Published online Apr 14, 2026. doi: 10.3748/wjg.v32.i14.116041
Revised: December 31, 2025
Accepted: February 6, 2026
Published online: April 14, 2026
Processing time: 153 Days and 7.7 Hours
The Liver Imaging Reporting and Data System (LI-RADS) is widely used for the diagnosis of hepatocellular carcinoma, but feature scoring by radiologists is subjective and time-consuming. An urgent need exists for an objective and efficient radiologist-supervised automated LI-RADS categorization system.
To develop an evidence-based radiologist-supervised automated LI-RADS grade 3 (LR-3), 4 (LR-4) and 5 (LR-5) categorization system (Evi-LIRADS) through quantitative feature characterization, following LI-RADS v2018.
This retrospective multicenter study (April 2012-November 2022) included untreated patients with suspected hepatocellular carcinoma undergoing gadoxetic acid-enhanced magnetic resonance imaging. Lesions from center 1 were partitioned into a development set (275 lesions used for five-fold cross-validation) and an internal testing set (62 lesions). Lesions from centers 2 (85 lesions) and 3 (104 lesions) constituted two external testing sets. Evi-LIRADS was designed by emulating the decision-making process of radiologists through a series of image processing algorithms, to recognize nonrim arterial phase hyper-enhancement, nonperipheral washout, and enhancing capsule, which provided detailed assessments of feature locations and patterns, improving the transparency of feature classification. Based on the three major image features and the automatically segmented lesion size, LI-RADS categories were assigned using LI-RADS v2018 algorithm. Feature classification was evaluated using area under the receiver operating characteristic curve. LI-RADS categorization was assessed by accuracy.
The internal dataset included 337 patients from center 1, while external datasets comprised 76 patients from center 2 and 97 patients from center 3. For feature classification, areas under the receiver operating characteristic curves were 0.975, 0.898, and 0.940 for arterial phase hyper-enhancement; 0.803, 0.824, and 0.850 for washout; 0.759, 0.800, and 0.784 for capsule across three datasets. Three-class LI-RADS categorization among LR-3, LR-4 and LR-5 achieved accuracies of 80.6%, 74.1%, and 77.9%, respectively, surpassing comparison methods (58.6%-69.6%). LI-RADS categorization between LR-3 and combined LR-4/LR-5 achieved 95.2%, 88.2%, and 90.4% accuracies for the three datasets, respectively. The visualization provided detailed feature locations and patterns. Evi-LIRADS saved an average of 21.1 seconds per patient (58.8% of the time) compared with radiologists, excluding radiologists’ quality control time.
Following LI-RADS guidelines and radiologists’ decision-making process, Evi-LIRADS was developed through quantitative feature characterization, demonstrating good accuracy, robust generalization, improved efficiency, enhanced clinical relevance, and improved transparency.
Core Tip: This study developed transparent classifiers for three of the major Liver Imaging Reporting and Data System (LI-RADS) features: Arterial phase hyper-enhancement, washout, and capsule. Then, LI-RADS categories were assigned in accordance with the LI-RADS v2018 guidelines. By following LI-RADS guidelines and emulating the decision-making process of radiologists, the model achieved radiologist-supervised automated LI-RADS categorization, through specialized feature characterization algorithms that provide explicit evidence for feature classification, thereby improving transparency for radiologists and patients. Categorization among LI-RADS grade 3, 4 and 5 achieved accuracies of 80.6%, 74.1%, and 77.9% for the internal testing set and two external testing sets, respectively.
- Citation: Xia XQ, Sheng RF, Zheng RC, Dai YX, Yang L, Chu YH, Zhang H, Wu XR, Shi NN, Wang CY, Zeng MS, Wang H. Evidence-based radiologist-supervised automated Liver Imaging Reporting and Data System categorization for the diagnosis of hepatocellular carcinoma. World J Gastroenterol 2026; 32(14): 116041
- URL: https://www.wjgnet.com/1007-9327/full/v32/i14/116041.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i14.116041
The incidence of liver cancer is increasing globally and is expected to exceed 35 million cases by 2050[1]. Hepatocellular carcinoma (HCC), the most common primary liver cancer, accounting for 75% to 90% of cases[2,3], has high morbidity and mortality[4]. Early detection and accurate diagnosis of HCC are essential for effective treatment. The Liver Imaging Reporting and Data System (LI-RADS), developed by the American College of Radiology, provides widely used guidelines for the diagnosis of HCC[5]. LI-RADS was first released in 2011 and updated in 2013, 2014, 2017, and 2018[5]. It classifies high-risk patients into categories LI-RADS grade 1 (LR-1) to 5 (LR-5) and M (LR-M). LI-RADS grade 3 (LR-3), 4 (LR-4), and 5 (LR-5) correspond to intermediate, high, and definite probabilities of HCC, respectively; however, distinguishing among these categories remains challenging[6]. Accurate identification of LR-3 to LR-5 lesions is crucial for guiding appropriate treatment strategies[7]. Several studies have addressed the classification of LR-3, LR-4, and LR-5 lesions[7-9]. Differentiation between LR-3 and combined LR-4/5 lesions is also important given that LR-3 lesions are generally managed less aggressively[10]. In LI-RADS v2018, the categorization of LR-3, LR-4 and LR-5 relies on nonrim arterial phase hyper-enhancement (APHE), nonperipheral washout, enhancing capsule, threshold growth and lesion size[5,11,12]. Threshold growth, defined by size increase, was not included in this study due to lack of follow-up data. This study focused on lesion size and three of the major features, APHE, washout, and capsule. However, the feature assessments by radiologists are subjective, heavily influenced by clinical experience with significant inter-observer variability (moderate agreement; intraclass correlation coefficient of 0.68[13]). Therefore, an urgent need exists for an objective and efficient radiologist-supervised automated LI-RADS LR-3, LR-4 and LR-5 categorization system.
Several LI-RADS categorization models have been proposed, employing radiomics-based models like random forest and support vector machine[14,15], and deep learning architectures such as DenseNet, AlexNet, and VGG16[10,15,16]. However, these models did not provide any evidence for the prediction of LI-RADS category and the decision-making processes lacked transparency. Moreover, they failed to follow clinical practice of using the three major LI-RADS features. Several other studies estimated these features using three deep learning models with the same architecture[7,17] or a single model through a multi-task learning framework[9]. However, the feature recognition process remained opaque, without any evidence. Lack of transparency made the model’s outputs hard to understand and trust, affecting the effectiveness and safety of treatment decisions. Additionally, the LI-RADS categorization accuracy remains low, ranging from 68.3% to 76.1%[7,9] for distinguishing LR-3, LR-4 and LR-5 in the internal testing set, which is insufficient for clinical practice. Moreover, most studies[9,10,14,15,17] did not use external validation datasets to verify the models’ robustness.
This study aimed to develop an evidence-based, radiologist-supervised automated LI-RADS categorization pipeline that provides transparent classifiers for three of the major LI-RADS features, APHE, washout, and capsule, using lesion segmentation, lesion subregion analysis for APHE and washout classification, and capsule enhancement for capsule classification, and that were used for automated LI-RADS categorization among LR-3, LR-4, and LR-5 in accordance with the LI-RADS v2018 guidelines.
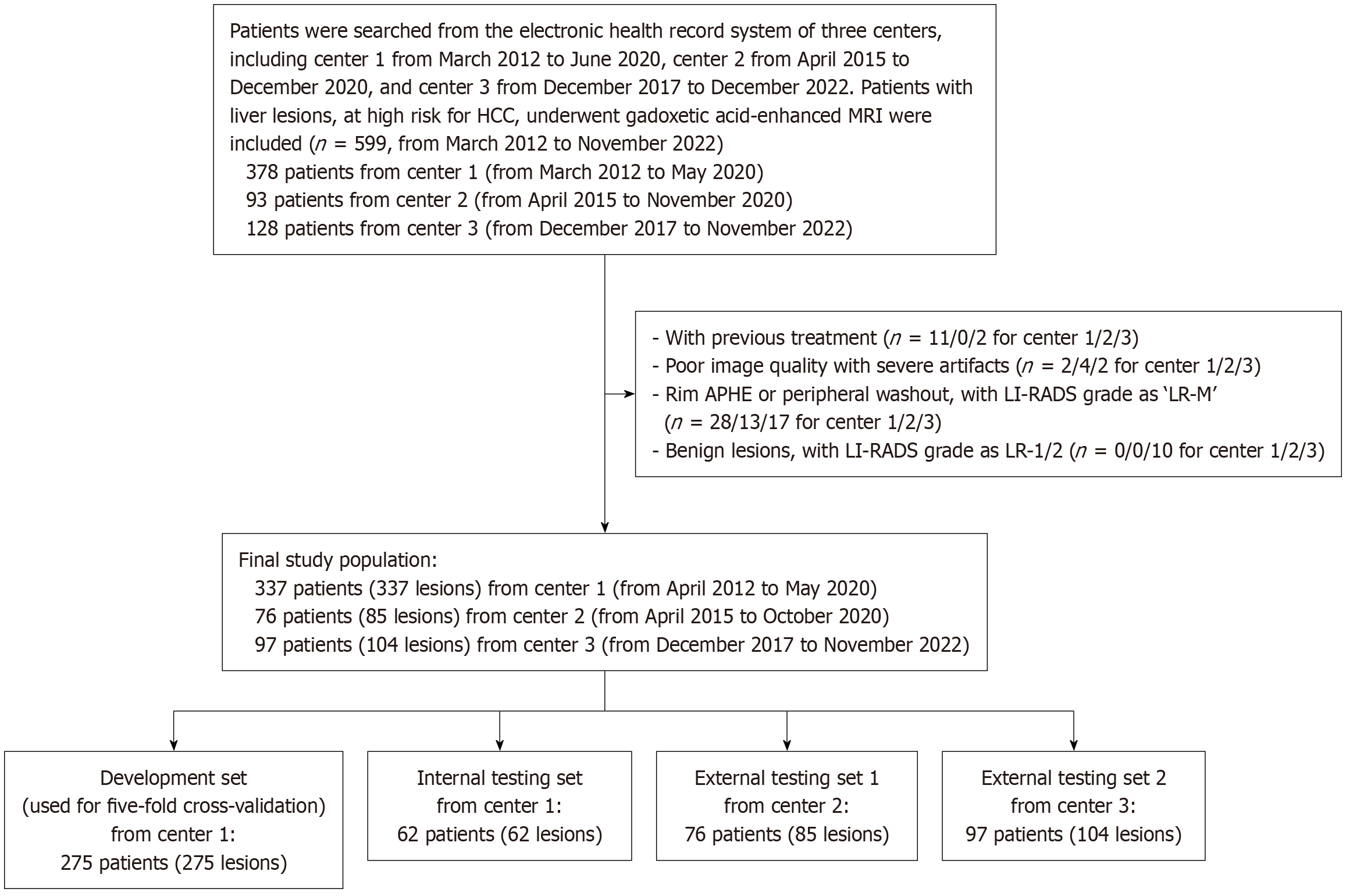
Patients at high risk for HCC with liver lesions who underwent gadoxetic acid (Primovist, Bayer HealthCare, Germany)-enhanced magnetic resonance imaging (MRI) were consecutively included. The inclusion and exclusion criteria are presented in Figure 1. Patients with previous treatment and poor image quality were excluded. This study focused on classifying LR-3, LR-4 and LR-5 lesions; thus, LR-1/2 and LR-M lesions were also excluded. Three datasets were included in this study: An internal dataset from center 1, and two external datasets from center 2 and 3 for external evaluation. Patients from center 1 were randomly divided into the training and validation set for algorithm development, and the testing set for internal evaluation.
All patients underwent gadoxetic acid-enhanced hepatic MRI. The contrast agent was injected at a dose of 0.025 mmol/kg and a flow rate of 2 mL/second for dynamic imaging. The arterial, portal venous, and transitional phases were scanned at 20-25 seconds, 60-70 seconds, and 120-180 seconds, respectively, followed by the hepatobiliary phase at 20 minutes. Detailed scanner models, scanning sequences, and parameters are provided in the Supplementary material and Supple
A senior radiologist with 12 years of liver MRI experience delineated lesions on transitional-phase axial images and measured the maximum diameters, serving as the reference standard for lesion size. These delineations were used to train and evaluate the segmentation model. If lesions exhibited iso-intensity on the transitional phase, the maximum diameters were measured on arterial phase images.
In terms of image features, for center 1, three radiologists with 6, 8, and 12 years of experience independently classified the three major LI-RADS features, APHE, washout, and capsule, while three radiologists with 8, 12, and 18 years of experience independently performed the feature classification for center 2 and 3. These radiologists were blinded to patient history, laboratory examination results, and pathologic results. Any disagreements were resolved through consensus review by the three radiologists to make final decisions, which served as the reference standard for image features. Reference standards for LI-RADS categories were established using the LI-RADS v2018 algorithm based on the three major image features classified by radiologists and the manually measured lesion size.
The proposed evidence-based radiologist-supervised automated LI-RADS LR-3, LR-4 and LR-5 categorization system (Evi-LIRADS) included registration, liver and lesion segmentation, lesion size calculation, feature characterization, and LI-RADS categorization following LI-RADS guidelines. The pipeline is radiologist-supervised end-to-end: Radiologists may verify/correct lesion masks as a quality-control step, whereas feature characterization and LI-RADS categorization are fully automated given a lesion mask.
The radiologist-supervised automated LI-RADS categorization involved liver and lesion segmentation (Figure 2A, left) using three-dimensional (3D) U-Net[18] and nnU-Net[19]. Before segmentation, the pre-contrast, arterial, and portal venous phases were registered to the transitional phase with advanced normalization tools[20] using SyNRA, which applies rigid, affine, and deformable transformations, with mutual information as the optimization criterion. For liver segmentation after registration, we leveraged a 3D U-Net architecture pre-trained on a dataset of gadoxetic acid-enhanced liver MRI[21]. The entire 3D image patch with a size of 64 × 224 × 256 was fed into the network for training. The total number of training epochs was set to 200, and the model with the best validation performance during training was retained for prediction on the testing sets. For the nnU-Net model used for lesion segmentation, the 3D full resolution U-Net configuration was applied. The nnU-Net was trained on registered images from all four phases together with manual masks. Five-fold cross-validation was performed in the development (training/validation) set, with 1000 epochs per fold. Other parameters were set as in the original study. Lesion segmentation performance was evaluated using mean Dice similarity coefficients. After lesion segmentation, radiologists reviewed and modified the labels as needed, which is limited to quality control of the mask, and maximum lesion diameters were automatically calculated. The continuous diameter values were mapped into three intervals as specified in LI-RADS v2018, including < 10 mm, 10-19 mm, and ≥ 20 mm. Accuracy and quadratic weighted Cohen’s kappa coefficient were used to evaluate the consistency between the calculated lesion diameter and the manually measured diameter through the classification of the three diameter intervals.
Before feature classification, we applied a fully automated preprocessing pipeline that included removal of fragmented, extremely small regions (fewer than 3 pixels), hole filling in lesion masks, boundary smoothing via dilation and erosion, and hole filling in the liver mask. Finally, the lesion region of interest was cropped on a per-slice basis (Figure 2A, right; Supplementary material).
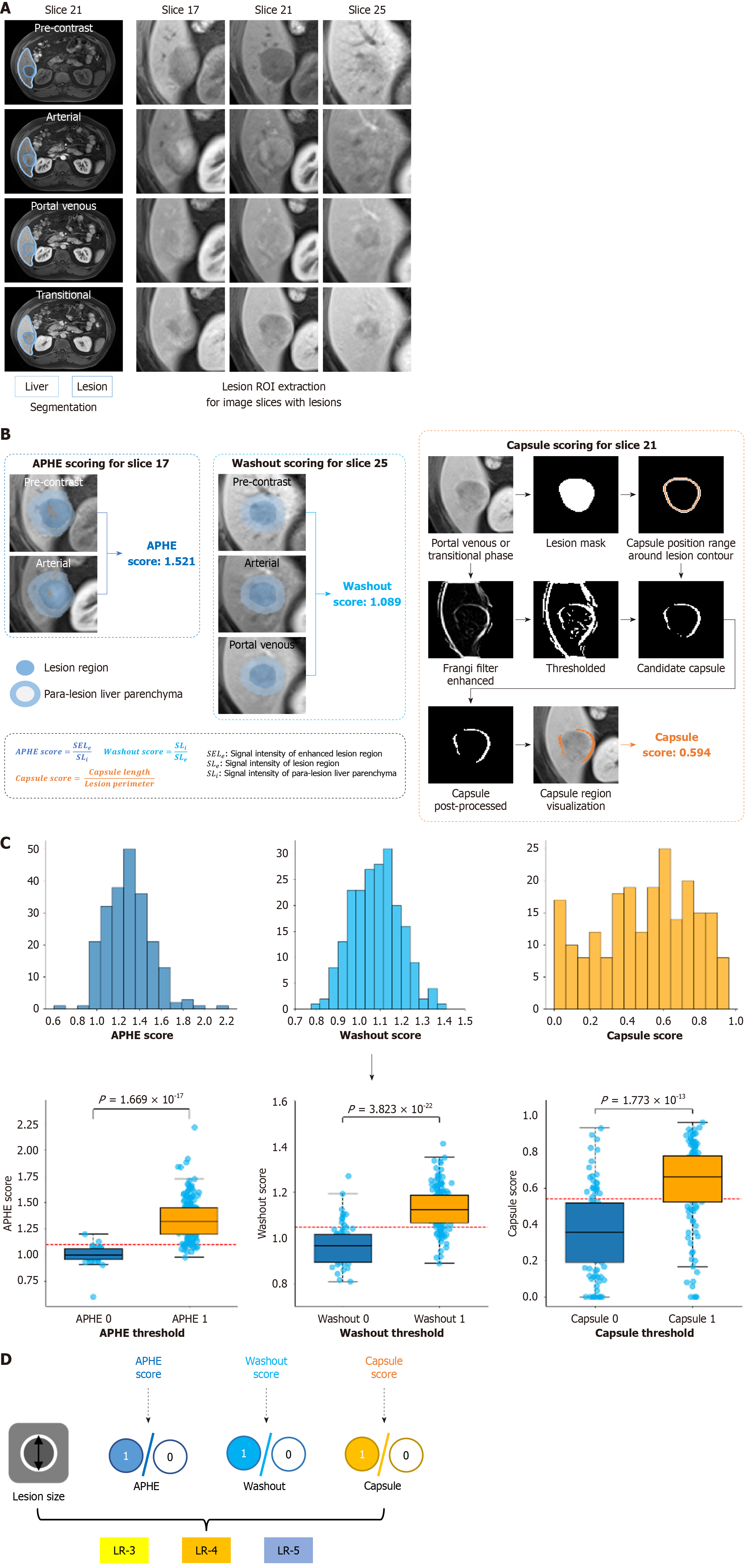
Three classifiers were developed to classify the three major LI-RADS features. The APHE classifier calculated the ratio of the 75th percentile signal intensity in the enhanced lesion region (extracted using Otsu’s method[22]; Figure 2B) to the mean signal intensity of para-lesion liver parenchyma, aligning better with LI-RADS guidelines when APHE just has enhancement in part of the lesion. The washout classifier used the ratio of the mean signal intensity of the lesion or dark region within the lesion (if there was strong enhancement in the arterial phase) to the mean signal intensity of para-lesion liver parenchyma (Figure 2B). The capsule classifier applied the Frangi filter[23] to enhance and extract capsule regions, followed by post-processing to remove small points and confirm capsule regions based on signal intensity. The capsule score was the ratio of capsule length to lesion perimeter (Figure 2B). For all features, higher scores indicated a greater probability of feature presence. Each feature was then classified as 0 (absent) or 1 (present) based on feature score thresholds (Figure 2C and D). More details on feature classification are provided in the Supplementary material.
LI-RADS categorization among LR-3, LR-4 and LR-5 was performed following LI-RADS v2018 guidelines by assessing the three major image features (APHE, washout and capsule) combined with lesion size (Figure 2D). Lesions without any major features were classified as LR-3. Without APHE, lesions with only washout or capsule were LR-3 if < 20 mm and LR-4 if ≥ 20 mm; those with both washout and capsule were LR-4 regardless of size. With APHE, absence of washout/capsule led to LR-3 (< 20 mm) or LR-4 (≥ 20 mm). APHE with only washout or only capsule was classified as LR-4 (< 10 mm), LR-5 (≥ 20 mm), and for 10-19 mm, LR-5 if washout-only or LR-4 if capsule-only. Lesions with all three features were LR-4 (< 10 mm) or LR-5 (≥ 10 mm). This study also assessed Evi-LIRADS’s ability to differentiate intermediate LR-3 lesions from more-likely malignant LR-4/LR-5 lesions.
To verify its effectiveness, Evi-LIRADS was compared with three existing LI-RADS categorization methods: A radiomics-based random forest classifier (Radiomics)[14], and deep learning methods AlexNet[10] and VGG16[16], learned by the same data as Evi-LIRADS used. For fair comparisons, radiomics and deep-learning baselines used the same registration results and radiologist-verified lesion masks. Lesion-centered region of interests were generated by bounding-box cropping with a fixed margin of 20 pixels (thus including perilesional liver parenchyma), with the same intensity normalization and multiphase input configuration. The deep learning baselines used multi-phase inputs (arterial, portal venous, and transitional phases) and were trained using the same train/validation/test splits as our study. Model selection was performed on the validation set (e.g., selecting the best checkpoint and using early stopping based on validation accuracy), with conventional training settings (optimizer, learning-rate schedule, and data aug
Continuous variables were reported as mean ± SD, and categorical variables as n (%). Inter-observer agreement was assessed using Fleiss kappa coefficient[24] for feature classification and quadratic weighted Fleiss kappa coefficient for LI-RADS categorization. A sample size of 73 patients was calculated using PASS (version 2021; NCSS, UT, United States) based on reported areas under the receiver operating characteristic curve (AUCs) for APHE (0.941), washout (0.859), and capsule (0.712), ensuring 90% power at a 0.05 significance level. Five-fold cross-validation was repeated five times, with means and 95% confidence intervals calculated. Feature classification and LI-RADS categorization between LR-3 and LR-4/5 was evaluated using AUC, sensitivity, specificity, accuracy, and F1 score. LI-RADS categorization among LR-3, LR-4 and LR-5 was assessed using overall accuracy and quadratic weighted Cohen’s kappa coefficient.
Accuracies were compared between Evi-LIRADS and the three LI-RADS categorization comparison methods using the paired sample t test. A P value < 0.05 was considered statistically significant. To examine whether inter-center heterogeneity in cirrhosis prevalence introduced within-center clustering of validation errors, we performed mixed-effects analyses using R 4.5.2. To evaluate model robustness under balanced subgroup representation across centers and etiologies, we performed a center × hepatitis B virus (HBV)-stratified resampling analysis using R 4.5.2. All other analyses were conducted using Python 3.8.11 with relevant libraries. Fleiss kappa ranges for various inter-observer agreement levels, details of mixed-effects analyses, and details of center × etiology-stratified resampling analysis are provided in the Supplementary material.
The flowchart of the study population is depicted in Figure 1. The internal dataset comprised 337 patients from center 1 (mean age, 53.9 ± 11.3 years; 286 males), the external datasets comprised 76 patients from center 2 (mean age, 59.5 ± 12.1 years; 59 males), and 97 patients from center 3 (mean age, 55.6 ± 11.5 years; 84 males). Detailed patient demographics are depicted in Table 1. The number of positive and negative samples for the three major features and sample numbers of LI-RADS categories (LR-3, LR-4, and LR-5) based on the consensus reads by the radiologists are presented in Supple
| Characteristics | Datasets from center 1 (n = 337) | Training and validation set from center 1 (n = 275) | Internal testing set from center 1 (n = 62) | External testing set 1 from center 2 (n = 76) | External testing set 2 from center 3 (n = 97) |
| Age (years) | 53.9 ± 11.3 (24-81) | 53.7 ± 11.1 (26-81) | 54.7 ± 12.2 (24-81) | 59.5 ± 12.1 (30-83) | 55.6 ± 11.5 (28-81) |
| Sex | |||||
| Male | 286 (84.9) | 229 (83.3) | 57 (91.9) | 59 (77.6) | 84 (86.6) |
| Female | 51 (15.1) | 46 (16.7) | 5 (8.1) | 17 (22.4) | 13 (13.4) |
| Etiology | |||||
| Hepatitis B | 265 (78.6) | 218 (79.3) | 47 (75.8) | 56 (73.7) | 86 (88.7) |
| Other | 72 (21.4) | 57 (20.7) | 15 (24.2) | 20 (26.3) | 11 (11.3) |
| With cirrhosis | 297 (88.1) | 246 (89.5) | 51 (82.3) | 31 (40.8) | 65 (67.0) |
| PLT (× 109/L) | 145.2 ± 122.9 | 146.9 ± 133.7 | 137.6 ± 57.8 | 143.8 ± 70.6 | 150.8 ± 75.7 |
| ALT (U/L) | 39.9 ± 43.6 | 39.8 ± 41.8 | 40.4 ± 51.3 | 27.6 ± 17.8 | 73.2 ± 206.5 |
| AST (U/L) | 36.2 ± 38.9 | 36.4 ± 39.4 | 35.5 ± 36.8 | 35.1 ± 18.8 | 75.4 ± 260.9 |
| ALB (g/L) | 43.0 ± 4.5 | 43.2 ± 4.3 | 41.7 ± 5.2 | 40.4 ± 5.3 | 39.5 ± 5.3 |
| TBIL (μmol/L) | 14.7 ± 8.0 | 15.0 ± 8.2 | 13.3 ± 6.8 | 19.0 ± 12.2 | 26.3 ± 41.7 |
| AKP (U/L) | 80.9 ± 29.5 | 82.3 ± 30.7 | 74.0 ± 22.2 | 113.0 ± 90.7 | 124.0 ± 60.8 |
| GGT (U/L) | 75.4 ± 110.9 | 79.5 ± 119.0 | 55.3 ± 52.8 | 104.9 ± 118.8 | 84.8 ± 81.6 |
Results of inter-observer agreement for feature classification and LI-RADS categorization are presented in Supplementary Table 3. For feature classification, the consistency of capsule from center 1 showed substantial agreement; other analyses showed almost perfect agreement. For LI-RADS categories, all the quadratic weighted Fleiss kappa coefficients for center 1, 2, and 3 indicated almost perfect agreement.
The mean Dice similarity coefficients for lesion segmentation were 0.892 ± 0.044 for the training and validation sets from center 1, 0.731 ± 0.168 for the internal testing set from center 1, 0.745 ± 0.235 for the external testing set 1 from center 2, and 0.728 ± 0.241 for the external testing set 2 from center 3. Representative segmentation results are illustrated in Supple
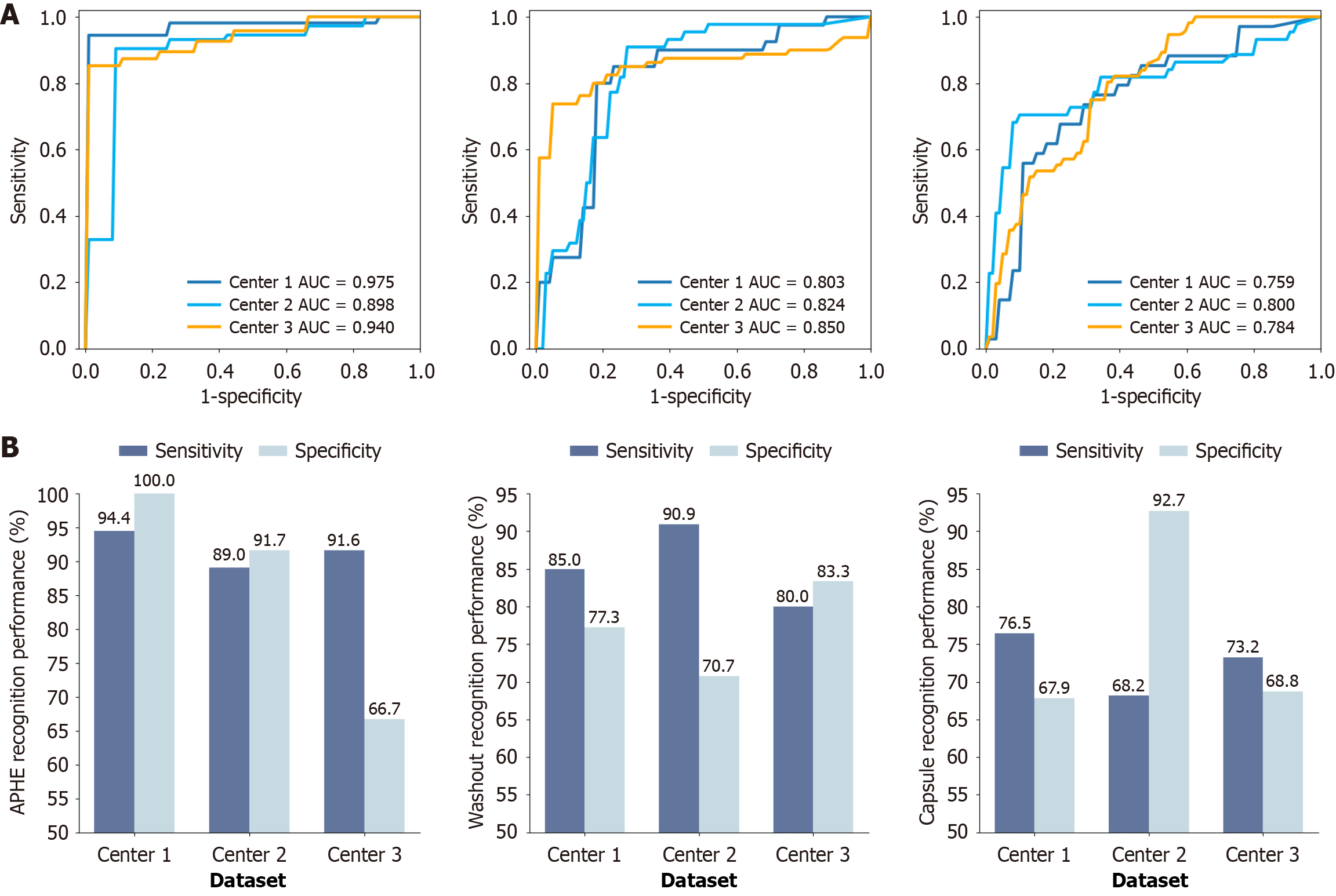
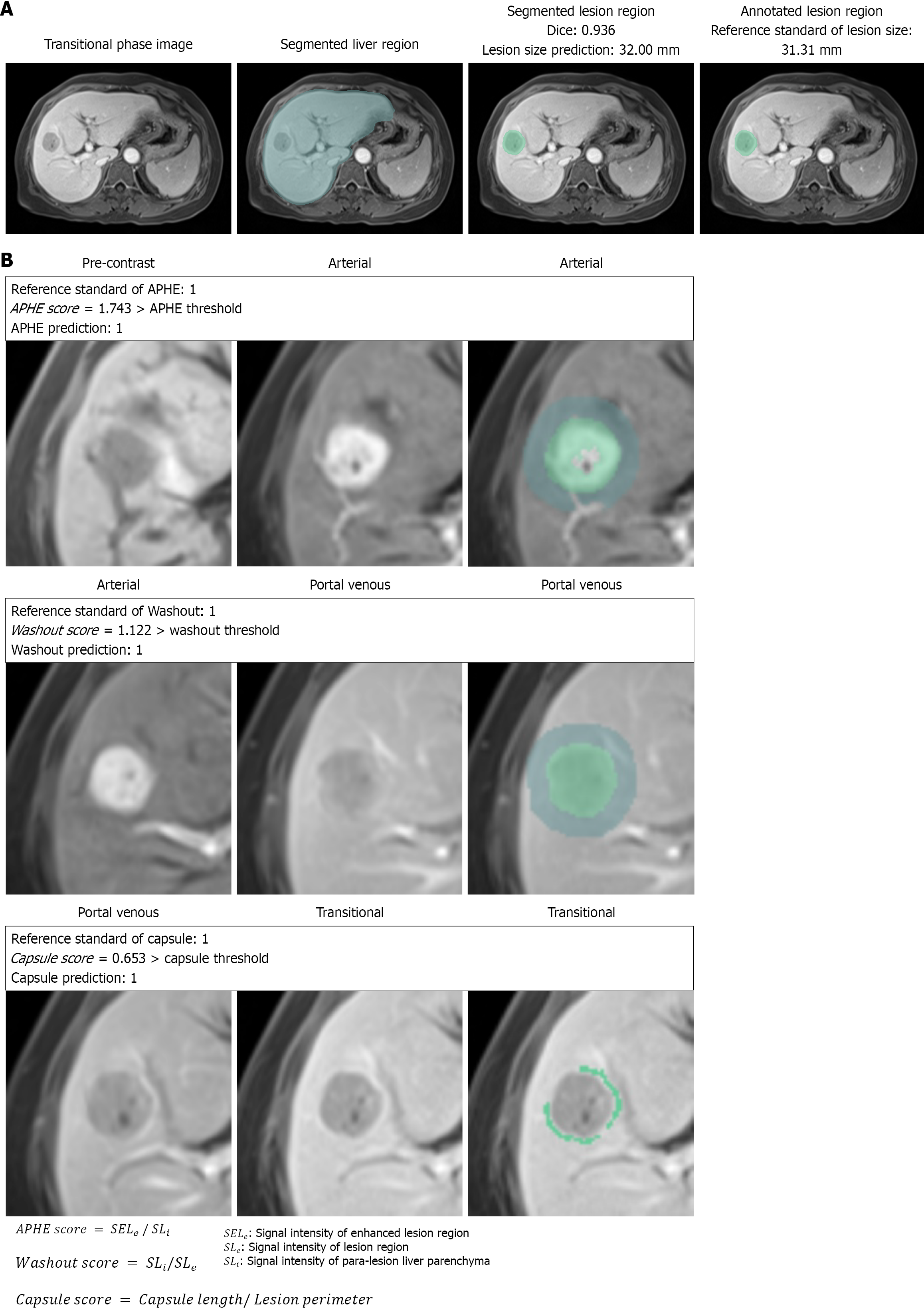
The classification results of the three major features were analyzed for the three datasets. APHE yielded outstanding performance with AUCs of 0.898-0.975 and sensitivities of 89.0%-94.4%; washout performed well with AUCs of 0.803-0.850 and sensitivities of 80.0%-90.9%; and capsule achieved moderate performance with AUCs of 0.759-0.800 and sensitivities of 68.2%-76.5%. The specificities, accuracies, and F1 scores of feature classification are presented in Table 2. The three feature classifiers demonstrated stable performance on the internal and two external testing sets, exhibiting robust generalization. Receiver operating characteristic curves are presented in Figure 3A, and the sensitivity and specificity are depicted in Figure 3B. Visualization results for liver/Lesion segmentation and feature characterization for a representative case are displayed in Figure 4, showing feature locations and detail patterns which are the evidence for the judgment of the three major features and final LI-RADS category. More cases for APHE, washout, and capsule characterization are displayed in Supplementary Figure 3. Feature locations, patterns, and quantitative feature score displayed in Figure 4 and Supplementary Figure 3 demonstrated the effectiveness of the algorithms, and improved the transparency of the algorithms, which can help radiologists understand how the model makes the decision.
| Data sets and metrics | APHE | Washout | Capsule |
| Center 1 | |||
| AUC | 0.975 | 0.803 | 0.759 |
| Sensitivity, % | 94.4 | 85.0 | 76.5 |
| Specificity, % | 100.0 | 77.3 | 67.9 |
| Accuracy, % | 95.2 | 82.3 | 72.6 |
| F1 score | 0.971 | 0.861 | 0.754 |
| Center 2 | |||
| AUC | 0.898 | 0.824 | 0.800 |
| Sensitivity, % | 89.0 | 90.9 | 68.2 |
| Specificity, % | 91.7 | 70.7 | 92.7 |
| Accuracy, % | 89.4 | 81.2 | 80.0 |
| F1 score | 0.935 | 0.833 | 0.779 |
| Center 3 | |||
| AUC | 0.940 | 0.850 | 0.784 |
| Sensitivity, % | 91.6 | 80.0 | 73.2 |
| Specificity, % | 66.7 | 83.3 | 68.8 |
| Accuracy, % | 89.4 | 80.8 | 71.2 |
| F1 score | 0.941 | 0.865 | 0.732 |
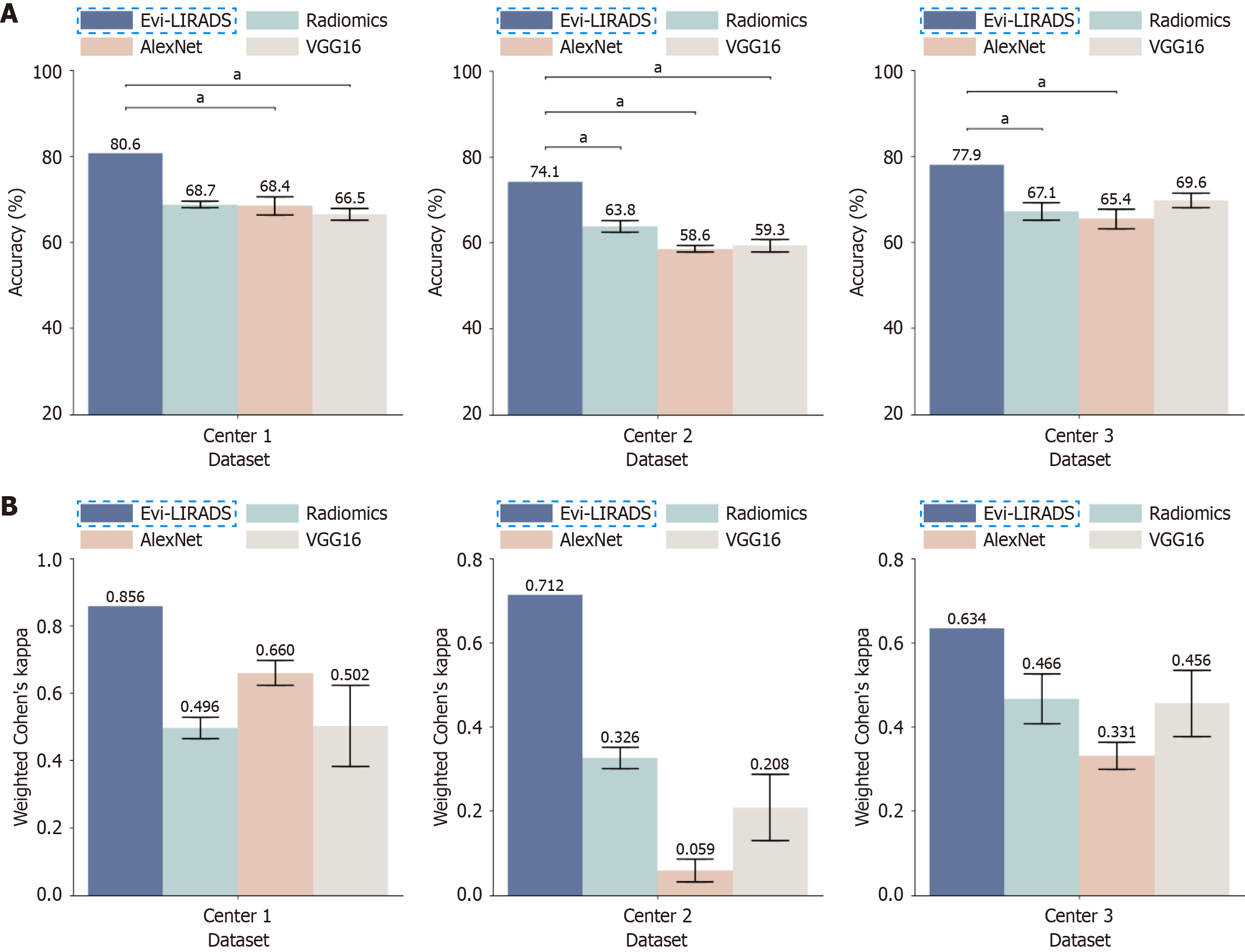
The overall accuracy of LI-RADS categorization among LR-3, LR-4 and LR-5 using Evi-LIRADS was 80.6%, 74.1%, and 77.9% on the internal testing set, external testing set 1, and external testing set 2, respectively. The quadratic weighted Cohen’s kappa coefficient was 0.856, 0.712, and 0.634, respectively. Evi-LIRADS was compared with Radiomics, AlexNet, and VGG16 and achieved the highest accuracy and quadratic weighted Cohen’s kappa coefficient across all datasets (Figure 5). Paired-sample t tests showed that Evi-LIRADS (accuracy = 80.6%) achieved significantly higher accuracy than AlexNet (68.4%) and VGG16 (66.5%) in center 1 (P = 0.045 and P = 0.024, respectively). In center 2, Evi-LIRADS outperformed all three comparison methods (accuracies = 63.8%, 58.6%, and 59.3%) with statistical significance (P = 0.049, P = 0.013, and P = 0.040, respectively). In center 3, Evi-LIRADS also performed significantly better than Radiomics (67.1%) and AlexNet (65.4%) (P = 0.033 and P = 0.016, respectively).
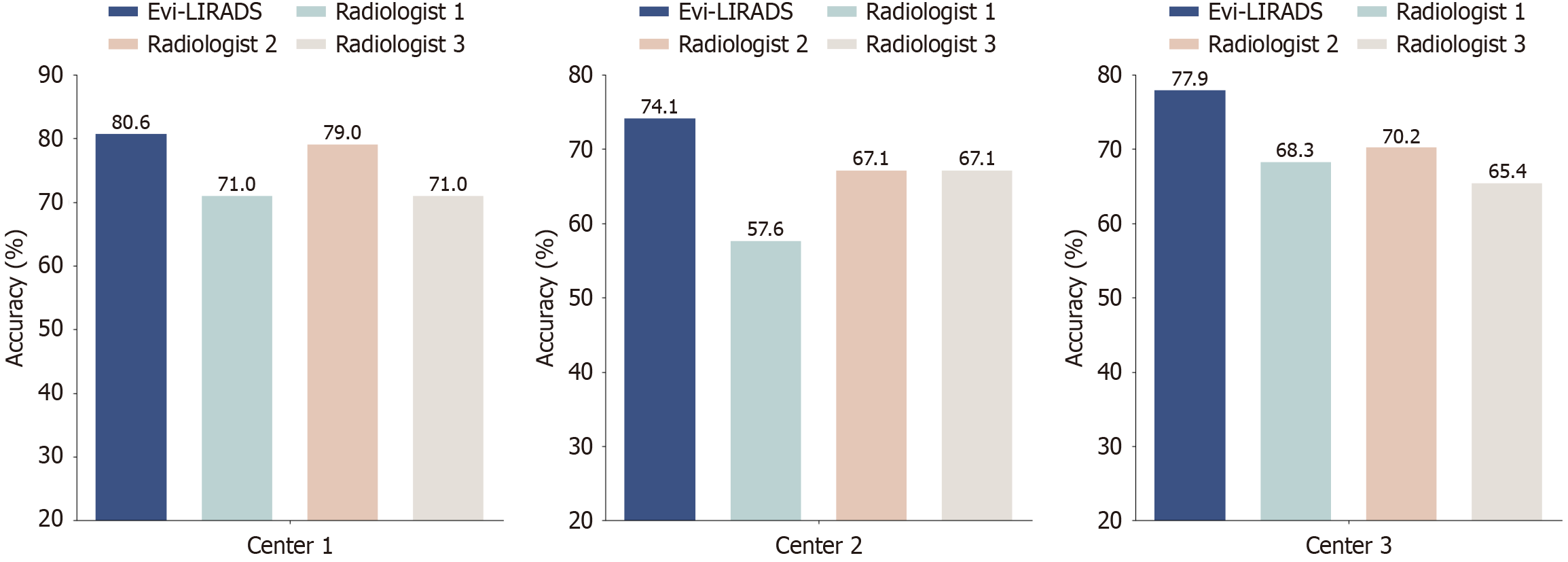
LI-RADS category classification between LR-3 and combined LR-4/LR-5 achieved AUCs of 0.826-0.918, sensitivities of 92.1%-97.9%, accuracies of 88.2%-95.2%, and F1 scores of 0.923-0.969 for the three datasets (Table 3). Compared with three junior radiologists with two years of liver MRI experience, the overall accuracies of LI-RADS categorization by Evi-LIRADS were comparable or better than that of the three junior radiologists for all the three datasets (Figure 6). The detail results of mixed-effects analyses and center × HBV-stratified resampling analyses are provided in the Supplementary material. The results of mixed-effects analyses suggest that the lower cirrhosis rate in center 2 did not induce meaningful clustering or systematic center effects that would bias validation performance or compromise generalizability. The center × HBV-stratified resampling analyses support robust ordinal grading performance under subgroup-balanced evaluation, without evidence that HBV materially increases the magnitude of grading errors after accounting for center and the true LI-RADS grade.
| Data sets and metrics | AUC | Sensitivity, % | Specificity, % | Accuracy, % | F1 score |
| Center 1 | 0.918 | 97.9 | 85.7 | 95.2 | 0.969 |
| Center 2 | 0.826 | 93.8 | 71.4 | 88.2 | 0.923 |
| Center 3 | 0.861 | 92.1 | 80.0 | 90.4 | 0.943 |
The radiologists took an average of 35.80 seconds per patient for feature classification, lesion measurement, and LI-RADS categorization. In contrast, our radiologist-supervised automated method required only 14.74 seconds, saving approximately 21.06 seconds (58.8%) per patient. The running time comparisons between Evi-LIRADS and radiologists are detailed in Table 4. The reported time saving reflects automated lesion diameter measurement, feature characterization and LI-RADS categorization given a lesion mask, and does not include optional radiologist time for mask review/editing. Radiologists reviewed (and corrected when needed) the segmentation masks as a quality-control step; this time was not included in the reported runtime.
| Data sets | Time of Evi-LIRADS (seconds per patient) | Time of radiologists (seconds per patient) |
| Center 1 | 6.6 ± 3.4 | 35.4 ± 15.1 |
| Center 2 | 18.8 ± 22.6 | 31.8 ± 7.6 |
| Center 3 | 18.8 ± 13.1 | 40.2 ± 16.1 |
| Average for three centers | 14.7 | 35.8 |
This study developed an evidence-based, transparent system for LI-RADS categorization among LR-3, LR-4, and LR-5 and demonstrated favorable performance across internal and external testing. Across external validations, the system achieved accuracies of 88.2%-95.2% for distinguishing LR-3 from LR-4/5 and 74.1%-80.6% for LR-3/4/5 categorization, supporting stable performance across centers under a radiologist-supervised workflow. Radiologist supervision was limited to quality control of the lesion mask when needed; feature characterization and categorization were fully auto
By following LI-RADS guidelines and emulating the decision-making process of radiologists, our system enabled radiologist-supervised automated LI-RADS categorization using transparent, feature-specific algorithms that provide explicit, phase-linked evidence for each major feature. Prior work has rarely implemented evidence-based algorithms for LI-RADS feature recognition. One study[8] developed evidence-based washout and capsule recognition algorithms but was limited to hyper-enhanced lesions and did not address APHE recognition; washout was assessed by comparing the intensity of the entire lesion with the liver parenchyma. Compared with that approach, we improved APHE assessment by applying Otsu’s method[22] to extract the enhancing subregion within a lesion, better reflecting partial enhancement. We further refined capsule assessment by confirming candidate capsule regions via signal intensity criteria rather than relying solely on Frangi filter[23]-based rim enhancement, and we introduced a capsule score based on capsule length, which is more clinically meaningful.
Prior studies have explored both computer-aided[8] and deep-learning[7,10,16,17,25] approaches for LI-RADS-related assessment, including differentiation of LR-3 from LR-4/5 lesions on multiphase MRI[10], LI-RADS v2014 category classification in a pilot setting[16], and major-feature classification using subtraction MRI images[17]. However, these studies vary in task formulation, LI-RADS version, and the extent to which phase-specific, guideline-linked evidence is provided. For example, a relatively transparent convolutional neural network inferred feature attribution from activation patterns[25], but the highlighted regions reflected combined multiphase information rather than explicit, phase-specific evidence aligned with LI-RADS definitions. In contrast, our framework explicitly links each feature decision to its corresponding phase, aligning more closely with LI-RADS definitions and routine radiologic interpretation.
Compared with existing studies[7,10,16,17], our study demonstrated superior performance. Independent external validation remains limited in the published literature, and reported results vary across study designs. Among deep learning-based studies reporting external testing, one study reported an overall accuracy of 60.4% internally and 47.7% on an external MRI dataset (LI-RADS v2014; multi-class categorization)[16], and another study reported LR-3/4/5 grading accuracies of 68.3% (internal) and 66.2% (external) in an expert-guided step-by-step framework[7]. Although task formulations and LI-RADS versions differ across studies, these findings highlight the ongoing challenge of robust external generalization. In our study, accuracies on two external test sets (74.1% and 77.9%) were higher than those reported in the above externally validated studies[7,16], while additionally providing phase-specific, auditable evidence for each major feature. Class imbalance is common in LI-RADS datasets and can bias end-to-end deep learning training toward majority classes. In our experiments, the deep learning baselines showed increased confusion for LR-3/LR-4 cases (Supplementary Figure 4). In contrast, our rule-based method applies consistent guideline-defined logic irrespective of class frequency, contributing to more stable categorization across categories.
We acknowledge that purely rule-based approaches have limitations for challenging features such as capsule (e.g., thin/discontinuous rims or boundary-adjacent lesions). A practical next step is a hybrid strategy: Rule-based logic for strictly defined features (e.g., APHE and washout) combined with deep learning-based refinement/enhancement (e.g., multi-scale attention and edge-aware enhancement) for challenging features (e.g., capsule).
This study has several limitations. First, following LI-RADS v2018, washout criteria differ between gadoxetic acid and other gadolinium-based contrast agents (GBCAs). For GBCAs, washout is assessed in the portal venous and delayed phases, whereas for gadoxetic acid it is evaluated only in the portal venous phase. Our algorithms are tailored for gadoxetic acid-enhanced MRI, and were not validated with other GBCAs; future studies will include other contrast agents. Second, this study currently focuses on learning and evaluating LI-RADS major features using pre-contrast, arterial, portal venous, and transitional phases of gadoxetic acid-enhanced MRI. Building on these results, future work will incorporate the hepatobiliary phase of gadoxetic acid-enhanced MRI and additional sequences (T2-weighted imaging and diffusion-weighted imaging) to analyze ancillary features[12,26] and further improve performance. Third, although the capsule feature extraction algorithm shows promise, its robustness remains limited in lesions with irregular tumor margins, thin or discontinuous capsules, and lesions located adjacent to vessels or the liver surface, which may partly explain its relatively modest AUC (0.759-0.800). The main sources of capsule misclassification include: (1) Ambiguous or fragmented capsule appearance, particularly when the enhancing rim is thin, incomplete, or mixed with perilesional enhancement; (2) Irregular lesion boundaries and partial-volume effects that blur the capsule-parenchyma interface; (3) Heterogeneous enhancement across phases and phase-timing variability that attenuate capsule conspicuity; and (4) Confounding structures (e.g., adjacent vessels, fibrotic bands, or peritumoral perfusion alterations) that can mimic capsule-like rim enhancement. To improve capsule recognition, we propose several optimization strategies: (1) Incor
In conclusion, we developed Evi-LIRADS, a LI-RADS LR-3, LR-4 and LR-5 categorization system that resembled clinical radiologists’ decision-making process to analyze the three major LI-RADS features in detail and followed the LI-RADS v2018 algorithm to assess LI-RADS category. The system demonstrated good accuracy, robust generalization, and improved efficiency for the diagnosis of HCC. Our evidence-based feature characterization algorithms provide clear evidence for the presence of each major feature, enhancing their clinical relevance and transparency. Evi-LIRADS shows strong potential for clinical application.
We wish to thank Ye Yao, PhD (Associate Professor in the Department of Biostatistics, School of Public Health, Fudan University), for his help in reviewing the statistical methods of this study.
| 1. | Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, Jemal A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229-263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16785] [Cited by in RCA: 15073] [Article Influence: 7536.5] [Reference Citation Analysis (23)] |
| 2. | Llovet JM, Kelley RK, Villanueva A, Singal AG, Pikarsky E, Roayaie S, Lencioni R, Koike K, Zucman-Rossi J, Finn RS. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7:6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1323] [Reference Citation Analysis (0)] |
| 3. | Petrick JL, Florio AA, Znaor A, Ruggieri D, Laversanne M, Alvarez CS, Ferlay J, Valery PC, Bray F, McGlynn KA. International trends in hepatocellular carcinoma incidence, 1978-2012. Int J Cancer. 2020;147:317-330. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 484] [Cited by in RCA: 447] [Article Influence: 74.5] [Reference Citation Analysis (4)] |
| 4. | Golabi P, Fazel S, Otgonsuren M, Sayiner M, Locklear CT, Younossi ZM. Mortality assessment of patients with hepatocellular carcinoma according to underlying disease and treatment modalities. Medicine (Baltimore). 2017;96:e5904. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 214] [Cited by in RCA: 197] [Article Influence: 21.9] [Reference Citation Analysis (3)] |
| 5. | Chernyak V, Fowler KJ, Kamaya A, Kielar AZ, Elsayes KM, Bashir MR, Kono Y, Do RK, Mitchell DG, Singal AG, Tang A, Sirlin CB. Liver Imaging Reporting and Data System (LI-RADS) Version 2018: Imaging of Hepatocellular Carcinoma in At-Risk Patients. Radiology. 2018;289:816-830. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 990] [Cited by in RCA: 904] [Article Influence: 113.0] [Reference Citation Analysis (4)] |
| 6. | Arribas Anta J, Moreno-Vedia J, García López J, Rios-Vives MA, Munuera J, Rodríguez-Comas J. Artificial intelligence for detection and characterization of focal hepatic lesions: a review. Abdom Radiol (NY). 2025;50:1564-1583. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 7. | Sheng R, Huang J, Zhang W, Jin K, Yang L, Chong H, Fan J, Zhou J, Wu D, Zeng M. A Semi-Automatic Step-by-Step Expert-Guided LI-RADS Grading System Based on Gadoxetic Acid-Enhanced MRI. J Hepatocell Carcinoma. 2021;8:671-683. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 7] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 8. | Kim Y, Furlan A, Borhani AA, Bae KT. Computer-aided diagnosis program for classifying the risk of hepatocellular carcinoma on MR images following liver imaging reporting and data system (LI-RADS). J Magn Reson Imaging. 2018;47:710-722. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 11] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 9. | Wang K, Liu Y, Chen H, Yu W, Zhou J, Wang X. Fully automating LI-RADS on MRI with deep learning-guided lesion segmentation, feature characterization, and score inference. Front Oncol. 2023;13:1153241. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 7] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 10. | Wu Y, White GM, Cornelius T, Gowdar I, Ansari MH, Supanich MP, Deng J. Deep learning LI-RADS grading system based on contrast enhanced multiphase MRI for differentiation between LR-3 and LR-4/LR-5 liver tumors. Ann Transl Med. 2020;8:701. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 40] [Article Influence: 6.7] [Reference Citation Analysis (5)] |
| 11. | Tang A, Bashir MR, Corwin MT, Cruite I, Dietrich CF, Do RKG, Ehman EC, Fowler KJ, Hussain HK, Jha RC, Karam AR, Mamidipalli A, Marks RM, Mitchell DG, Morgan TA, Ohliger MA, Shah A, Vu KN, Sirlin CB; LI-RADS Evidence Working Group. Evidence Supporting LI-RADS Major Features for CT- and MR Imaging-based Diagnosis of Hepatocellular Carcinoma: A Systematic Review. Radiology. 2018;286:29-48. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 261] [Cited by in RCA: 238] [Article Influence: 29.8] [Reference Citation Analysis (4)] |
| 12. | Cerny M, Bergeron C, Billiard JS, Murphy-Lavallée J, Olivié D, Bérubé J, Fan B, Castel H, Turcotte S, Perreault P, Chagnon M, Tang A. LI-RADS for MR Imaging Diagnosis of Hepatocellular Carcinoma: Performance of Major and Ancillary Features. Radiology. 2018;288:118-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 117] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 13. | Hong CW, Chernyak V, Choi JY, Lee S, Potu C, Delgado T, Wolfson T, Gamst A, Birnbaum J, Kampalath R, Lall C, Lee JT, Owen JW, Aguirre DA, Mendiratta-Lala M, Davenport MS, Masch W, Roudenko A, Lewis SC, Kierans AS, Hecht EM, Bashir MR, Brancatelli G, Douek ML, Ohliger MA, Tang A, Cerny M, Fung A, Costa EA, Corwin MT, McGahan JP, Kalb B, Elsayes KM, Surabhi VR, Blair K, Marks RM, Horvat N, Best S, Ash R, Ganesan K, Kagay CR, Kambadakone A, Wang J, Cruite I, Bijan B, Goodwin M, Moura Cunha G, Tamayo-Murillo D, Fowler KJ, Sirlin CB. A Multicenter Assessment of Interreader Reliability of LI-RADS Version 2018 for MRI and CT. Radiology. 2023;307:e222855. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 14. | Alksas A, Shehata M, Saleh GA, Shaffie A, Soliman A, Ghazal M, Khalifeh HA, Razek AA, El-baz A. A Novel Computer-Aided Diagnostic System for Early Assessment of Hepatocellular Carcinoma. 2020 25th International Conference on Pattern Recognition (ICPR); 2021 Jan 10-15, Milan, Italy. NJ, United States, IEEE, 2021: 10375-10382. [DOI] [Full Text] |
| 15. | Du L, Yuan J, Gan M, Li Z, Wang P, Hou Z, Wang C. A comparative study between deep learning and radiomics models in grading liver tumors using hepatobiliary phase contrast-enhanced MR images. BMC Med Imaging. 2022;22:218. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 18] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 16. | Yamashita R, Mittendorf A, Zhu Z, Fowler KJ, Santillan CS, Sirlin CB, Bashir MR, Do RKG. Deep convolutional neural network applied to the liver imaging reporting and data system (LI-RADS) version 2014 category classification: a pilot study. Abdom Radiol (NY). 2020;45:24-35. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 30] [Article Influence: 5.0] [Reference Citation Analysis (4)] |
| 17. | Park J, Bae JS, Kim JM, Witanto JN, Park SJ, Lee JM. Development of a deep-learning model for classification of LI-RADS major features by using subtraction images of MRI: a preliminary study. Abdom Radiol (NY). 2023;48:2547-2556. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 4] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 18. | Çiçek Ö, Abdulkadir A, Lienkamp SS, Brox T, Ronneberger O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. 2016. Available from: arXiv:1606.06650. [DOI] [Full Text] |
| 19. | Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods. 2021;18:203-211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8060] [Cited by in RCA: 3952] [Article Influence: 790.4] [Reference Citation Analysis (3)] |
| 20. | Tustison NJ, Cook PA, Holbrook AJ, Johnson HJ, Muschelli J, Devenyi GA, Duda JT, Das SR, Cullen NC, Gillen DL, Yassa MA, Stone JR, Gee JC, Avants BB. The ANTsX ecosystem for quantitative biological and medical imaging. Sci Rep. 2021;11:9068. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 362] [Cited by in RCA: 258] [Article Influence: 51.6] [Reference Citation Analysis (4)] |
| 21. | Zheng R, Wang Q, Lv S, Li C, Wang C, Chen W, Wang H. Automatic Liver Tumor Segmentation on Dynamic Contrast Enhanced MRI Using 4D Information: Deep Learning Model Based on 3D Convolution and Convolutional LSTM. IEEE Trans Med Imaging. 2022;41:2965-2976. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 38] [Article Influence: 9.5] [Reference Citation Analysis (5)] |
| 22. | Otsu N. A Threshold Selection Method from Gray-Level Histograms. IEEE Trans Syst Man Cybern. 1979;9:62-66. [DOI] [Full Text] |
| 23. | Frangi AF, Niessen WJ, Vincken KL, Viergever MA. Multiscale vessel enhancement filtering. Lect Notes Comput Sci. 1998;1496:130-137. |
| 24. | McHugh ML. Interrater reliability: the kappa statistic. Biochem Med (Zagreb). 2012;22:276-282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15465] [Cited by in RCA: 10219] [Article Influence: 729.9] [Reference Citation Analysis (3)] |
| 25. | Wang CJ, Hamm CA, Savic LJ, Ferrante M, Schobert I, Schlachter T, Lin M, Weinreb JC, Duncan JS, Chapiro J, Letzen B. Deep learning for liver tumor diagnosis part II: convolutional neural network interpretation using radiologic imaging features. Eur Radiol. 2019;29:3348-3357. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 134] [Cited by in RCA: 112] [Article Influence: 16.0] [Reference Citation Analysis (5)] |
| 26. | Dawit H, Lam E, McInnes MDF, van der Pol CB, Bashir MR, Salameh JP, Levis B, Sirlin CB, Chernyak V, Choi SH, Kim SY, Fraum TJ, Tang A, Jiang H, Song B, Wang J, Wilson SR, Kwon H, Kierans AS, Joo I, Ronot M, Song JS, Podgórska J, Rosiak G, Kang Z, Allen BC, Costa AF. LI-RADS CT and MRI Ancillary Feature Association with Hepatocellular Carcinoma and Malignancy: An Individual Participant Data Meta-Analysis. Radiology. 2024;310:e231501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (0)] |