Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.114778
Revised: November 27, 2025
Accepted: February 2, 2026
Published online: April 21, 2026
Processing time: 185 Days and 19.1 Hours
Pyrrolizidine-alkaloid induced hepatic sinusoidal obstruction syndrome (PA-HSOS) is a rare and severe drug-induced liver injury with nonspecific manifestations. Its diagnosis currently relies on exclusive strategies and often necessitates invasive examinations, posing significant clinical challenges. The potential role of artificial intelligence algorithms in diagnosing PA-HSOS remains to be established.
To develop and validate a deep-learning-based diagnostic model for PA-HSOS using computed tomography images.
This multicenter case-control study compared PA-HSOS patients with Budd-Chiari syndrome and hepatitis B cirrhosis patients as controls. Patients from Zhongshan Hospital, Fudan University were retrospectively assigned to training or internal test cohorts, while those from the First Affiliated Hospital of Zhengzhou University formed an external cohort. We constructed the diagnostic models using multiscale convolutional modules. Model per
Diagnostic models with deep learning methods using computed tomography images for PA-HSOS were deve
The deep learning model demonstrates promising diagnostic performance for PA-HSOS and can effectively assist clinicians in improving diagnostic accuracy and efficiency.
Core Tip: This study developed the first deep learning model for diagnosing pyrrolizidine-alkaloid induced hepatic sinusoidal obstruction syndrome based on computed tomography images. The model integrates multiscale convolutional modules and an anatomy-based region of interest sampling strategy. Initial validation showed promising diagnostic performance, with potential to improve diagnostic consistency among clinicians and reduce image interpretation time, suggesting its possible utility as a clinical decision-support tool.
- Citation: Wang SY, Yin SQ, Yang JY, Ji MY, Zeng XQ, Rao SX, Lv MZ, Bao J, Wang MN, Gao H. Development and validation of a deep-learning-based diagnostic model for drug-induced liver injury using computed tomography images. World J Gastroenterol 2026; 32(15): 114778
- URL: https://www.wjgnet.com/1007-9327/full/v32/i15/114778.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i15.114778
Drug-induced liver injury (DILI) is a leading cause of acute liver failure[1]. The diagnostic strategy for DILI is exclusive, which is usually challenging for doctors. Pyrrolizidine alkaloids (PAs), widely present plant toxins, can cause severe vascular DILI, known as hepatic sinusoidal obstruction syndrome (PA-HSOS). The manifestations of HSOS are nonspecific, such as hepatomegaly, hyperbilirubinemia and ascites with high mortality rates ranging from 16% to 40%[2-4]. Accurate medication history review is important for diagnosis of PA-HSOS. However, patients often struggle to accurately recall their history of taking herbs containing PAs. Therefore, the diagnosis of HSOS relies on excluding other diseases; for example, Budd-Chiari syndrome using invasive methods such as liver biopsy, and digital subtraction angiography.
Enhanced computed tomography (CT) and magnetic resonance imaging (MRI) findings are essential for the diagnosis of PA-HSOS[3]. Compared with MRI, enhanced CT is more commonly utilized because of its high quality as well as time and cost saving. More than 90% of patients with PA-HSOS exhibit imaging features such as liver enlargement, uneven low-density, delayed phase heterogeneous enhancement of liver maps, and compression thinning of the inferior vena cava in the liver segment[4]. Typical imaging findings have been regarded as one of the diagnostic items for nonpathological pathways[5], especially for those who cannot tolerate such aggressive examinations such as liver biopsy and angiography due to abnormal coagulation function, thrombocytopenia, and large amounts of ascites. CT-patchy liver enhancement showed a sensitivity of 93.0% and specificity of 92.8%, and CT-heterogeneous hypoattenuation showed a sensitivity of 100% and specificity of 95.1%, which can distinguish HSOS from hepatic cirrhosis and Budd-Chiari syndrome[6,7]. However, in clinical practice, the radiological diagnosis of PA-HSOS often poses a challenge due to its reliance on the differing levels of experience and expertise among radiologists. In addition, early, mild or moderate abnormalities can be missed or difficult to determine.
With the advancement of deep learning technology, artificial intelligence (AI)-assisted techniques have been widely applied in hepatology and have shown potential in classifying liver diseases[8,9]. These techniques have demonstrated satisfactory results for hepatocellular carcinoma, nonalcoholic fatty liver disease, tumors and fibrosis[10], validating the powerful feature extraction and information mining capabilities of deep learning models.
Some studies have also applied deep learning algorithms to DILI prediction[11], imaging analysis[12], and pathological findings in rat DILI models[13,14]. Unlike liver-tumor-related diseases, clinical diagnosis of PA-HSOS is not contingent on any specific, predefined pattern such as patchy enhancement or heterogeneous hypoattenuation on CT. Traditional clinical deep learning models typically rely on commonly used medical imaging architectures such as U-Net[15] or SwinUNet[16] for feature extraction, modeling and prediction. However, these architectures generally require large training datasets and lack specialized mechanisms for extracting and integrating multiscale information in one module (typically only relying on hierarchical layers) within a single image. Thus, constructing a deep-learning-based diagnosis model for PA-HSOS is challenging; particularly when differentiating it from easily confused conditions such as hepatitis B virus-related liver disease or Budd-Chiari syndrome. Given the difficulty in collecting PA-HSOS cases, the dataset is limited, and imaging diagnosis itself is challenging, deep-learning-based diagnostic models for PA-HSOS have not been explored to date. The deep-learning-based models have the potential to substantially improve the efficiency and accuracy of PA-HSOS diagnosis, offering considerable clinical value. The primary challenge lies in effectively extracting imaging features related to diffuse parenchymal abnormalities and aligning the model design with the expert diagnostic workflow to enable accurate prediction. This study aimed to establish and validate a deep learning-based PA-HSOS diagnostic model that can provide valuable auxiliary diagnostic value.
A case-control study was performed using PA-HSOS patients as cases, and Budd-Chiari syndrome and hepatitis B cirrhosis patients, which are difficult to distinguish from PA-HSOS, as controls. Patients with PA-HSOS, Budd-Chiari syndrome or hepatitis B cirrhosis admitted to Zhongshan Hospital, Fudan University, between June 2008 and September 2022 were retrospectively enrolled into the training cohort or the internal test cohort. Patients retrospectively and prospectively enrolled at the First Affiliated Hospital of Zhengzhou University were evaluated as an external cohort. Considering the low prevalence of PA-HSOS, all eligible patients were included to maximize feasibility. Additionally, controls were selected from Zhongshan Hospital database during the same study period to roughly match the cases on key demographic factors and to ensure the availability of high-quality contrast-enhanced CT images.
All patients were aged 18-75 years, with contrast-enhanced abdominal CT or hepatic vascular enhanced CT images. PA-HSOS was diagnosed based on the Nanjing criteria[5]. Budd-Chiari syndrome and hepatitis B cirrhosis were diagnosed based on the clinical practice guidelines of European Association for Liver Research[17,18].
The detailed diagnostic criteria: (1) PA-HSOS. Confirmed history of PA-containing plant ingestion, plus meeting all three of the following criteria or having pathological evidence, while excluding other known causes of liver injury: Abdominal distension and/or right upper quadrant pain, hepatomegaly, ascites; elevated serum total bilirubin or abnormal liver function tests; and characteristic findings on contrast-enhanced CT or MRI. CT imaging was evaluated by two radiologists specializing in abdominal imaging with over 10 years of experience. A consensus between the two radiologists was required. In instances of discordance, a senior radiologist’s perspective was sought for consultation. Pathological confirmation includes swelling, damage, and shedding of hepatic sinusoidal endothelial cells in the hepatic acinus zone III and significant dilation and congestion of hepatic sinusoids; (2) Budd-Chiari syndrome. Diagnosis was established with unequivocal radiological confirmation of hepatic venous outflow obstruction, including Doppler ultrasound, MRI, CT evaluation or venography; and (3) Hepatitis B cirrhosis. Diagnosis was confirmed by a documented history or serological evidence of hepatitis B virus infection and liver biopsy or imaging findings consistent with cirrhosis, after exclusion of other etiologies. The control groups were screened to ensure they had no history of PA-containing drug or Chinese herbal medicine use.
The exclusion criteria: (1) More than two types of chronic liver diseases, such as concomitant primary sclerosing cho
To mitigate the risk of circular reasoning, the diagnostic labels (the “ground truth”) for all patients were established independently of the CT imaging features used for deep learning. The reference standard for PA-HSOS, Budd-Chiari syndrome and hepatitis B cirrhosis was based solely on comprehensive clinical criteria prior to and without any reliance on the model’s analysis.
Dual-phase enhanced abdominal scanning was performed using scanners from two medical centers. The internal cohort (Zhongshan Hospital) utilized SOMATOM Definition AS, SOMATOM Definition AS+ (Siemens Medical Systems, Germany), or Toshiba Aquilion ONE (Toshiba Medical Systems, Japan), while the external cohort (The First Affiliated Hospital of Zhengzhou University) used Discovery CT 750HD (GE Medical Systems, WI, United States) and Somatom Force (Siemens Medical Systems, Germany). The scanning range extended from the top of the diaphragm to the lower renal edges. Scanning parameters included: Tube voltage 120 kV; tube current 150-250 mA (internal cohort) or 220-330 mA/automatic modulation (external cohort); scanning layer thickness 1.5 mm (internal cohort) or 1.25 mm (external cohort); rotation time 0.5 second (internal cohort) or 0.5-0.8 second (external cohort). A nonionic point contrast agent was administered intravenously at a rate of 2.5 mL/second (internal cohort) or 3.5 mL/second (external cohort), at a dose of 1.5 mL/kg. The internal cohort underwent triphasic scanning at 35-40 seconds, 80-90 seconds and 120 seconds post-injection for arterial, portal venous and delayed phases, respectively, with the external cohort being scanned at 25-30 seconds (arterial phase) and 60 seconds (venous phase).
ITK-SNAP (version 3.6.0, http://www.itksnap.org) was used to randomly annotate one point in each liver segment (nine locations in total) and three hepatic venous reflux areas, resulting in 12 points for each patient. This process was conducted across three-dimensional (3D) volumes containing the liver in both portal and venous phases, using a reconstruction layer thickness of 5 mm, and excluding conspicuous cysts and large blood vessels. Annotation outcomes are reported in the NIFTI format. All images were resampled at a spacing of 1 mm × 1 mm × 1 mm and reoriented.
Development of a diagnostic model for PA-HSOS is challenging primarily due to two factors: (1) Feature extraction, effectively capturing imaging characteristics associated with diffuse parenchymal abnormalities; and (2) Alignment with the clinical diagnostic workflow, improving the interpretability and clinical relevance of the model.
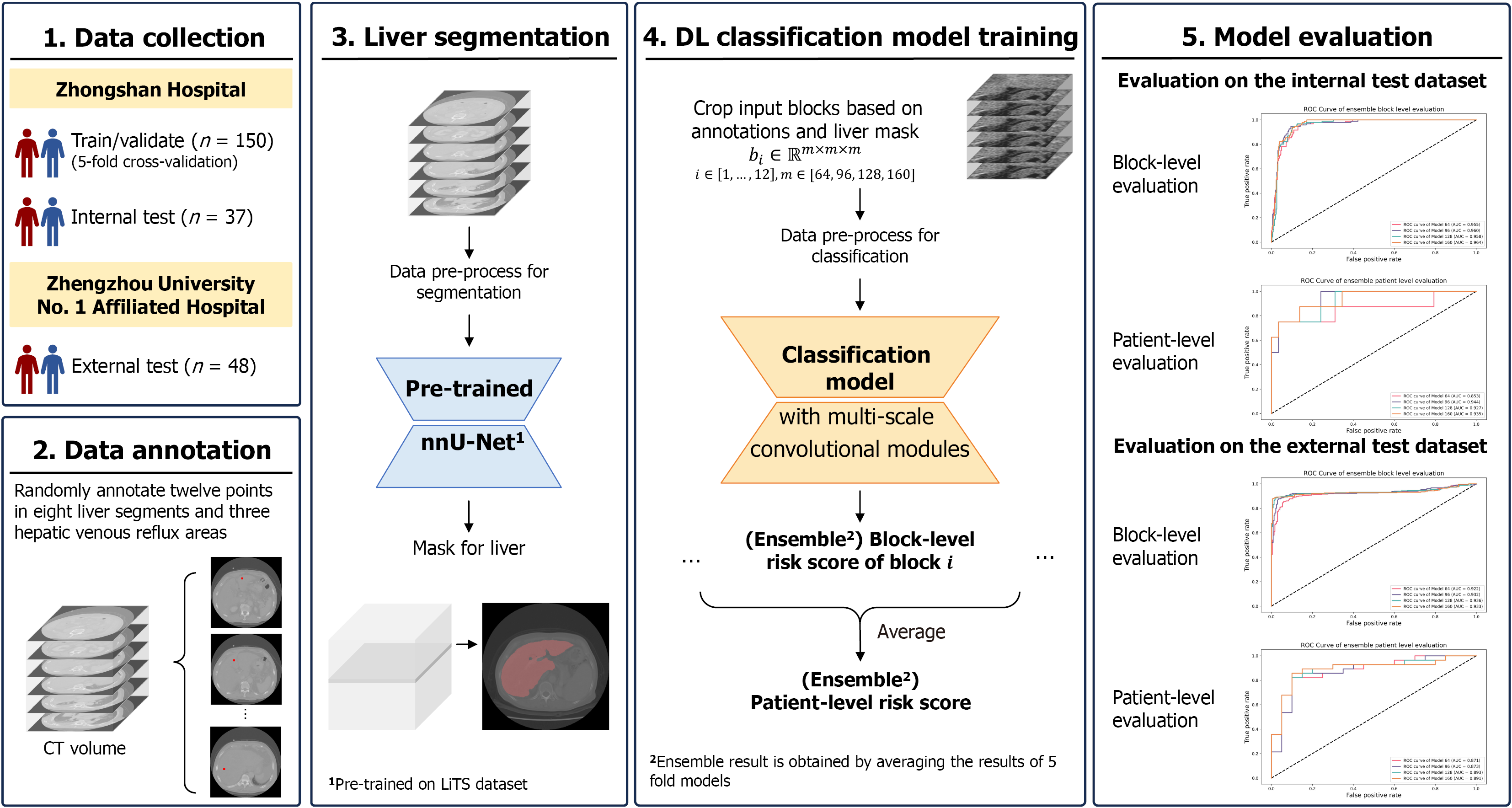
To address these challenges, we designed a two-stage framework (auto-segmentation + classification) (Figure 1). Stage 1, a transfer learning-based auto-segmentation model was used to extract the liver region from CT images. This step did not require any manual delineation or additional training, and the resulting liver mask was used to suppress nonhepatic regions during subsequent region of interest (ROI) block sampling, thereby reducing irrelevant noise. ROIs were sampled using anatomical annotation points-based sampling strategy, which collected ROIs that corresponded to regions typically examined by clinical experts. This design enhanced the consistency between the deep learning pipeline and the clinical diagnostic process, thereby improving interpretability. Stage 2, the classification model was built upon multiscale convolutional modules to perform ROI block level PA-HSOS risk prediction. This design enabled the extraction of informative features across multiple receptive fields, thereby enhancing robustness in identifying diffuse parenchymal abnormalities. For each patient, the model outputted an aggregated patient-level risk score as well as a final binary classification result.
To minimize the impact of extraneous organs on PA-HSOS diagnosis in the deep learning model, liver segmentation was first conducted on the abdominal CT images. As manual annotation of the liver is both time-consuming and labor-intensive, nnU-Net[19] based on transfer learning was used to facilitate automatic end-to-end liver segmentation. nnU-Net uses the classic U-net[15] as the backbone for semantic segmentation of 3D medical image data. With portal phase enhanced CT full image data as input, the model produced segmented portal phase data for the entire liver. Model weights pretrained on the LiTS dataset were utilized for liver segmentation. The segmentation results were used as masks on the original CT images, based on which 12 3D ROIs were cropped for each patient centered at every point annotation as inputs for the classification model. For each ROI, window widths and levels of 150 and 45, respectively, were utilized to adjust the image, which was then normalized as 0 to 1. Visualization of ROIs can be found in Supplementary Figure 1.
The deep-learning-based diagnostic model consisted of five layers of multiscale convolution modules. Each multiscale convolutional module used convolution kernels of different sizes (3 and 5, respectively) to extract image features; after which, the outputs were concatenated along the channel dimension. This process was repeated to achieve the goal of extracting and integrating information across multiple scales, improving the performance of the classification model. The detailed model architecture is shown in Supplementary Figure 2.
For each patient, 12 ROIs were used as inputs, and the model predicted the probability value within the range of 0 to 1 for each ROI, indicating the likelihood of the presence of PA-HSOS within the input ROI. The average result of the 12 ROIs was used for the patient-level prediction. To investigate the impact of the ROI size on model performance, four different sizes of ROI were cropped to cube lengths of 64 mm, 96 mm, 128 mm, and 160 mm. Finally, four deep-learning-based diagnostic models for PA-HSOS were trained based on ROIs of different input sizes, denoted as model 64, model 96, model 128 and model 160, respectively. The model ultimately outputs a probability value, representing the likelihood that the current input belongs to a PA-HSOS patient. The model was optimized using binary cross-entropy focal loss.
Patients in Zhongshan Hospital, Fudan University were randomly divided into training and internal test cohorts in an 8:2 ratio. For all the models, K-fold cross validation (K = 5) was applied to the training cohort. The data were further divided into five sets; each of which was used as a validation cohort, leaving the remaining four as a training cohort. The splitting of K-fold cross-validation was achieved at the patient level, ensuring that several ROIs (blocks) of one patient were divided into the same set. Subsequently, the models were tested using an external test cohort at the First Affiliated Hospital of Zhengzhou University. The external validation cohort was completely held out from all training and validation processes. The final model weights and preprocessing pipelines were fixed prior to evaluation on this cohort.
The learning rate for the classification task was set to 3 × 10-5, and the model was trained for 200 epochs using the stochastic gradient descent optimizer. An early stop strategy with 50 epochs was implemented. To prevent overfitting, a data augmentation strategy was used for the training dataset, with a probability of 0.5 for horizontal flipping and random rotation of each image. The experiments were conducted on a single NVIDIA 3090 GPU with PyTorch 1.12.1.
To evaluate the diagnostic performance of the developed models, three attending gastroenterologists (AGs), three residents of internal medicine (RIMs), three attending radiologists and three residents of radiology (RRs) were invited to diagnose the patients in the internal test cohort (n = 37), with or without model assistance (with at least 1-month interval). The readers were selected based on their professional roles and clinical seniority, including board-certified specialists with > 5 years of experience and residents in their second or third year of training. All readers were blinded to clinical data, laboratory results and original diagnoses, reviewing only anonymized DICOM images using ITK-SNAP (version 3.6.0) under standardized viewing conditions. To minimize recall bias, the case order was fully randomized for each reading session, with a washout period of at least 1 month between unassisted and model-assisted readings.
Prior to the model-assisted reading session, all participating physicians received a brief, standardized instruction. They were informed that they would review their initial diagnoses with the assistance of the output of the AI model, which would consist of a binary classification (PA-HSOS or non-PA-HSOS) and a corresponding probability score. It was emphasized that the output of the model should be considered as a decision-support tool, and the final diagnostic decision remained entirely at their discretion. No specific training on the features or rationale of the model was provided. During the model-assisted reading session, the output of model 96 was presented to the physicians in a numerical and categorical format without any visual overlays (e.g., heatmaps or ROI delineations) on the original CT images. For each patient, the provided output included: (1) A binary prediction (PA-HSOS or non-PA-HSOS) determined by the optimal threshold from the Youden index; and (2) A continuous probability value (ranging from 0 to 1) indicating the reliability of the model in prediction of PA-HSOS. The doctors independently reviewed the images, and none were involved in the original diagnostic process of the enrolled patients. Diagnostic results and time consumption were recorded.
During the training phase, the model with the highest accuracy in the validation cohort was selected as the optimal model for each fold. During the testing phase, the best model for each fold was used to predict the probability value for each ROI. Each model was evaluated at block level and patient level. Block-level prediction refers to the prediction probability value of each ROI input that was directly obtained from the model. Patient-level prediction refers to the averaged probability across the 12 ROIs for a given patient. Subsequently, an ensemble strategy was used, averaging the prediction results of the fivefold models to obtain a single result for each ROI input as the ensembled block-level prediction. Similarly, an ensembled patient-level prediction was obtained by averaging the ensembled block-level results for each patient. The receiver operating characteristic curve and area under the curve (AUC) were calculated using the ensembled result and label.
Statistical analysis was performed using SPSS Statistics 26.0, Python 3.8.12 and GraphPad Prism 8. Quantitative data following a normal distribution are presented as mean ± SD, while data not conforming to a normal distribution are expressed as median (25th percentile, 75th percentile). The Youden index was used to determine the optimal threshold for binarization of the predicted positive (PA-HSOS) or negative (non-PA-HSOS) results. Model performance was assessed using metrics including AUC, accuracy, sensitivity, specificity, Youden index, kappa value, positive predictive value and negative predictive value. The McNemar test was used to compare the model accuracy, sensitivity and specificity with the corresponding values from the doctors. The difference in physicians’ diagnostic time with and without model assistance was compared using a paired samples t test. Statistical significance was set at P < 0.05.
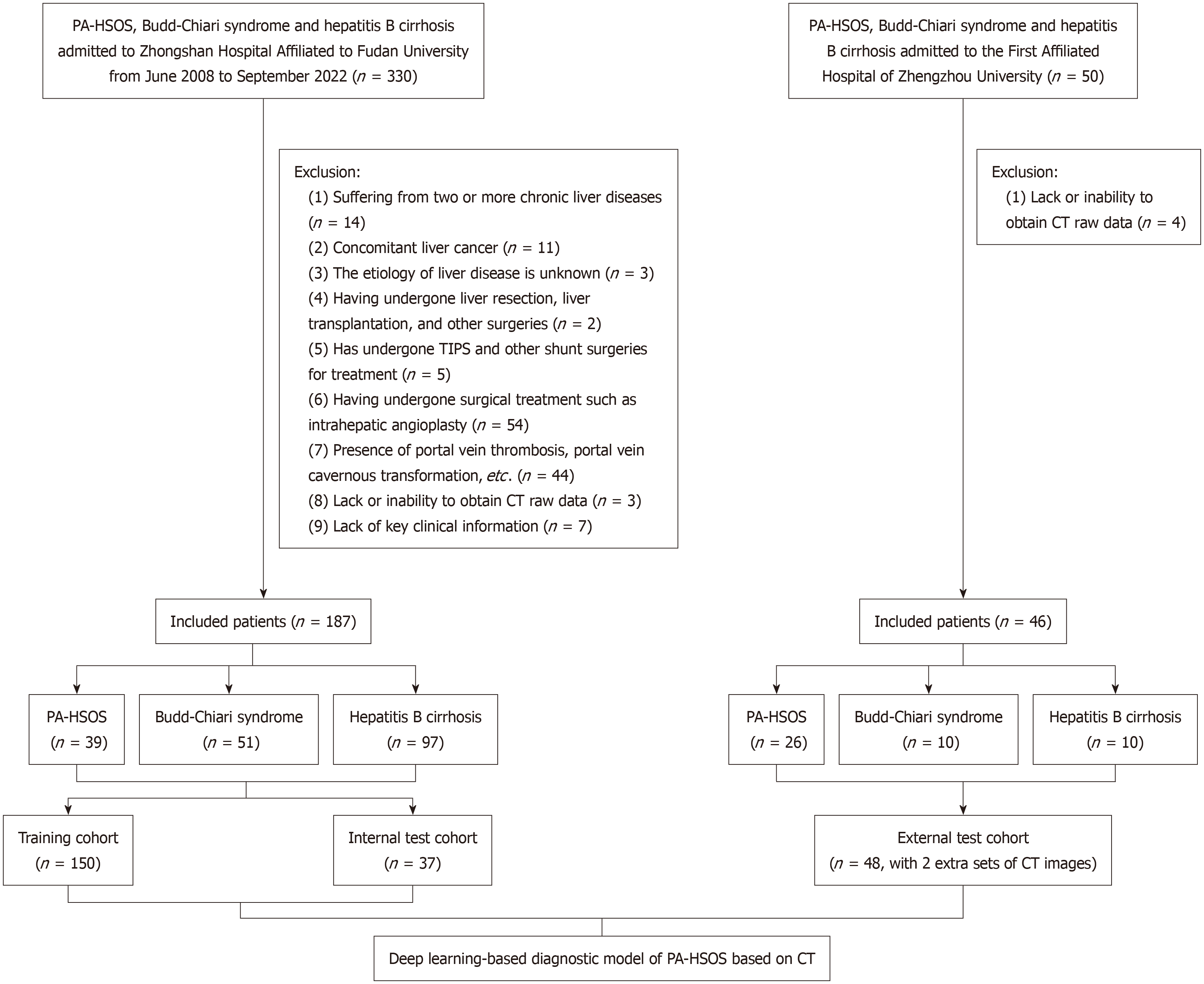
A total of 235 sets of abdominal CT images were collected from 67 patients with PA-HSOS, 61 with Budd-Chiari syndrome and 107 with hepatitis B cirrhosis (Figure 2). From the portal venous phase of each patient’s contrast-enhanced CT examination, which was selected for its optimal demonstration of the heterogeneous parenchymal enhancement characteristic of PA-HSOS, 12 3D ROIs were extracted. The deep learning analysis was performed on 2820 ROIs (235 patients × 12 ROIs) across the entire dataset. Of the 235 patients, 187 from Zhongshan Hospital were randomly assigned to the training (n = 150) and internal test (n = 37) cohorts, while 48 from The First Affiliated Hospital of Zhengzhou University formed the external test cohort. The demographic characteristics are shown in Table 1. Distribution of patient cohorts and disease types are shown in Table 2.
| Hospital group | Diagnostic group | Male | Age (years) |
| Zhongshan cohort | PA-HSOS (n = 39) | 27 (69.23) | 62.62 ± 12.33 |
| Budd-Chiari syndrome (n = 51) | 29 (56.86) | 44.98 ± 11.84 | |
| Hepatitis B cirrhosis (n = 97) | 73 (75.26) | 53.29 ± 11.12 | |
| Total (n = 187) | 129 (68.98) | 52.97 ± 13.02 | |
| Zhengzhou cohort | PA-HSOS (n = 28) | 19 (67.86) | 66.07 ± 9.25 |
| Budd-Chiari syndrome (n = 10) | 6 (60.00) | 55.9 ± 11.06 | |
| Hepatitis B cirrhosis (n = 10) | 8 (80) | 49.60 ± 13.28 | |
| Total (n = 48) | 33 (68.75) | 60.52 ± 12.43 | |
| Total | (n = 235) | 162 (68.94) | 54.37 ± 13.23 |
| Zhongshan cohort | Zhengzhou cohort | ||
| Training | Internal test | External test | |
| PA-HSOS | 31 | 8 | 28 |
| Hepatitis B cirrhosis | 78 | 19 | 10 |
| Budd-Chiari syndrome | 41 | 10 | 10 |
| Total | 150 | 37 | 48 |
Since the manual pixel-level liver annotation was not available in our dataset, the qualitative visualization results are shown in Supplementary Figure 3. Most samples achieved segmentation masks that closely matched the liver region (Supplementary Figure 3A). We also showed failures (Supplementary Figure 3B), including under-segmentation and mis-segmentation. In the under-segmentation cases (white boxes), although parts of the liver boundary were omitted, most of the liver region remained intact, so the impact on ROI extraction was minimal. In mis-segmentation cases, some nonliver tissues were incorrectly included; however, because our ROI sampling was centered on radiologist-annotated points, these areas did not fall within the ROI and therefore had no effect on the classification. Overall, from a qualitative standpoint, our segmentation results were sufficiently accurate for downstream tasks.
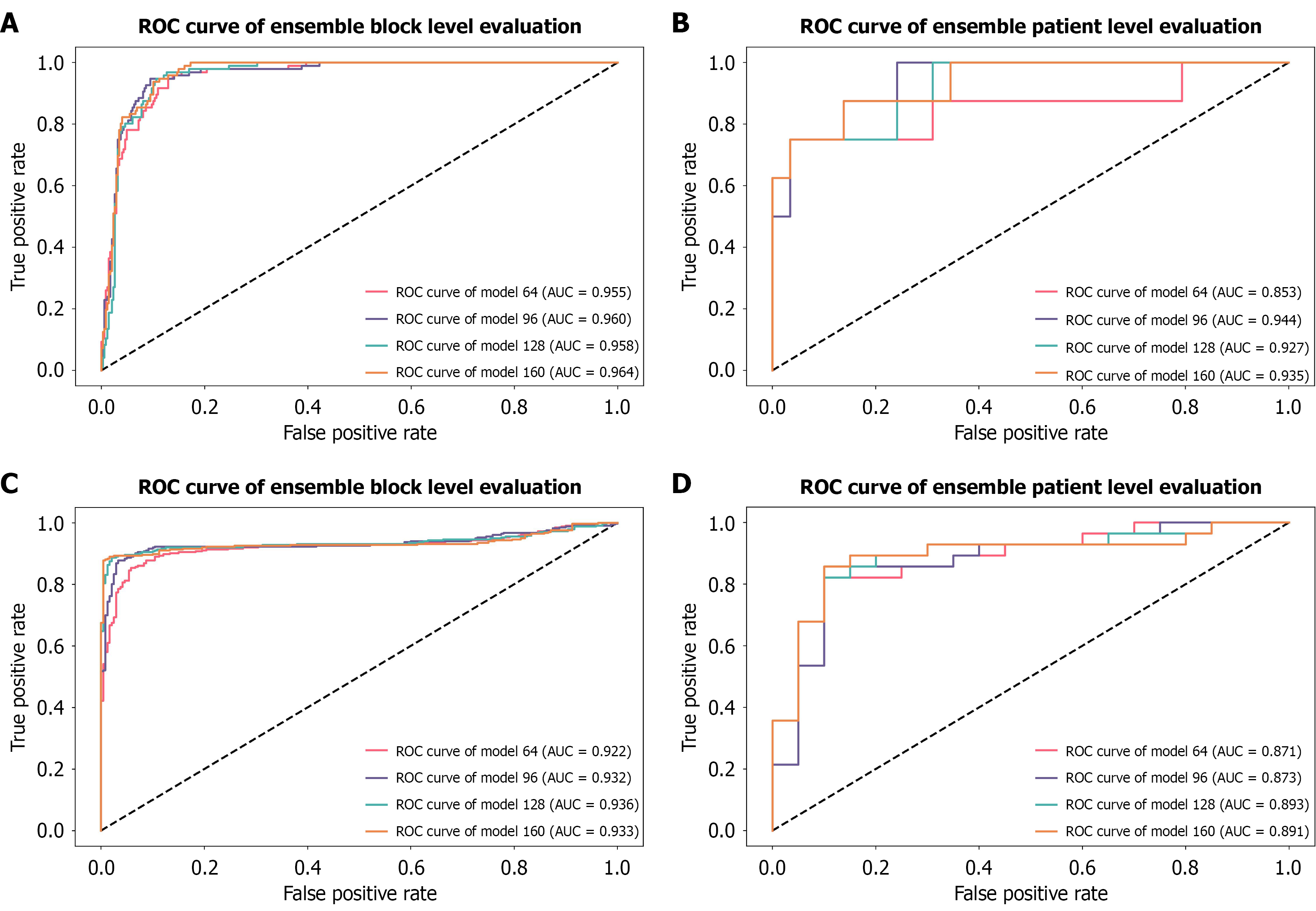
During model training, the model with the best accuracy in the validation cohort was selected and tested on internal and external test cohorts. In the internal test cohort, the block-level AUC of the four models with different input sizes ranged between 0.926 and 0.973 on the five folds (Supplementary Figure 4). After integrating the fivefold models, the block-level AUC improved, ranging between 0.955 and 0.964 (Figure 3A). And at the patient level, the AUC of the four models with different input sizes ranged between 0.853 and 0.944 (Figure 3B). The model 96 performed best, with an AUC of 0.944 (Table 3). The accuracy, sensitivity and specificity of model 96 were 0.865, 0.875 and 0.862, respectively, with a positive predictive value of 0.636 and negative predictive value of 0.962. In the external test cohort, the block-level AUC was between 0.922 and 0.936 (Figure 3C), demonstrating that the models exhibited strong diagnostic performance at the block level. The patient-level AUC of model 96 was 0.873 (Figure 3D), further suggesting that the proposed deep learning model showed satisfactory performance for PA-HSOS diagnosis (Table 3). The inference speed of the classification model was evaluated using model 96 on an NVIDIA 3090 GPU. The model required approximately 0.0738 second to predict the risk probability for a single ROI block, which corresponds to 813 ROI predictions/minute. Under the setup used in this study, where each patient had 12 ROIs, our model generated patient-level risk predictions for approximately 67 patients/minute.
| Cohort | Model | AUC (95%CI) | ACC | SEN | SPE | PPV | NPV |
| Internal | Model 64 | 0.853 (0.678-1.000) | 0.919 | 0.750 | 0.966 | 0.857 | 0.933 |
| Model 96 | 0.944 (0.829-1.000) | 0.865 | 0.875 | 0.862 | 0.636 | 0.962 | |
| Model 128 | 0.927 (0.797-1.000) | 0.919 | 0.750 | 0.966 | 0.857 | 0.933 | |
| Model 160 | 0.935 (0.813-1.000) | 0.865 | 0.875 | 0.862 | 0.636 | 0.962 | |
| External | Model 64 | 0.871 (0.772-0.971) | 0.854 | 0.821 | 0.900 | 0.920 | 0.783 |
| Model 96 | 0.873 (0.774-0.972) | 0.854 | 0.821 | 0.900 | 0.920 | 0.783 | |
| Model 128 | 0.893 (0.802-0.984) | 0.854 | 0.821 | 0.900 | 0.920 | 0.783 | |
| Model 160 | 0.891 (0.799-0.983) | 0.875 | 0.857 | 0.900 | 0.923 | 0.818 |
To quantitatively demonstrate the effectiveness of the proposed method for diagnosis of PA-HSOS, ablative experiments were conducted on key components, including presegmentation process, multiscale convolutional kernels, and ROI sampling strategy. The results of the ablation experiments are presented in Supplementary Table 1. The contribution of each component in our proposed deep-learning-based PA-HSOS diagnostic models demonstrated their effectiveness in PA-HSOS classification.
The accuracy (0.865) and specificity (0.862) of model 96 were significantly higher than those of RIMs (0.541 and 0.483, respectively) and RRs (0.676 and 0.621, respectively; P < 0.05), and similar to those of AGs and attending radiologists (P > 0.05). There was no significant difference between the sensitivity of model 96 and that of the four categories of doctors. Youden index (0.737) and kappa value (0.649) of model 96 were higher than those of all doctors (Table 4).
| Item | Accuracy (95%CI) | P value1 | Sensitivity (95%CI) | P value | Specificity (95%CI) | P value | Youden index | Kappa value |
| Residents of internal medicine | 0.541 (0.369-0.705) | 0.002b | 0.750 (0.349-0.968) | 1.000 | 0.483 (0.294-0.675) | 0.003b | 0.233 | 0.147 |
| Attending physicians | 0.730 (0.559-0.862) | 0.227 | 0.750 (0.349-0.968) | 1.000 | 0.724 (0.528-0.873) | 0.289 | 0.474 | 0.373 |
| Residents of radiology | 0.676 (0.502-0.820) | 0.039a | 0.875 (0.473-0.997) | 1.000 | 0.621 (0.423-0.793) | 0.039a | 0.496 | 0.341 |
| Attending radiologists | 0.865 (0.712-0.955) | 1.000 | 0.625 (0.245-0.915) | 0.500 | 0.931 (0.772-0.992) | 0.687 | 0.556 | 0.582 |
| Model 96 | 0.865 (0.712-0.955) | 0.875 (0.473-0.997) | 0.862 (0.683-0.961) | 0.737 | 0.649 |
The role of model 96 in assisting the performance of physicians across different professional categories in diagnosing PA-HSOS using CT imaging was evaluated. The results demonstrated significant improvements in diagnostic accuracy when RIMs and AGs were assisted by model 96. Accuracy improved from 0.541 to 0.757 for RIMs (P < 0.05), from 0.730 to 0.892 for AGs (P < 0.05), and showed an increasing, nonsignificant trend for RRs and APs. Sensitivity and specificity also improved, with notable enhancements in specificity for RIM (0.483 to 0.759, P < 0.01). Model 96 reduced diagnostic time across all professional categories (P < 0.05), with the most pronounced reduction for APs. These findings suggest that model 96 enhances both the efficiency and accuracy of HSOS diagnosis (Table 5).
| Item | Professional category | Doctor only | Doctor + model 96 | P value |
| Accuracy | RIM | 0.541 | 0.757 | 0.021a |
| AG | 0.730 | 0.892 | 0.031a | |
| RR | 0.676 | 0.811 | 0.063 | |
| AR | 0.865 | 0.973 | 0.125 | |
| Sensitivity | RIM | 0.750 | 0.875 | 1.000 |
| AG | 0.750 | 1.000 | ||
| RR | 0.875 | 0.875 | 1.000 | |
| AR | 0.625 | 0.875 | 0.500 | |
| Specificity | RIM | 0.483 | 0.759 | 0.008b |
| AG | 0.724 | 0.862 | 0.125 | |
| RR | 0.621 | 0.793 | 0.063 | |
| AR | 0.931 | 1.000 | ||
| Time (seconds) | RIM | 1807.00 ± 249.49 | 1561.33 ± 306.64 | 0.023a |
| AG | 1488.67 ± 462.48 | 924.67 ± 248.62 | 0.045a | |
| RR | 916.33 ± 187.22 | 832.00 ± 179.10 | 0.004b | |
| AR | 1071.00 ± 164.16 | 900.33+135.07 | 0.016a |
In recent years, AI has been widely used for diagnosing patients with nonalcoholic fatty liver disease or hepatocellular carcinoma, but there is no application for imaging diagnosis of DILI yet[20]. This study provides the first evidence that deep learning methods could be a useful tool to establish diagnostic models based on CT images for DILI. A multiscale convolutional module was used to extract and integrate features using receptive fields of different sizes, which was suitable for capturing the characteristic imaging patterns of PA-HSOS.
Among the deep learning models, model 96 (96-mm input size) showed an AUC of 0.944, accuracy of 0.865, sensitivity of 0.875, specificity of 0.862, positive predictive value of 0.636 and negative predictive value of 0.962, which demonstrated the superior diagnostic performance for PA-HSOS. The model can be used as a tool for RIMs and AGs as it can improve diagnostic accuracy. By combining medical history and laboratory tests, more accurate diagnosis might be made without further invasive examinations. The model significantly shortened diagnostic time. Considering that radiologists face a high demand for reading films every day, saving time is especially important.
The integration of model 96 into the diagnostic workflow demonstrated substantial benefits, particularly in improving diagnostic accuracy and reducing interpretation time. The observed enhancements in specificity for RIMs suggest that the model mitigates diagnostic uncertainties among less-experienced clinicians, while the reduced time across all groups indicates streamlined decision-making. The lack of statistical significance in some metrics (e.g., sensitivity for AP and RR) may reflect ceiling effects or limited sample size, warranting further investigation. Notably, the consistent reduction in diagnostic time without compromising accuracy underscores the potential of the model to optimize clinical workflows. These results align with prior studies highlighting the value of AI assistance in radiology[21], particularly for complex conditions like HSOS. Future research should explore long-term impacts, including effects on inter-reader variability and clinical outcomes, to validate the broader utility of the model.
In this study a transfer learning-based liver segmentation was utilized to extract ROIs containing only the liver region, so as to minimize the impact of surrounding organs in the classification tasks. From a qualitative perspective, we visualized the results in Supplementary Figure 3. The ablation study without using the auto-segmentation results in Supplementary Table 1 showed a decrease in AUC from 0.944 to 0.922, indicating that using areas other than the liver has a negative effect on the PA-HSOS classification task. Thus, automatic segmentation addresses this issue without introducing additional labeling costs. However, to further refine the segmentation results, manual correction may be considered following the automatic segmentation process.
The main innovations of the classification model lie in the multiscale convolutional modules and the anatomy-based ROI sampling strategy. The multiscale convolutional kernels help to capture information from different receptive fields, aiding in the detection of diffuse lesions and the classification. The ablation result of using only single scale convolution kernel highlighted the importance of using the multiscale convolutional kernels for extracting richer information for the model. The ablation experiments were conducted to illustrate the importance and necessity of sampling ROIs based on liver segment annotation. AUC decreased to 0.853 using the random sampling strategy. This suggested that using ROIs based on the liver segment was superior. On the one hand, it provided information relevant to the anatomical prior, improving the consistency between the deep learning prediction workflow and the real-world clinical diagnostic process, as well as the interpretability of the deep learning model. On the other hand, the selected ROI was more representative and could better reflect the characteristics of the PA-HSOS radiology data. Using a combination fivefold models results contributed to improved model performance.
In patient-level prediction, model 96 achieved the highest AUC of 0.944 compared with model 64 (AUC 0.853), model 128 (AUC 0.927) and model 160 (AUC 0.935) in the internal test cohort. By comparing the result of models with different input sizes, larger inputs may capture more lesion-related information, contributing to better performance. However, continuously increasing the input size may introduce redundant information into the model, potentially leading to a decline in performance. We performed paired DeLong tests on the ensemble prediction results of all models in both the internal and external test cohorts. All P values were > 0.05, indicating that models trained with different input sizes did not exhibit significant differences. The results demonstrate the effectiveness of our multiscale model, showing that the deep learning model was robust to variations in input size. We primarily selected model 64, which achieved the highest AUC on the internal test set, for subsequent analyses including comparisons with expert evaluations. We also visualized several samples that were consistently misclassified by all five cross-validation models, which were hard samples and were misclassified. This was potentially due to factors such as imaging features where diffuse parenchymal abnormalities mimicked HSOS or suboptimal image quality (Supplementary Figure 5). These misclassified hard samples also suggest that more training data will be needed in the future to improve the model performance.
The clinical translation of our model requires careful consideration of its positive predictive value, which is dependent on disease prevalence and would be lower in real-world settings than in our enriched cohort. Consequently, the model is positioned not as a screening tool, but as a decision-support aid for evaluating symptomatic patients. To mitigate the risk of false positives, its output must be integrated with clinical history and laboratory findings. Potential applications include serving as a triage alert for junior clinicians and a second opinion for specialists, thereby enhancing diagnostic efficiency without replacing clinical judgment.
This study had several limitations. First, the case-control design with two specific control diseases (Budd-Chiari syndrome and hepatitis B cirrhosis), while targeting the most critical differential diagnoses, may introduce spectrum bias and potentially overestimate performance. Future studies should include a broader spectrum of liver diseases (e.g., other forms of DILI, congestive hepatopathy) for validation. Second, although the sample size represented a substantial effort for this rare condition, it remained a constraint. Third, the retrospective design necessitates prospective validation. Fourth, despite external testing, performance should be further assessed on images from a wider array of scanner manufacturers and acquisition protocols. Finally, the performance of the model was contingent on the initial automated liver segmentation. Manual labels for evaluation or training may enhance the credibility of the results. The performance of our model indicates its potential for future clinical translation. With further development, such a tool could potentially be integrated into healthcare systems to provide decision-support data for patients exhibiting hepatic abnormalities. However, any clinical application would require rigorous multicenter validation. Ultimately, this tool should be viewed as an aid to clinical judgment rather than a replacement for physician expertise.
The deep-learning-based diagnostic model demonstrated promising performance for diagnosis of PA-HSOS using CT images and has the potential to assist clinicians in improving diagnostic accuracy and efficiency.
| 1. | Andrade RJ, Chalasani N, Björnsson ES, Suzuki A, Kullak-Ublick GA, Watkins PB, Devarbhavi H, Merz M, Lucena MI, Kaplowitz N, Aithal GP. Drug-induced liver injury. Nat Rev Dis Primers. 2019;5:58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 699] [Cited by in RCA: 559] [Article Influence: 79.9] [Reference Citation Analysis (3)] |
| 2. | Zhu L, Zhang CY, Li DP, Chen HB, Ma J, Gao H, Ye Y, Wang JY, Fu PP, Lin G. Tu-San-Qi (Gynura japonica): the culprit behind pyrrolizidine alkaloid-induced liver injury in China. Acta Pharmacol Sin. 2021;42:1212-1222. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 61] [Article Influence: 12.2] [Reference Citation Analysis (1)] |
| 3. | Yang XQ, Ye J, Li X, Li Q, Song YH. Pyrrolizidine alkaloids-induced hepatic sinusoidal obstruction syndrome: Pathogenesis, clinical manifestations, diagnosis, treatment, and outcomes. World J Gastroenterol. 2019;25:3753-3763. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 64] [Cited by in RCA: 65] [Article Influence: 9.3] [Reference Citation Analysis (4)] |
| 4. | Zhuge YZ, Wang Y, Zhang F, Zhu CK, Zhang W, Zhang M, He Q, Yang J, He J, Chen J, Zou XP. Clinical characteristics and treatment of pyrrolizidine alkaloid-related hepatic vein occlusive disease. Liver Int. 2018;38:1867-1874. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 60] [Article Influence: 7.5] [Reference Citation Analysis (1)] |
| 5. | Zhuge Y, Liu Y, Xie W, Zou X, Xu J, Wang J; Chinese Society of Gastroenterology Committee of Hepatobiliary Disease. Expert consensus on the clinical management of pyrrolizidine alkaloid-induced hepatic sinusoidal obstruction syndrome. J Gastroenterol Hepatol. 2019;34:634-642. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 86] [Article Influence: 12.3] [Reference Citation Analysis (1)] |
| 6. | Kan X, Ye J, Rong X, Lu Z, Li X, Wang Y, Yang L, Xu K, Song Y, Hou X. Diagnostic performance of Contrast-enhanced CT in Pyrrolizidine Alkaloids-induced Hepatic Sinusoidal Obstructive Syndrome. Sci Rep. 2016;6:37998. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 48] [Article Influence: 4.8] [Reference Citation Analysis (2)] |
| 7. | Ravaioli F, Colecchia A, Alemanni LV, Vestito A, Dajti E, Marasco G, Sessa M, Pession A, Bonifazi F, Festi D. Role of imaging techniques in liver veno-occlusive disease diagnosis: recent advances and literature review. Expert Rev Gastroenterol Hepatol. 2019;13:463-484. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 34] [Article Influence: 4.9] [Reference Citation Analysis (1)] |
| 8. | Park HJ, Park B, Lee SS. Radiomics and Deep Learning: Hepatic Applications. Korean J Radiol. 2020;21:387-401. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 139] [Cited by in RCA: 117] [Article Influence: 19.5] [Reference Citation Analysis (5)] |
| 9. | Xiang K, Jiang B, Shang D. The overview of the deep learning integrated into the medical imaging of liver: a review. Hepatol Int. 2021;15:868-880. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (1)] |
| 10. | Nam D, Chapiro J, Paradis V, Seraphin TP, Kather JN. Artificial intelligence in liver diseases: Improving diagnostics, prognostics and response prediction. JHEP Rep. 2022;4:100443. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 162] [Cited by in RCA: 120] [Article Influence: 30.0] [Reference Citation Analysis (1)] |
| 11. | Yang Q, Zhang S, Li Y. Deep Learning Algorithm Based on Molecular Fingerprint for Prediction of Drug-Induced Liver Injury. Toxicology. 2024;502:153736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 16] [Reference Citation Analysis (1)] |
| 12. | Dana J, Venkatasamy A, Saviano A, Lupberger J, Hoshida Y, Vilgrain V, Nahon P, Reinhold C, Gallix B, Baumert TF. Conventional and artificial intelligence-based imaging for biomarker discovery in chronic liver disease. Hepatol Int. 2022;16:509-522. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 33] [Article Influence: 8.3] [Reference Citation Analysis (1)] |
| 13. | Baek EB, Hwang JH, Park H, Lee BS, Son HY, Kim YB, Jun SY, Her J, Lee J, Cho JW. Artificial Intelligence-Assisted Image Analysis of Acetaminophen-Induced Acute Hepatic Injury in Sprague-Dawley Rats. Diagnostics (Basel). 2022;12:1478. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 8] [Reference Citation Analysis (1)] |
| 14. | Baek EB, Lee J, Hwang JH, Park H, Lee BS, Kim YB, Jun SY, Her J, Son HY, Cho JW. Application of multiple-finding segmentation utilizing Mask R-CNN-based deep learning in a rat model of drug-induced liver injury. Sci Rep. 2023;13:17555. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (1)] |
| 15. | Ronneberger O, Fischer P, Brox T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab N, Hornegger J, Wells W, Frangi A, editors. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. MICCAI 2015. Lecture Notes in Computer Science. Cham: Springer, 2015: 234-241. [RCA] [DOI] [Full Text] [Cited by in Crossref: 13000] [Cited by in RCA: 11480] [Article Influence: 1043.6] [Reference Citation Analysis (1)] |
| 16. | Cao H, Wang Y, Chen J, Jiang D, Zhang X, Tian Q, Wang M. Swin-Unet: Unet-Like Pure Transformer for Medical Image Segmentation. In: Karlinsky L, Michaeli T, Nishino K, editors. Computer Vision – ECCV 2022 Workshops. ECCV 2022. Lecture Notes in Computer Science. Cham: Springer, 2023: 205-218. [DOI] [Full Text] |
| 17. | European Association for the Study of the Liver. EASL Clinical Practice Guidelines: Vascular diseases of the liver. J Hepatol. 2016;64:179-202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 678] [Cited by in RCA: 577] [Article Influence: 57.7] [Reference Citation Analysis (2)] |
| 18. | European Association for the Study of the Liver. EASL Clinical Practice Guidelines for the management of patients with decompensated cirrhosis. J Hepatol. 2018;69:406-460. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2259] [Cited by in RCA: 2061] [Article Influence: 257.6] [Reference Citation Analysis (6)] |
| 19. | Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods. 2021;18:203-211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8060] [Cited by in RCA: 3932] [Article Influence: 786.4] [Reference Citation Analysis (3)] |
| 20. | Lu F, Meng Y, Song X, Li X, Liu Z, Gu C, Zheng X, Jing Y, Cai W, Pinyopornpanish K, Mancuso A, Romeiro FG, Méndez-Sánchez N, Qi X. Artificial Intelligence in Liver Diseases: Recent Advances. Adv Ther. 2024;41:967-990. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 14] [Article Influence: 7.0] [Reference Citation Analysis (3)] |
| 21. | Yamada A, Kamagata K, Hirata K, Ito R, Nakaura T, Ueda D, Fujita S, Fushimi Y, Fujima N, Matsui Y, Tatsugami F, Nozaki T, Fujioka T, Yanagawa M, Tsuboyama T, Kawamura M, Naganawa S. Clinical applications of artificial intelligence in liver imaging. Radiol Med. 2023;128:655-667. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 28] [Reference Citation Analysis (1)] |