Published online Apr 7, 2026. doi: 10.3748/wjg.v32.i13.115810
Revised: November 29, 2025
Accepted: February 4, 2026
Published online: April 7, 2026
Processing time: 152 Days and 14.9 Hours
The landscape and clinical utility of comprehensive genomic investigations for a wide range of pediatric gastrointestinal (GI) disorders have not been fully characterized in the Middle East.
To characterize the diagnostic yield and clinical utility of genomic investigations in a Middle Eastern pediatric cohort of GI disorders, and to dissect the pathogenic landscape of those disorders in this region.
Sixty-nine pediatric patients of diverse Arab and Asian origins, were clinically and genetically assessed for a spectrum of GI diseases, including liver disease, inflammatory bowel disease, chronic diarrhea, and pancreatitis. Clinical genomic investigations included mainly (87%) next generation sequencing-based gene panels and whole exome or genome sequencing. Clinical information, including demographics, symptoms, management and clinical outcomes, was extracted from medical records.
The overall positive yield was 55%, whereas multiple molecular diagnoses were made in 3 patients (4%) including 2 with triple genetic findings, highlighting the utility of genetic investigations in delineating the phenotypic complexity in this cohort. A secondary medically actionable finding (MYBPC3-associated cardiomyopathy) was identified in one out of 12 patients (8%) who re
Our study provides new insights into the pathogenic variation landscape in pediatric GI disorders in the Middle East and emphasizes the clinical utility of genomic investigations in managing those patients.
Core Tip: Here we characterize the diagnostic and clinical utility of genomic investigations for a wide range of pediatric gastrointestinal disorders in a pediatric cohort from the Middle East. We show that the cumulative diagnostic yield was 55%, including 6% having multiple molecular diagnoses, mostly attributed to autosomal recessive disorders (66%). Yield was highest for patients with congenital diarrhea (73.3%) and cholestasis (62.5%). Genomic findings guided management plans in 97% of diagnosed patients. We propose a novel gene-disease association most likely due to biallelic loss-of-function variants in the NR1I3 gene.
- Citation: Alsarhan A, Alloush R, Jain R, Abou Tayoun A, Tzivinikos C. Clinical utility of genomic investigations in a Middle Eastern pediatric gastroenterology disease cohort. World J Gastroenterol 2026; 32(13): 115810
- URL: https://www.wjgnet.com/1007-9327/full/v32/i13/115810.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i13.115810
The rapid advances in the genomics field have democratized clinical genetic testing which became widely accessible for diagnosing a variety of challenging diseases. The ability to sequence and analyze comprehensive gene lists, up to the whole exome, in a short period of time has shifted clinicians’ practice from ‘confirmatory’, often phenotype-driven testing, into a genotype-driven diagnostic paradigm especially for diseases with marked phenotypic and genetic heterogeneity. As of June 16, 2025, there are 7000 phenotypes for which the genetic and molecular basis are known[1], and there are ongoing studies to identify additional novel gene-disease associations. This is particularly important in the Arab population of the Middle East considering the high rate of consanguinity which can reach up to 20%-50%[2-6].
Several studies investigated the utility of genomics in gastroenterology with diagnostic rates varying based on clinical presentations. Jeyaraj et al[7] described a pediatric cholestatic cohort for which genetic testing yielded positive results in 8.5%, and emphasized the potential genotype-phenotype correlation. In pediatric Inflammatory bowel disease (IBD), the chance of getting a positive genetic result is higher with younger patients’ ages at which the diagnosis was established. The pathogenic variation spectrum in this cohort mostly impacts the immunological pathway rendering patients with unpredictable response to conventional treatment[8]. Mayerle et al[9] described the genetic basis of recurrent and chronic pancreatitis, and showed that targeting these receptors, receptors like inositol-tri-phosphate-receptor (types 2 and 3) where pathogenic variants have been identified, may help ameliorate pancreatitis. Another study showed genetic testing for congenital diarrhea or enteropathy was able to identify a molecular diagnosis in 64.2 % of the cohort (n = 137). The study showed that age and clinical presentation affected diagnostic yield, which was higher in neonates (75.4% vs 57.6%) and in cases of fatty (71.4%) or bloody diarrhea (68.0%) compared to watery diarrhea (48.1%)[10].
Besides diagnostic utility, genetic testing can assist in the management of various conditions and predict progression. For example, mutation class underlying cystic fibrosis disease can assist in anticipating and predicting the risk of developing pancreatitis[11]. In patients with microvillous inclusion disease (MVID), identifying pathogenic variants in MYO5B helps determine whether the disease will present as isolated intestinal involvement, a mixed intestinal-hepatic phenotype, or mimic other conditions such as progressive familial intrahepatic cholestasis (PFIC). This influences decisions around initiating parenteral nutrition, monitoring for cholestasis, and considering early intestinal or liver transplantation[12].
Currently, there are limited studies investigating the landscape and clinical utility of genomics in gastrointestinal (GI) diseases in pediatric populations across different ancestries, specifically those of Middle Eastern origin.
Here, we characterize the diagnostic yield, molecular findings, and clinical outcomes of genomic investigations in 69 pediatric patients, of primarily Middle Eastern origin, who presented with a range of GI associated symptoms.
This study includes gastroenterology patients referred for clinical genomic testing from April 2019 to December 2023. Review of electronic medical records was performed by gastroenterologists to retrieve all clinical and demographics data and to identify interventions guided by genetic results.
Clinical indications included six groups: (1) Liver disease: Defined in patients with direct hyperbilirubinemia, persis
This study was reviewed and approved by the Dubai Scientific Research Ethics Committee, Dubai Health Authority (Approval No. DSREC-08/2025_03). All patients were consented for clinical genetic testing under an approved de-identified research protocol, which permits the publication of de-identified analyses.
Genomic testing was performed in our College of American Pathologists-accredited genomics facility. Exome sequencing was performed as previously described[6]. Briefly, following DNA fragmentation by ultrasonication (Covaris, United States), the coding regions of the genome, also known as the exome, were captured using the Agilent Clinical Research Exome V2 capture probes (Agilent, United States). Libraries were prepared using the SureSelectXT protocol (Agilent, Agilent) and then sequenced (2 × 150 bp) using the NovaSeq6000 system (Illumina, Agilent) to a minimum average depth of 100 ×.
Sequencing data were processed using an in-house custom-made bioinformatics pipeline to retain high-quality sequen
For indication-based analysis, only rare, known pathogenic, or novel variants in the relevant genes associated with the patient’s indication, including cholestasis, pancreatitis, congenital diarrhea, and VEOIBD (Supplementary Table 1) were retained for interpretation.
For whole exome sequencing (WES), all known pathogenic variants in ClinVar/HGMD and novel loss-of-function variants in disease genes were retained. In addition, segregation analysis was performed for trio WES to identify domi
Chromosomal microarray analysis (CMA) was preformed using the Affymetrix CytoScanHD.
All retained sequence and copy number variants were classified following the American College of Medical Genetics and Genomics/Association for Molecular Pathology or the American College of Medical Genetics and Genomics/Clinical genome Resource variant interpretation guidelines, respectively[18-20]. Pathogenic and likely pathogenic variants in genes relevant to the patients’ primary indications were reported and were considered diagnostic if the patient’s phenotype (based on physician’s notes and feedback), disease mechanism, and inheritance were all consistent. Clinically significant heterozygous variants in genes with recessive inheritance and all variants of uncertain significance relevant to patients’ primary indications were also reported, though were not considered diagnostic, leading to inconclusive reports.
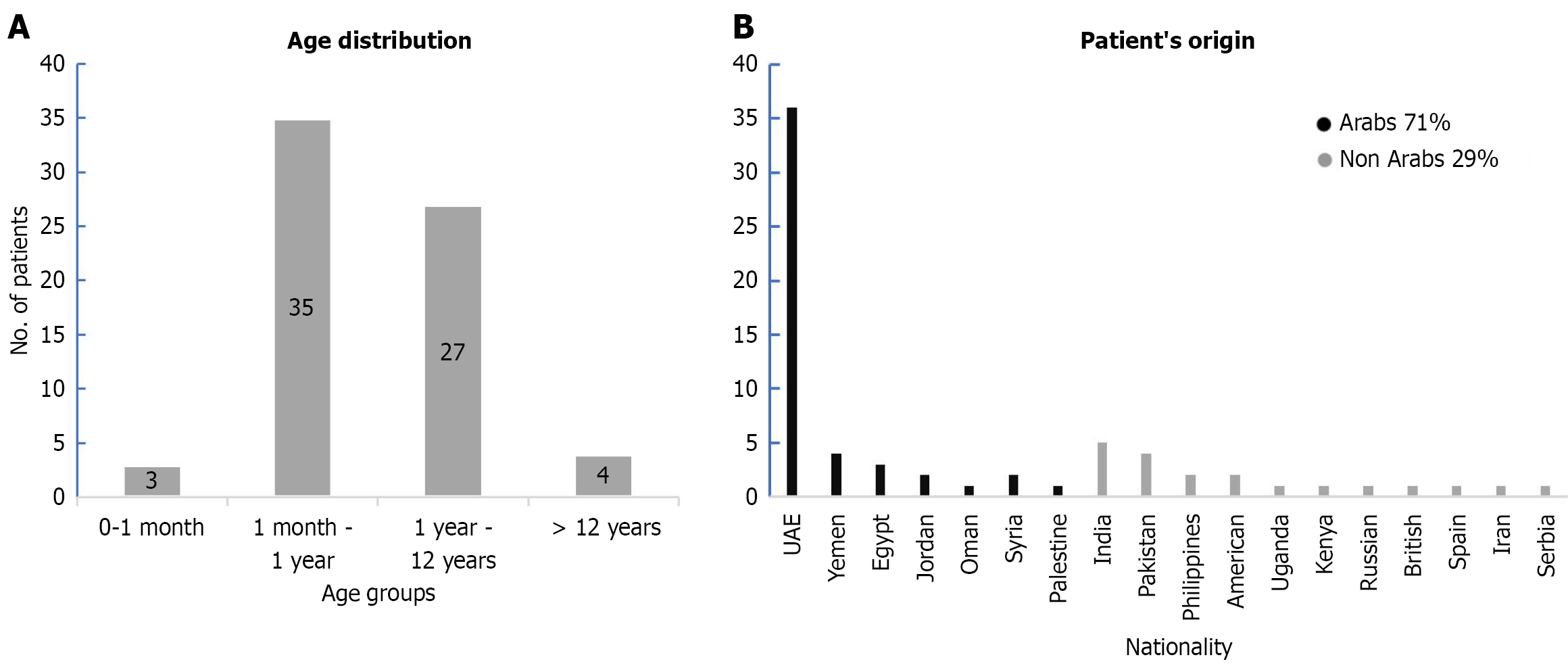
Sixty-nine patients (average age: 2.8 years, range 2 weeks-18 years; 62.3% males; Figure 1) underwent genetic testing for a range of GI indications. Most patients (71%) were Arabs with majority being Emiratis (52.1%) and Yemenis (5.7%); 29% were mostly non-Arabs Asians (Figure 1).
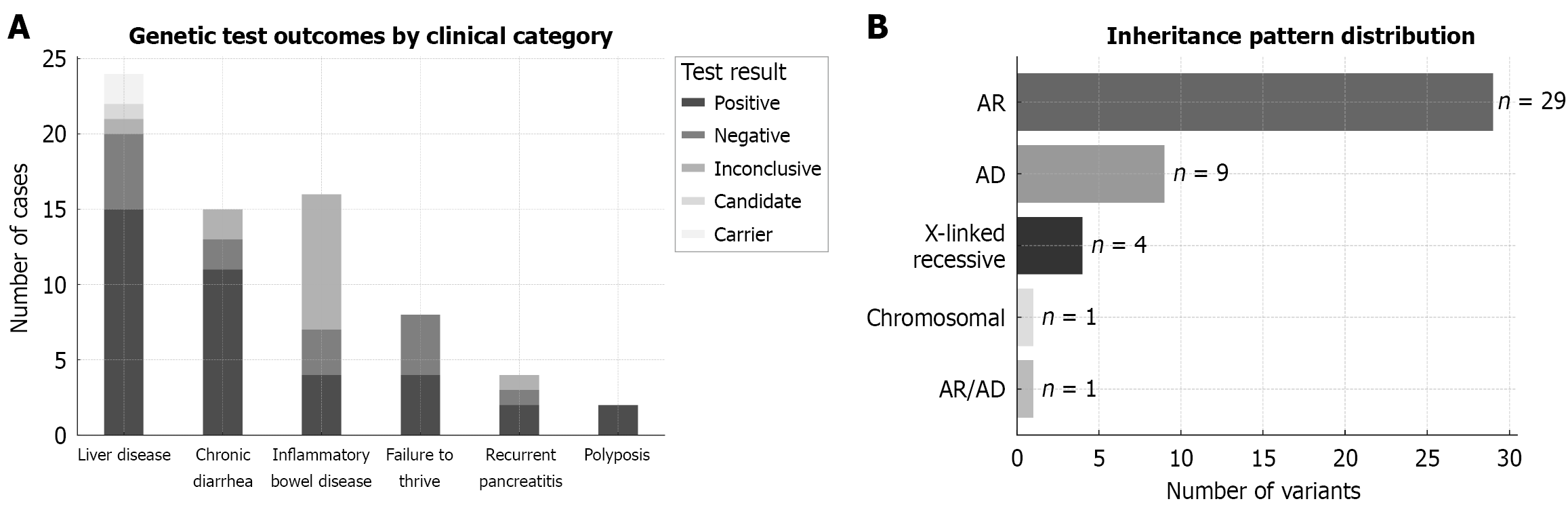
Patients mainly presented with liver disease (34.8%), chronic diarrhoea (21.8%), IBD (23.2%), failure to thrive (11.6%), recurrent pancreatitis (5.8%), and polyposis (2.9%; Figure 2A). Most patients were tested by genomic sequencing (86.9%), while CMA was done in 7 patients (10.1%; Supplementary Table 1).
The overall positive diagnostic yield was 55% (38/69), whereas multiple molecular diagnoses were made in 3 patients (4%) including 2 with triple genetic findings. In the liver (cholestasis) disease group, 15 patients (62.5%) had positive genetic results, while 4 patients with IBD (25%) were positive. A secondary medically actionable finding (MYBPC3-associated cardiomyopathy) was identified in one out of 12 patients (8%) who received exome or genome sequencing. Aside from the two patients with polyposis and positive findings, the highest diagnostic yield in this cohort was in the chronic diarrhea group where 11 patients (73.3%) had diagnostic results. The diagnostic yields in patients with pancreatitis and failure to thrive, were 50% each (Figure 2A). A list of all genetic findings is summarized in Table 1.
| ID | Primary indication | Technology used | Test done | Variants | Inheritance | Zygosity | Classification |
| 6 | Very early onset IBD | NGS | WESTRIO | NM_000377.3(WAS):c.383T>C; p.(Phe128Ser) | X-linked recessive | Hemizygous | LP |
| 14 | Very early onset IBD | CMA | CMA | Xp21.1p11.4 (CYBB gene) deletion 1.83 Mb | X-linked recessive | Hemizygous | Pathogenic |
| 15 | Recurrent peri-anal abscess | CMA | CMA | Xp21.1p11.4 (CYBB gene) deletion 1.83 Mb | X-linked recessive | Hemizygous | Pathogenic |
| 16 | Very early onset IBD | NGS | WES | TTC7A: NM_020458.4, c.133_166del (p.Gly45SerfsTer23) | AR | Heterozygous | LP |
| 17 | Liver failure and direct hyperbilirubinemia | NGS | Custom gene panel | NM_002437.5(MPV17):c.280G>C; p.(Gly94Arg) | AR | Homozygous | LP |
| 19 | Direct hyperbilirubinemia, multiple congenital anomalies | CMA | CMA | arr[GRCh37] 18p11.32q23(136,227_78,014,123)x3 | Chromosomal disorder | - | Pathogenic |
| 20 | Persistent indirect hyperbilirubinemia | NGS | UGT1A1 full gene sequencing | NM_000463.3(UGT1A1):c.625C>T; p.(Arg209Trp) NM_000463.3(UGT1A1): C.-41_-40dupTA; p.? | AR AR | Homozygous homozygous | Pathogenic LP |
| 21 | Persistent indirect hyperbilirubinemia | NGS | Cholestasis panel | NM_000463.3(UGT1A1): C.-41_-40dupTA; p.? | AR | Homozygous | Pathogenic |
| 23 | Persistent indirect hyperbilirubinemia | NGS | Cholestasis panel | NM_005603.4(ATP8B1):c.3040C>T; p.(Arg1014*) | AR | Homozygous | Pathogenic |
| 24 | Persistent indirect hyperbilirubinemia | NGS | Cholestasis panel | NM_000463.2(UGT1A1): C.1021C>T; p.R341* | AR | Homozygous | Pathogenic |
| 25 | Persistent elevated liver enzymes, congenital heart anomalies | NGS | Alagile syndrome | NM_000214.2(JAG1): C.1052delG; p.(Cys351 Leufs*61) | AD | Heterozgous | Pathogenic |
| 26 | Persistent elevated liver enzymes, congenital heart anomalies | NGS | Unknown | NM_000214.2(JAG1): C.1052delG; p.(Cys351 Leufs*61) | AD | Heterozgous | Pathogenic |
| 27 | Elevated liver enzymes and hepatosplenomegaly | NGS | WES | NM_000443.4(ABCB4):c.3634-4A>G; p.? NM_000443.4(ABCB4):c.1864G>T; p.(Gly622Trp) | AD/AR | Compound heterozygous | VUS |
| 28 | Elevated liver enzymes and hepatosplenomegaly | NGS | Custom gene panel | NM_000443.4(ABCB4):c.158A>T; p.(Asp53Val) NM_004004.6(GJB2):c.-23+1G>A; p.?NM_001042351.3(G6PD):c.563C>T; p.(Ser188Phe) | AR AR X-linked recessive | Homozygous homozygous hemizygous | VUS pathogenic pathogenic |
| 34 | Direct hyperbilirubinemia | Sanger | Targeted variant analysis | NM_025193.4(HSD3B7):c.45_46del; p.(Gly17 Leufs*26) | AR | Homozygous | Pathogenic |
| 35 | Persistent indirect hyperbilirubinemia | NGS | Gilbert syndrome genetic test | NM_000463.3(UGT1A1): C.-41_-40dupTA; p.? | AR | Homozygous | Pathogenic |
| 37 | Persistent indirect hyperbilirubinemia | NGS | Crigler-Najjar syndrome genetic test | NM_000463.3(UGT1A1): C.-41_-40dupTA; p.? | AR | Homozygous | Pathogenic |
| 38 | Persistent indirect hyperbilirubinemia | NGS | UGT1A1 full gene sequencing | NM_000463.3(UGT1A1): C.-41_-40dupTA; p.? | AR | Homozygous | Pathogenic |
| 40 | Direct hyperbilirubinemia | NGS | WGS | 4.70 Mb deletion JAG1 gene 20p12 | AD | Heterozygous | Pathogenic |
| 43 | Chronic pancreatitis | NGS | WES | NM_002769.4(PRSS1):c.365G>A; p.(Arg122His) | AD | Heterozygous | Pathogenic |
| 44 | Recurrent pancreatitis | NGS | Pancreatitis panel | NM_007272.3 (CTRC):c.738_761del; p.(Lys247_Arg254del) | AD | Heterozygous | Pathogenic |
| 45 | Chronic congenital diarrhea | NGS | Custom gene panel | NM_021102.4(SPINT2):c.442C>T; p.(Arg148Cys) NM_000277.3(PAH):c.157C>T; p.(Arg53Cys) NM_144687.4(NLRP12):c.1854C>G;p.(Tyr618*) | AR AR AD | Homozygous homozygous heterozygous | LP LP LP |
| 48 | Chronic congenital diarrhea | NGS | Unknown | NM_001080467.3(MYO5B):c.1966C>T; p.(Arg656Cys) | AR | Homozygous | LP |
| 49 | Chronic congenital diarrhea | NGS | Unknown | 17 kb deletion in EPCAM gene 2p21 | - | - | - |
| 50 | Chronic congenital diarrhea | NGS | Unknown | NM_001080467.3(MYO5B):c.1966C>T; p.(Arg656Cys) | AR | Homozygous | LP |
| 51 | Chronic congenital diarrhea, albinism, dysmorphism | NGS | Unknown | NM_014639.4(SKIC3):c.4070del; p.(Pro1357 Leufs*10) | AR | Homozygous | LP |
| 52 | Chronic congenital diarrhea | NGS | Chronic Congenital Diarrhea panel | NM_001080467.3(MYO5B):c.82del; p.(Thr28Profs*47) | AR | Homozygous | LP |
| 53 | FTT, persistent diarrhea, direct hyperbilirubinemia | NGS | Cholestasis panel | NM_020198.3(CCDC47):c.1234C>T; p.(Arg412*) | AR | Homozygous | LP |
| 56 | Non-mechanical intestinal obstruction, diarrhea | NGS | WES | NM_000111.2(SLC26A3):c.559G>T; p.(Gly187*) | AR | Homozygous | Pathogenic |
| 57 | FTT | NGS | Custom gene panel | NM_006408.4(AGR2):c.104del; p.(Asp35Alafs*38) | AR | Homozygous | LP |
| 58 | Chronic congenital diarrhea | NGS | WESTRIO | NM_001080467.2(MYO5B):c.2062C>T; p.(Arg688*) NM_000463.2(UGT1A1):c.1075G>A; p.(Asp359Asn) NM_000492.3(CFTR):c.1163C>T; p.(Thr388Met) | AR | Homozygous heterozygous heterozygous | Pathogenic VUS VUS |
| 59 | FTT, and chronic congenital diarrhea | NGS | WES | NM_012079.5(DGAT1):c.1374G>A; p.(Trp458*) | AR | Homozygous | LP |
| 61 | FTT | NGS | Cystic fibrosis panel | NM_000492.4(CFTR):c.1521_1523del; p.(Phe508del) | AR | Homozygous | Pathogenic |
| 63 | FTT | Sanger | Targeted variant analysis | NM_001012331.1(NTRK1):c.1624del; p.(Glu542Argfs*110) | AR | Homozygous | LP |
| 64 | FTT | NGS | Comprehensive lung panel | NM_001013838.3(CARMIL2):c.950dup; p.(Pro318Thrfs*44) | AR | Homozygous | LP |
| 65 | FTT | NGS | WESTRIO | NM_004333.6(BRAF):c.1574T>C; p.(Leu525Pro) | AD | Heterozygous | Pathogenic |
| 68 | Ileal mass and polyps | NGS | Unknown | NM_000455.5(STK11):c.300dup; p.(Gly622Trp) | AD | Heterozygous | LP |
| 69 | Multiple Polyp on multiple occasions | NGS | Unknown | STK11 gene deletion (unknown) | AD | Heterozygous | Pathogenic |
Sequencing-based testing (exome, genome, targeted gene panels) yielded an overall diagnostic rate of 49% (26/53) while CMA resulted in a lower diagnostic yield of 42.9% (3/7). Notably, the VEOIBD panel was performed in 12 patients but did not identify a positive result in any case (Table 2).
| Test/panel | Positive | Total | % |
| WES | 5 | 8 | 62.5 |
| WESTRIO | 3 | 3 | 100.0 |
| WGS | 1 | 1 | 100.0 |
| VEOIBD panel | 0 | 12 | 0.0 |
| Custom gene panel | 4 | 4 | 100.0 |
| Cholestasis panel | 4 | 10 | 40.0 |
| CMA | 3 | 7 | 42.9 |
| UGT1A1 full gene sequencing | 2 | 2 | 100.0 |
| Gilbert syndrome genetic test | 1 | 2 | 50.0 |
| Crigler-Najjar syndrome genetic test | 1 | 1 | 100.0 |
| Targeted variant analysis | 2 | 2 | 100.0 |
| Pancreatitis panel | 1 | 2 | 50.0 |
| Chronic congenital diarrhea panel | 1 | 3 | 33.3 |
| Cystic fibrosis panel | 1 | 1 | 100.0 |
| Comprehensive lung panel | 1 | 1 | 100.0 |
| Alagille syndrome | 1 | 1 | 100.0 |
| Congenital mono- and disaccharide disorders panel | 0 | 2 | 0.0 |
| Unknown | 7 | 7 | 100.0 |
Consistent with the higher consanguinity rates in the Middle East[4], conditions with autosomal recessive (AR) in
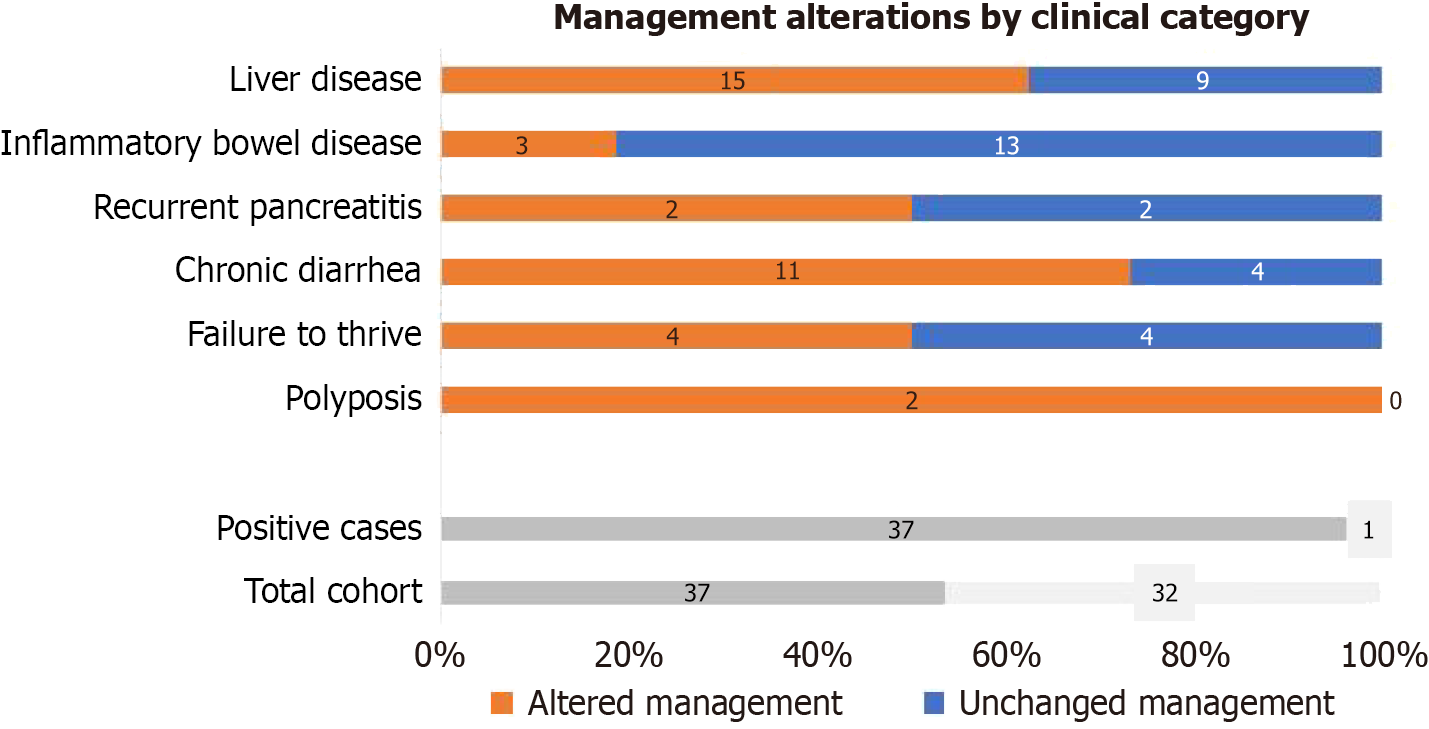
When considering the clinical context, diagnostic genetic results altered the clinical approach and management in 97% (37/38) of the positive cases and 53.6% of the whole cohort. Clinical management varied by indication and the genetic findings. Figure 3 illustrates the proportion of cases with altered management within each disease group, as well as among the total cohort and total genetically positive cases. In addition, all families with positive findings received genetic counselling to assess recurrence risks and to guide families in considering preventive strategies for future pregnancies. Table 3 summarizes the clinical diagnoses and corresponding management changes applied to each patient.
| ID | Clinical diagnosis | Clinical implication |
| 6 | Wiskott-Aldrich syndrome or related phenotypes | Nil |
| 14 | X-linked chronic granulomatous disease | Requires BMT; no response to conventional therapy |
| 15 | X-linked chronic granulomatous disease | Requires BMT; no response to conventional therapy |
| 16 | Combined immunodeficiency with multiple intestinal atresias | Initiated leflunomide, counseled on immunodeficiency risk |
| 17 | Mitochondrial DNA depletion syndrome | Liver transplant referral; neurology referral; prenatal screening recommended |
| 19 | Trisomy 18 | Liver transplant not pursued |
| 20 | Crigler-Najjar (and Gilbert syndrome) | Intensive phototherapy; phenobarbital; transplant counseling; potential gene therapy |
| 21 | Gilbert syndrome | Avoidance of further testing |
| 23 | PFIC 1 | UDCA, IBAT inhibitor; hearing test; anticipate cirrhosis and possible liver transplant |
| 24 | Crigler-Najjar | Intensive phototherapy; phenobarbital; transplant counseling; potential gene therapy |
| 25 | Alagille syndrome | Screening for associated disorders: Cardiac, ocular, vascular, and renal |
| 26 | Alagille syndrome | Screening for associated disorders: Cardiac, ocular, vascular, and renal |
| 27 | PFIC 3 | UDCA, IBAT inhibitor; slower disease progression |
| 28 | PFIC 3/autosomal recessive nonsyndromic hearing loss/glucose-6-phosphate dehydrogenase deficiency | UDCA, IBAT inhibitor; slower disease progression; counseling on post–liver transplant recurrence; G6PD precautions; hearing test; ENT referral |
| 34 | Congenital bile acid synthesis defect | Bile acid replacement therapy; liver transplant considered |
| 35 | Gilbert syndrome | Avoidance of further testing |
| 37 | Gilbert syndrome | Avoidance of further testing |
| 38 | Gilbert syndrome | Avoidance of further testing |
| 40 | Alagille syndrome | IBAT inhibitor, screening for associated disorders: Cardiac, ocular, vascular, and renal |
| 43 | Hereditary pancreatitis | Annual pancreatic cancer screening; regular monitoring of exocrine/endocrine function |
| 44 | Chronic pancreatitis | Regular monitoring of exocrine/endocrine function |
| 45 | TE, hyperphenylalaninemia, familial cold autoinflammatory syndrome | Family counseling for future pregnancies; MVT not favored as symptoms may improve |
| 48 | MVID | Family counseling for future pregnancies; renal screening; MVT likely as symptoms unlikely to improve; lifelong TPN anticipated |
| 49 | TE | Family counseling for future pregnancies; MVT not favored as symptoms may improve |
| 50 | MVID | Family counseling for future pregnancies; renal screening; MVT likely as symptoms unlikely to improve; lifelong TPN anticipated |
| 51 | Trichohepatoenteric syndrome | Screening for hypogammaglobulinemia, screening for liver disease |
| 52 | MVID | Family counseling for future pregnancies; renal screening; MVT likely as symptoms unlikely to improve; lifelong TPN anticipated |
| 53 | Trichohepatoneurodevelopmental syndrome | Screening for developmental delay, screening for liver disease |
| 56 | Congenital chloride diarrhea | Chloride supplementation |
| 57 | Recurrent respiratory infections and failure to thrive with or without diarrhea | Family counseling for future pregnancies |
| 58 | MVID | Family counseling for future pregnancies; renal screening; MVT likely as symptoms unlikely to improve; lifelong TPN anticipated |
| 59 | Congenital diarrheal disorder due to DGAT1 deficiency | Family counseling for future pregnancies; considering MVT |
| 61 | Cystic fibrosis | Annual screening for pancreatic insufficiency, started Trikafta |
| 63 | Autosomal recessive TRK 1 positive congenital insensitivity to pain with anhydrosis | Antipyretics and cooling measures; trauma precautions due to absent pain; screen for immunoglobulin deficiency |
| 64 | Immunodeficiency | Screening for immunodeficiency |
| 65 | BRAF gene related disorders (cardiofaciocutaneous syndrome, Noonan syndrome and LEOPARD syndrome) | Family counseling regarding high risk of developing tumors, referral to cardiology, started Growth hormone |
| 68 | Peutz-Jeghers syndrome | Regular endoscopic screening; genetic testing for first-degree relatives |
| 69 | Peutz-Jeghers syndrome | Regular endoscopic screening; genetic testing for first-degree relatives |
This was largest group of patients in our cohort where 24 presented mainly with elevated liver enzymes, hyperbilirubinemia (both direct and indirect) and liver failure. Majority of these cases were encountered during infancy, and the genetic testing was considered at some point the only modality left to establish the diagnosis. Several pathologies were identified such as: Mitochondrial DNA depletion syndrome, Trisomy 18, Crigler-Najjar syndrome, Gilbert syndrome, PFIC, Alagille syndrome, and congenital bile acid synthesis defect. As shown in Figure 2A, the yield of the genetic testing in this group was high (62.5%) and it helped to identify the diagnosis where other modalities have failed. The manage
Patients with unexplained indirect hyperbilirubinemia often undergo frequent testing, and possibly invasive pro
Cholestasis or itching can manifest as presenting symptoms in variety of cases. In PFIC (n = 2), molecular findings can confirm the diagnosis and aids in identifying subtypes, which significantly influence prognosis. This information helped to counsel the parents and to decide management. One patient had ATP8B1 homozygous pathogenic variants (PFIC 1) and needed referral for hearing assessment since this disease is associated with hearing loss[24]. Family was informed that the prognosis remains poor despite liver transplantation patient No. 27 with ABCB4 compound heterozygous variant diagnosed with PFIC 3 disease had a milder disease and responded well to ursodeoxycholic acid. Both families were informed that ileal bile acid transporter (IBAT) inhibitors may be considered in cases of inadequate treatment response or disease progression. On the other hand, patient No. 34 was diagnosed with bile acid synthesis defects, and was started on bile acid replacement therapy.
Genetic testing has proven invaluable in guiding the management of complex and multisystem presentations. In confirmed Alagille syndrome cases (n = 3), identifying associated defects of the heart, eyes, vascular system (including the risk of intracranial bleeding), and kidneys are crucial for comprehensive care planning[25]. Accordingly, those patients were referred for several speciality clinics to screen for the associated defects. Also, patient No. 40 was started on IBAT inhibitors. In the case of mitochondrial DNA depletion syndrome (n = 1), the patient presented with acute liver failure and she was referred for liver transplant at another center[25]. Patient No. 19 was diagnosed with trisomy 18 and was ineligible for liver transplant due to poor outcomes associated with the syndrome and we did not offer the family this therapy option[26]. These examples highlight the importance of genetic testing in taking vital and critical decisions which can be quite significant to patients and their families.
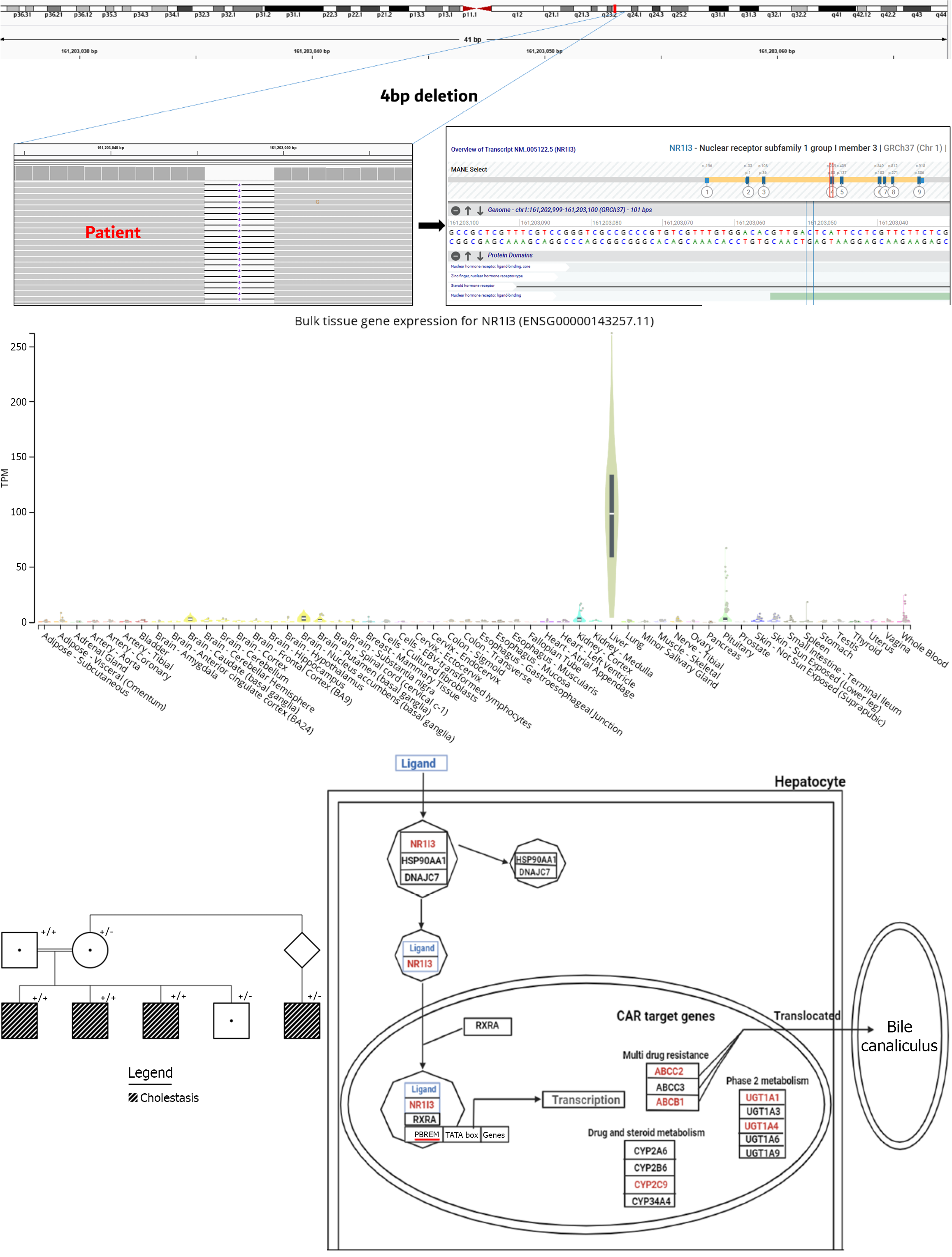
Patient No. 29 presenting with transient episodic itching and cholestasis had negative exome sequencing, though further analysis identified a homozygous 4 bp deletion in exon 4 (NM_005122.5) of a novel gene, NR1I3 (Figure 4). This 4 bp deletion (c.318_321del), which is extremely rare (allele frequency 0.0022%) in the general population, namely in the gnomAD (causes a frameshift which is predicted to alter the protein’s amino acid sequence beginning at position 107 and lead to a premature termination codon 6 amino acids downstream (p.Ser107Argfs*6). This alteration is then predicted to lead to a truncated receptor lacking the ligand-binding domain or, most likely, an absent protein given the expected nonsense mediated decay.
The NR1I3 gene encodes a nuclear receptor which is strictly expressed in the liver, based on human gene expression data, and is thought to regulate the transcription of genes involved in drug metabolism and bilirubin clearance (Figure 4). A previous study suggested a link between NR1I3 variants and hyperbilirubinemia, with preliminary evidence indicating a possible role in bilirubin metabolism[27].
Familial testing showed that the similarly affected two siblings were also homozygous for the NR1I3 frameshift variant. However, their father, who does not recall a history of liver disease, was also homozygous while an affected cousin was heterozygous carrier for this variant (Figure 4). While this information may suggest non-segregation and may not support a causal role for NRI13 in cholestasis, functional analyses are needed to characterize the penetrance expressivity, and inheritance of this potentially novel gene in light of the above evidence and given the progressive and episodic nature of this disease. Notably, the patient’s older sibling’s symptoms resolved over time, raising the possibility that the father may have experienced a similar self-limited disease course during childhood.
A total of 16 patients presented with IBD in this cohort. Patient No. 16 presented with diarrhoea and early onset of bowel inflammation, ultimately leading to bowel atresia and short gut syndrome necessitating lifelong parenteral nutrition. Exome sequencing (targeted gene panel sequencing) identified a homozygous pathogenic variant in the TTC7A gene pathogenic variant was identified. TTC7A pathogenic variants typically coincide with immunodeficiency in up to 90% of cases and necessitate screening, our patient did not display overt immunodeficiency[28]. This genetic finding was helpful for anticipating prognosis, highlighting the possible associated immunodeficiency problems reassuring the family that GI symptoms might slightly improve with time[28]. Furthermore, the patient was treated with leflunomide without clinical benefit and was later offered a MVT, which could potentially provide long-term survival and independence from parenteral nutrition.
A significant outcome of genetic testing in VEOIBD patients lies in identifying underlying immunodeficiency as a primary condition. We documented the case of two male siblings (patients No. 14 and No. 15) harbouring an X-linked recessive CYBB pathogenic deletion, confirming the diagnosis of chronic granulomatous disease. This finding was impor
Genetic testing was performed for four cases with pancreatitis, revealing positive results in two individuals. Specifically, pathogenic variants were identified in the PRSS1 and CTRC genes. PRSS1 pathogenic variants are the most common cause of pancreatitis. This gene plays a role in regulating trypsin function, which can lead to immature and excessive activation leading to inflammation and pancreatitis[31]. On the other hand, the CTRC gene is responsible for facilitating trypsin lysis. Loss of function variants in this gene result in the loss of its protective role against trypsin activation[31]. Both cases were scheduled for regular follow up to screen for pancreatic exocrine and endocrine function, in addition to ultrasound screening for pancreatic cancer in the patient with the PRSS1 variant which has increased risk of malignancy.
In patient No. 41, although a definitive diagnosis explaining pancreatitis was not identified, a homozygous variant of uncertain clinical significance was identified in the TULP3 gene which has been linked to liver and kidney disease, as well as cardiomyopathy[32]. Interestingly, the same variant has been described in the literature in two patients with chronic pancreatitis[32]. This patient also had a secondary pathogenic variant in MYBPC3, which is associated with risk of dilated cardiomyopathy and was therefore referred for additional surveillance and familial testing[33].
A total of 15 patients were included in this cohort. They presented with chronic diarrhea and were genetically confirmed to have the following diagnoses: MVID, tufting enteropathy (TE), tricho-hepato-neurodevelopmental syndrome (THNS), tricho-hepato-enteric syndrome (THES), glucose galactose malabsorption, congenital sucrase-isomaltase deficiency and others. These patients typically present with diarrhea, which can vary from mild to severe. However, it is not possible to differentiate the underlying diagnosis based solely on the clinical presentation.
In this cohort, 4 patients with MVID exhibited homozygous pathogenic variants in the MYO5B gene. Clinical pre
TE can be considered a favourable diagnosis since its prognosis is good with long-term survival reaching up to 90%, and enteral autonomy can range between 50%-75% at age of 25 years[36]. In patient No. 45, two homozygous likely pathogenic variants were identified in SPINT2 and PAH, explaining the chronic diarrhoea and suggesting phenylketonu
In patient No. 49, who presented with diarrhoea shortly after birth, a pathogenic deletion in the EPCAM gene con
The main reason for performing genetic testing in this group, which included 8 patients, was failure to thrive, yet all the patients had other symptoms such as diarrhea, vomiting, developmental delay, and dysmorphism and therefore genetic findings, which were positive in 4 out of the 8 patients (50%) in this group, helped characterize the underlying disease. For example, the identification of a mutation in the NTRK1 gene confirmed the diagnosis of AR congenital insensitivity to pain with anhidrosis. This condition requires special care of the limbs and appropriate temperature control. A BRAF gene mutation was found in another patient which is associated with Noonan syndrome characterized with failure to thrive, short stature, dysmorphic features, and congenital heart defects[37]. Therefore, patient was referred to cardiology for further evaluation and was started on growth hormone therapy. A pathogenic variant in the CARMIL2 gene was found in patient No. 64 who presented with recurrent chest infection and failure to thrive. Loss of function of this gene causes primary immunodeficiency along with failure to thrive, and recently was linked to the development of VEOIBD[38]. This patient was therefore referred for immunological screening. Finally, a homozygous pathogenic variant in the CFTR gene was identified in patient No. 61. Subsequently, the patient was diagnosed with cystic fibrosis and referred to a specialized clinic for regular follow-up and surveillance.
This small group comprised only two patients, both presenting with multiple polyps, and one experiencing intussusception. Both patients exhibited heterozygous pathogenic variants in the STK11 gene, which causes an AD Peutz-Jeghers syndrome. These individuals displayed distinctive polyp characteristics on histopathological analysis, though the diagnosis was confirmed by genetic testing. Following diagnosis, patients required routine endoscopic surveillance and regular screening for extraintestinal cancers due to the increased risk. Moreover, genetic test screening was offered to first-degree relatives of affected individuals with known genetic mutations[39].
This study has several limitations. First, the sample size is relatively small, which may limit the generalizability of our findings across the broader pediatric gastroenterology population. Second, we were unable to obtain long-term follow-up for many patients. A significant proportion of families relocated to other countries or transferred care to other insti
In conclusion, genomic investigations serve as a critical tool in pediatric GI diseases, aiding in diagnosis, etiological understanding, and treatment guidance. Despite ongoing gaps in our knowledge of many genes, the expanding genomic databases offer promising prospects for further insights. Our findings underscore the significant impact of genetic testing on altering management strategies and providing valuable prognostic information. Furthermore, we identified a novel gene potentially associated with a new form cholestasis. Continued research and technological advancements in genetic testing hold the promise of improving diagnostic and therapeutic approaches, ultimately enhancing patient outcomes and quality of life.
| 1. | OMIM Entry Statistics. Number of Entries in OMIM. [cited 25 July 2025]. Available from: https://omim.org/statistics/entry. |
| 2. | Abou Tayoun AN, Rehm HL. Genetic variation in the Middle East-an opportunity to advance the human genetics field. Genome Med. 2020;12:116. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 3. | Abou Tayoun AN, Fakhro KA, Alsheikh-Ali A, Alkuraya FS. Genomic medicine in the Middle East. Genome Med. 2021;13:184. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 19] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 4. | Chekroun I, Shenbagam S, Almarri MA, Mokrab Y, Uddin M, Alkhnbashi OS, Zaki MS, Najmabadi H, Kahrizi K, Fakhro KA, Almontashiri NAM, Ali FR, Özbek U, Reversade B, Alkuraya FS, Alsheikh-Ali A, Abou Tayoun AN. Genomics of rare diseases in the Greater Middle East. Nat Genet. 2025;57:505-514. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 6] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 5. | Chekroun I, Rabea F, Jain R, Alsheikh-Ali A, Abou Tayoun A. Premarital genomic screening in Arab populations of the Middle East. Nat Med. 2025;31:364-365. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 6. | El Naofal M, Ramaswamy S, Alsarhan A, Nugud A, Sarfraz F, Janbaz H, Taylor A, Jain R, Halabi N, Yaslam S, Alfalasi R, Shenbagam S, Rabea F, Bitzan M, Yavuz L, Wafadari D, Abulhoul H, Shankar S, Al Maazmi M, Rizk R, Alloub Z, Elbashir H, Babiker MOE, Chencheri N, AlBanna A, Sultan M, El Bitar M, Kherani S, Thalange N, Alshryda S, Di Donato R, Tzivinikos C, Majid I, Freeman AF, Gonzalez C, Khan AO, Hamdan H, Abuhammour W, AlAwadhi M, AlKhayat A, Alsheikh-Ali A, Abou Tayoun AN. The genomic landscape of rare disorders in the Middle East. Genome Med. 2023;15:5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 26] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 7. | Jeyaraj R, Bounford KM, Ruth N, Lloyd C, MacDonald F, Hendriksz CJ, Baumann U, Gissen P, Kelly D. The Genetics of Inherited Cholestatic Disorders in Neonates and Infants: Evolving Challenges. Genes (Basel). 2021;12:1837. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 2.2] [Reference Citation Analysis (2)] |
| 8. | Nameirakpam J, Rikhi R, Rawat SS, Sharma J, Suri D. Genetics on early onset inflammatory bowel disease: An update. Genes Dis. 2020;7:93-106. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 30] [Article Influence: 5.0] [Reference Citation Analysis (4)] |
| 9. | Mayerle J, Sendler M, Hegyi E, Beyer G, Lerch MM, Sahin-Tóth M. Genetics, Cell Biology, and Pathophysiology of Pancreatitis. Gastroenterology. 2019;156:1951-1968.e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 320] [Cited by in RCA: 285] [Article Influence: 40.7] [Reference Citation Analysis (1)] |
| 10. | Ye Z, Huang Y, Zheng C, Wang Y, Lu J, Wang H, Wu B, Wang X, Zhang R, Wang J. Clinical and genetic spectrum of children with congenital diarrhea and enteropathy in China. Genet Med. 2019;21:2224-2230. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 11. | Fanen P, Wohlhuter-Haddad A, Hinzpeter A. Genetics of cystic fibrosis: CFTR mutation classifications toward genotype-based CF therapies. Int J Biochem Cell Biol. 2014;52:94-102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 84] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 12. | Andreassen BU, Aunsholt L, Østergaard E, Ek J, Maroun LL, Jørgensen MH. Microvillus Inclusion Disease Caused by MYO5B: Different Presentation and Phenotypes Despite Same Mutation. JPGN Rep. 2023;4:e309. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 13. | Fathalla BM, Alsarhan A, Afzal S, El Naofal M, Abou Tayoun A. The genomic landscape of pediatric rheumatology disorders in the Middle East. Hum Mutat. 2021;42:e1-e14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 13] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 14. | Ramaswamy S, Jain R, El Naofal M, Halabi N, Yaslam S, Taylor A, Tayoun AA. Middle Eastern Genetic Variation Improves Clinical Annotation of the Human Genome. J Pers Med. 2022;12:423. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 8] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 15. | Balciuniene J, DeChene ET, Akgumus G, Romasko EJ, Cao K, Dubbs HA, Mulchandani S, Spinner NB, Conlin LK, Marsh ED, Goldberg E, Helbig I, Sarmady M, Abou Tayoun A. Use of a Dynamic Genetic Testing Approach for Childhood-Onset Epilepsy. JAMA Netw Open. 2019;2:e192129. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 57] [Cited by in RCA: 51] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 16. | Tayoun AN, Mason-Suares H, Frisella AL, Bowser M, Duffy E, Mahanta L, Funke B, Rehm HL, Amr SS. Targeted Droplet-Digital PCR as a Tool for Novel Deletion Discovery at the DFNB1 Locus. Hum Mutat. 2016;37:119-126. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 40] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 17. | Amr SS, Murphy E, Duffy E, Niazi R, Balciuniene J, Luo M, Rehm HL, Abou Tayoun AN. Allele-Specific Droplet Digital PCR Combined with a Next-Generation Sequencing-Based Algorithm for Diagnostic Copy Number Analysis in Genes with High Homology: Proof of Concept Using Stereocilin. Clin Chem. 2018;64:705-714. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 26] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 18. | Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL; ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405-424. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27361] [Cited by in RCA: 25496] [Article Influence: 2317.8] [Reference Citation Analysis (10)] |
| 19. | Riggs ER, Andersen EF, Cherry AM, Kantarci S, Kearney H, Patel A, Raca G, Ritter DI, South ST, Thorland EC, Pineda-Alvarez D, Aradhya S, Martin CL. Technical standards for the interpretation and reporting of constitutional copy-number variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics (ACMG) and the Clinical Genome Resource (ClinGen). Genet Med. 2020;22:245-257. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1362] [Cited by in RCA: 1187] [Article Influence: 197.8] [Reference Citation Analysis (4)] |
| 20. | Abou Tayoun AN, Pesaran T, DiStefano MT, Oza A, Rehm HL, Biesecker LG, Harrison SM; ClinGen Sequence Variant Interpretation Working Group (ClinGen SVI). Recommendations for interpreting the loss of function PVS1 ACMG/AMP variant criterion. Hum Mutat. 2018;39:1517-1524. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 751] [Cited by in RCA: 684] [Article Influence: 85.5] [Reference Citation Analysis (1)] |
| 21. | Fretzayas A, Moustaki M, Liapi O, Karpathios T. Gilbert syndrome. Eur J Pediatr. 2012;171:11-15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 73] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 22. | van der Veere CN, Sinaasappel M, McDonagh AF, Rosenthal P, Labrune P, Odièvre M, Fevery J, Otte JB, McClean P, Bürk G, Masakowski V, Sperl W, Mowat AP, Vergani GM, Heller K, Wilson JP, Shepherd R, Jansen PL. Current therapy for Crigler-Najjar syndrome type 1: report of a world registry. Hepatology. 1996;24:311-315. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 99] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 23. | D'Antiga L, Beuers U, Ronzitti G, Brunetti-Pierri N, Baumann U, Di Giorgio A, Aronson S, Hubert A, Romano R, Junge N, Bosma P, Bortolussi G, Muro AF, Soumoudronga RF, Veron P, Collaud F, Knuchel-Legendre N, Labrune P, Mingozzi F. Gene Therapy in Patients with the Crigler-Najjar Syndrome. N Engl J Med. 2023;389:620-631. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 55] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 24. | Paulusma CC, Elferink RP, Jansen PL. Progressive familial intrahepatic cholestasis type 1. Semin Liver Dis. 2010;30:117-124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 33] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 25. | Mitchell E, Gilbert M, Loomes KM. Alagille Syndrome. Clin Liver Dis. 2018;22:625-641. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 133] [Cited by in RCA: 107] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 26. | Murase N, Kaneko K, Hama A, Yoshida N, Sakaguchi H, Chiba K, Oshiro M. Hepatoblastoma associated with trisomy 18. J Pediatr Surg Case Rep. 2020;52:101342. [RCA] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 27. | Cheung TP, Van Rostenberghe H, Ismail R, Nawawi NN, Abdullah NA, Ramli N, Ibrahim NR, Hj Abd Majid N, Mohd Yusoff N, Nishio H, Yusoff S. High resolution melting analysis of the NR1I3 genetic variants: Is there an association with neonatal hyperbilirubinemia? Gene. 2015;573:198-204. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 28. | Jardine S, Dhingani N, Muise AM. TTC7A: Steward of Intestinal Health. Cell Mol Gastroenterol Hepatol. 2019;7:555-570. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 72] [Cited by in RCA: 63] [Article Influence: 7.9] [Reference Citation Analysis (5)] |
| 29. | Bhattacharya S, Marciano B, Malech H, Holland S, De Ravin SS, Zerbe C, Heller T. Ustekinumab for chronic granulomatous disease-associated inflammatory bowel disease. Gastroenterol. 2021;160:S80-S81. |
| 30. | Slatter MA, Gennery AR. Haematopoietic Stem Cell Transplantation for Chronic Granulomatous Disease. J Clin Med. 2023;12:6083. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 31. | Panchoo AV, VanNess GH, Rivera-Rivera E, Laborda TJ. Hereditary pancreatitis: An updated review in pediatrics. World J Clin Pediatr. 2022;11:27-37. [PubMed] [DOI] [Full Text] |
| 32. | Devane J, Ott E, Olinger EG, Epting D, Decker E, Friedrich A, Bachmann N, Renschler G, Eisenberger T, Briem-Richter A, Grabhorn EF, Powell L, Wilson IJ, Rice SJ, Miles CG, Wood K; Genomics England Research Consortium, Trivedi P, Hirschfield G, Pietrobattista A, Wohler E, Mezina A, Sobreira N, Agolini E, Maggiore G, Dahmer-Heath M, Yilmaz A, Boerries M, Metzger P, Schell C, Grünewald I, Konrad M, König J, Schlevogt B, Sayer JA, Bergmann C. Progressive liver, kidney, and heart degeneration in children and adults affected by TULP3 mutations. Am J Hum Genet. 2022;109:928-943. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 41] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 33. | Tudurachi BS, Zăvoi A, Leonte A, Țăpoi L, Ureche C, Bîrgoan SG, Chiuariu T, Anghel L, Radu R, Sascău RA, Stătescu C. An Update on MYBPC3 Gene Mutation in Hypertrophic Cardiomyopathy. Int J Mol Sci. 2023;24:10510. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 56] [Article Influence: 18.7] [Reference Citation Analysis (0)] |
| 34. | Ruemmele FM, Schmitz J, Goulet O. Microvillous inclusion disease (microvillous atrophy). Orphanet J Rare Dis. 2006;1:22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 121] [Cited by in RCA: 103] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 35. | Kalashyan M, Raghunathan K, Oller H, Theres MB, Jimenez L, Roland JT, Kolobova E, Hagen SJ, Goldsmith JD, Shub MD, Goldenring JR, Kaji I, Thiagarajah JR. Therapy Development for Microvillus Inclusion Disease using Patient-derived Enteroids. bioRxiv. 2023;2023.01.28.526036. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 36. | Ashworth I, Wilson A, Aquilina S, Parascandalo R, Mercieca V, Gerada J, Macdonald S, Simchowitz V, Hill S. Reversal of Intestinal Failure in Children With Tufting Enteropathy Supported With Parenteral Nutrition at Home. J Pediatr Gastroenterol Nutr. 2018;66:967-971. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 11] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 37. | Sarkozy A, Carta C, Moretti S, Zampino G, Digilio MC, Pantaleoni F, Scioletti AP, Esposito G, Cordeddu V, Lepri F, Petrangeli V, Dentici ML, Mancini GM, Selicorni A, Rossi C, Mazzanti L, Marino B, Ferrero GB, Silengo MC, Memo L, Stanzial F, Faravelli F, Stuppia L, Puxeddu E, Gelb BD, Dallapiccola B, Tartaglia M. Germline BRAF mutations in Noonan, LEOPARD, and cardiofaciocutaneous syndromes: molecular diversity and associated phenotypic spectrum. Hum Mutat. 2009;30:695-702. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 264] [Cited by in RCA: 224] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 38. | Bosa L, Batura V, Colavito D, Fiedler K, Gaio P, Guo C, Li Q, Marzollo A, Mescoli C, Nambu R, Pan J, Perilongo G, Warner N, Zhang S, Kotlarz D, Klein C, Snapper SB, Walters TD, Leon A, Griffiths AM, Cananzi M, Muise AM. Novel CARMIL2 loss-of-function variants are associated with pediatric inflammatory bowel disease. Sci Rep. 2021;11:5945. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (4)] |
| 39. | Leggett BA, Young JP, Barker M. Peutz-Jeghers syndrome: genetic screening. Expert Rev Anticancer Ther. 2003;3:518-524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 8] [Article Influence: 0.3] [Reference Citation Analysis (0)] |