant effectiveness and durability of multi-class pretraining, McNemar testing indicated no statistically significant superiority compared to direct binary training. Further study employing larger, thoroughly annotated patient datasets is essential to determine if multi-class learning provides a significant performance benefit. The 10-class CNN has marginally superior accuracy compared to the direct 2-class CNN (98.6% vs 96.7%); nonetheless, the McNemar test indicated no statistically significant difference between the two models (P = 1.0). The data indicate that multi-class training produces comparable performance instead of a statistically significant benefit in the current experimental setting. This study serves as a proof-of-concept technical investigation demonstrating the efficacy of CNN-LSTM-based automated bleeding detection. Future endeavors must prioritize comprehensive comparative benchmarking, employ a broader array of evaluation metrics, and conduct reader studies centered on clinical applications to ascertain the genuine clinical value and readiness for implementation.
| 1. | Peery AF, Dellon ES, Lund J, Crockett SD, McGowan CE, Bulsiewicz WJ, Gangarosa LM, Thiny MT, Stizenberg K, Morgan DR, Ringel Y, Kim HP, DiBonaventura MD, Carroll CF, Allen JK, Cook SF, Sandler RS, Kappelman MD, Shaheen NJ. Burden of gastrointestinal disease in the United States: 2012 update. Gastroenterology. 2012;143:1179-1187.e3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1614] [Cited by in RCA: 1485] [Article Influence: 106.1] [Reference Citation Analysis (6)] |
| 2. | Wang R, Li Z, Liu S, Zhang D. Global, regional, and national burden of 10 digestive diseases in 204 countries and territories from 1990 to 2019. Front Public Health. 2023;11:1061453. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 60] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 3. | Liu L, Towfighian S, Hila A. A Review of Locomotion Systems for Capsule Endoscopy. IEEE Rev Biomed Eng. 2015;8:138-151. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 38] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 4. | Hosoe N, Takabayashi K, Ogata H, Kanai T. Capsule endoscopy for small-intestinal disorders: Current status. Dig Endosc. 2019;31:498-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 68] [Article Influence: 9.7] [Reference Citation Analysis (1)] |
| 5. | Aoki T, Yamada A, Kato Y, Saito H, Tsuboi A, Nakada A, Niikura R, Fujishiro M, Oka S, Ishihara S, Matsuda T, Nakahori M, Tanaka S, Koike K, Tada T. Automatic detection of blood content in capsule endoscopy images based on a deep convolutional neural network. J Gastroenterol Hepatol. 2020;35:1196-1200. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 77] [Article Influence: 12.8] [Reference Citation Analysis (1)] |
| 6. | Boal Carvalho P, Magalhães J, Dias DE Castro F, Monteiro S, Rosa B, Moreira MJ, Cotter J. Suspected blood indicator in capsule endoscopy: a valuable tool for gastrointestinal bleeding diagnosis. Arq Gastroenterol. 2017;54:16-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 17] [Article Influence: 1.9] [Reference Citation Analysis (1)] |
| 7. | Kim SE, Kim HJ, Koh M, Kim MC, Kim JS, Nam JH, Cho YK, Choe AR; Research Group for Capsule Endoscopy and Enteroscopy of the Korean Society of Gastrointestinal Endoscopy. A practical approach for small bowel bleeding. Clin Endosc. 2023;56:283-289. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 12] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 8. | Dally WJ, Keckler SW, Kirk DB. Evolution of the Graphics Processing Unit (GPU). IEEE Micro. 2021;41:42-51. [DOI] [Full Text] |
| 9. | Wang YE, Wei GY, Brooks D. Benchmarking TPU, GPU, and CPU platforms for deep learning. 2019 Preprint. Available from: arXiv:1907.10701. [DOI] [Full Text] |
| 10. | Kim H, Nam H, Jung W, Lee J. Performance analysis of CNN frameworks for GPUs. 2017 IEEE International Symposium on Performance Analysis of Systems and Software (ISPASS); 2017 Apr 24-25; Santa Rosa, CA, United States. IEEE, 2017. [DOI] [Full Text] |
| 11. | Salehinejad H, Sankar S, Barfett J, Colak E, Valaee S. Recent advances in recurrent neural networks. 2017 Preprint. Available from: arXiv:1801.01078. [DOI] [Full Text] |
| 12. | Medsker L, Jain LC. Recurrent Neural Networks: Design and Applications. 1st ed. Boca Raton: CRC Press, 1999. [DOI] [Full Text] |
| 13. | Tarwani KM, Edem S. Survey on Recurrent Neural Network in Natural Language Processing. Int J Eng Trends Technol. 2017;48:301-304. [DOI] [Full Text] |
| 14. | Zhao JY, Huang FQ, Lv J, Duan YJ, Qin Z, Li GD, Tian GJ. Do RNN and LSTM have long memory? 2020 Preprint. Available from: arXiv:2006.03860. [DOI] [Full Text] |
| 15. | Jeon W, Ko G, Lee J, Lee H, Ha D, Ro WW. Deep learning with GPUs. Adv Comput. 2021;12:167-215. [DOI] [Full Text] |
| 16. | Mukhtorov D, Rakhmonova M, Muksimova S, Cho YI. Endoscopic Image Classification Based on Explainable Deep Learning. Sensors (Basel). 2023;23:3176. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 75] [Cited by in RCA: 23] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 17. | Al-Bayati K, Stone JK, Berzin TM. The Use of Artificial Intelligence for Endoscopic Evaluation of the Small Bowel. Gastrointest Endosc Clin N Am. 2025;35:355-366. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 18. | Hayat MT, Allawi YM, Alamro W, Sultan SM, Abadleh A, Kang H, Zreikat AI. A Hybrid Convolutional Neural Network-Long Short-Term Memory (CNN-LSTM)-Attention Model Architecture for Precise Medical Image Analysis and Disease Diagnosis. Diagnostics (Basel). 2025;15:2673. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 19. | Paderno A, Gennarini F, Sordi A, Montenegro C, Lancini D, Villani FP, Moccia S, Piazza C. Artificial intelligence in clinical endoscopy: Insights in the field of videomics. Front Surg. 2022;9:933297. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 20] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 20. | Florencio de Mesquita C, Antunes VL, Milioli NJ, Fernandes MV, Correa TL, Martins OC, Chavan R, Baraldo S. EUS-guided coiling plus glue injection compared with endoscopic glue injection alone in endoscopic treatment for gastric varices: a systematic review and meta-analysis. Gastrointest Endosc. 2025;101:331-340.e8. [RCA] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 19.0] [Reference Citation Analysis (1)] |
| 21. | Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, van der Laak JAWM, van Ginneken B, Sánchez CI. A survey on deep learning in medical image analysis. Med Image Anal. 2017;42:60-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11818] [Cited by in RCA: 5454] [Article Influence: 606.0] [Reference Citation Analysis (8)] |
| 22. | Mascarenhas Saraiva M, Ribeiro T, Afonso J, Ferreira JPS, Cardoso H, Andrade P, Parente MPL, Jorge RN, Macedo G. Artificial Intelligence and Capsule Endoscopy: Automatic Detection of Small Bowel Blood Content Using a Convolutional Neural Network. GE Port J Gastroenterol. 2022;29:331-338. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 12] [Article Influence: 3.0] [Reference Citation Analysis (2)] |
| 23. | Wu G, Guo Z, Li L, Wang C. Video Abnormal Event Detection Based on CNN and LSTM. 2020 IEEE 5th International Conference on Signal and Image Processing (ICSIP); 2020 Oct 23-25; Nanjing, China. IEEE, 2020: 334-338. [DOI] [Full Text] |
| 24. | Gupta R, Gupta A, Aswal R. Time-CNN and Stacked LSTM for Posture Classification. 2021 International Conference on Computer Communication and Informatics (ICCCI); 2021 Jan 27-29; Coimbatore, India. IEEE, 2021: 1-5. [DOI] [Full Text] |
| 25. | Zan H, Zhao G. Human Action Recognition Research Based on Fusion TS-CNN and LSTM Networks. Arab J Sci Eng. 2023;48:2331-2345. [RCA] [DOI] [Full Text] [Reference Citation Analysis (0)] |
| 26. | Nie J, Liu R, Mahasseni B, Mitra V. Model-Driven Heart Rate Estimation and Heart Murmur Detection Based On Phonocardiogram. 2024 IEEE 34th International Workshop on Machine Learning for Signal Processing (MLSP); 2024 Sep 22-25; London, United Kingdom. IEEE, 2024: 1-6. [DOI] [Full Text] |
| 27. | Omori T, Hara T, Sakasai S, Kambayashi H, Murasugi S, Ito A, Nakamura S, Tokushige K. Does the PillCam SB3 capsule endoscopy system improve image reading efficiency irrespective of experience? A pilot study. Endosc Int Open. 2018;6:E669-E675. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 28. | Smedsrud PH, Thambawita V, Hicks SA, Gjestang H, Nedrejord OO, Næss E, Borgli H, Jha D, Berstad TJD, Eskeland SL, Lux M, Espeland H, Petlund A, Nguyen DTD, Garcia-Ceja E, Johansen D, Schmidt PT, Toth E, Hammer HL, de Lange T, Riegler MA, Halvorsen P. Kvasir-Capsule, a video capsule endoscopy dataset. Sci Data. 2021;8:142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 221] [Cited by in RCA: 73] [Article Influence: 14.6] [Reference Citation Analysis (2)] |
| 29. | Ma Y, Feng P, He P, Long Z, Wei B. Low-dose CT with a deep convolutional neural network blocks model using mean squared error loss and structural similar loss. Eleventh International Conference on Information Optics and Photonics (CIOP 2019); 2019; Xi'an, China. Society of Photo-Optical Instrumentation Engineers (SPIE), 2019: 112090I. [DOI] [Full Text] |
| 30. | Ruby AU, Theerthagiri P, Jacob IJ, Vamsidhar Y. Binary cross entropy with deep learning technique for Image classification. Int J Adv Trends Comput Sci Eng. 2020;9:5393-5397. [DOI] [Full Text] |
| 31. | Kim Y, Kim S, Kim T, Kim C. CNN-Based Semantic Segmentation Using Level Set Loss. 2019 IEEE Winter Conference on Applications of Computer Vision (WACV); 2019 Jan 07-11; Waikoloa, HI, United States. IEEE, 2019: 1752-1760. [DOI] [Full Text] |
| 32. | Mostafa AH, Abdel-Galil H, Belal M. Ensemble Model-based Weighted Categorical Cross-entropy Loss for Facial Expression Recognition. 2021 Tenth International Conference on Intelligent Computing and Information Systems (ICICIS); 2021 Dec 05-07; Cairo, Egypt. IEEE, 2021: 165-171. [DOI] [Full Text] |
| 33. | Mishra M, Patil A. Sentiment Prediction of IMDb Movie Reviews Using CNN-LSTM Approach. 2023 International Conference on Control, Communication and Computing (ICCC); 2023 May 19-21; Thiruvananthapuram, India. IEEE, 2023: 1-6. [DOI] [Full Text] |
| 34. | Suresh V, Aksan F, Janik P, Sikorski T, Revathi BS. Probabilistic LSTM-Autoencoder Based Hour-Ahead Solar Power Forecasting Model for Intra-Day Electricity Market Participation: A Polish Case Study. IEEE Access. 2022;10:110628-110638. [DOI] [Full Text] |