Published online Mar 14, 2026. doi: 10.3748/wjg.v32.i10.116128
Revised: December 6, 2025
Accepted: January 14, 2026
Published online: March 14, 2026
Processing time: 117 Days and 13.8 Hours
The presence of human leukocyte antigen (HLA)-DQ2 and/or HLA-DQ8 alleles is necessary but not sufficient for the development of celiac disease (CeD). This suggests that additional environmental and biological factors, including bacteria and, above all, the still rarely studied fungal gut microbiota, play key roles in disease onset and progression.
To characterize and compare the intestinal bacteriobiota and mycobiota profiles of children with newly diagnosed CeD and their unaffected siblings, in comparison with a healthy control group.
The study included children and adolescents aged 1 to 18 years. Participants were divided into three groups: (1) 14 patients with newly diagnosed CeD; (2) 16 asymptomatic siblings of CeD patients; and (3) 19 healthy children (control group). Stool samples were collected from all eligible participants. Next-generation sequencing was performed, followed by analysis of the relationship between the gut microbiota and genetic predisposition to CeD, with attention to the HLA DQ2/8 alleles.
Regarding alpha diversity, the CeD and sibling groups differed significantly from the control group (bacteria), and the CeD group differed from siblings (fungi). Significant dissimilarities in beta diversity were observed between siblings and both CeD and control groups. In comparisons between CeD group and their siblings, 13 indicator bacterial species were identified, whereas in comparisons between the CeD group and their siblings and controls, 8 indicator fungal species were detected. No significant correlation was found between bacterial species and the presence of the HLA DQ2.5 allele, or between fungal species and HLA DQ2.2. A strong (r = 0.8-0.9) positive relationship was found between Subdoligranulum variabile and several bacterial species. A moderate (r = 0.4-0.7) positive correlation was observed between the fungal species Microidium phyllanthi and Bifidobacterium longum, Clostridium leptum and Romboutsia timonensis.
While DQ2.5 plays a central role in disease pathogenesis, it appears to have less direct influence on microbial composition. The distinct fungal signatures observed in siblings may serve as early indicators of risk and warrant further investigation.
Core Tip: This is the first comprehensive assessment of the bacteriobiome and mycobiome in children with celiac disease (CeD) and their first-degree relatives (siblings). Our findings highlight the potential of both bacterial and fungal signatures as early biomarkers of CeD susceptibility and suggest that host human leukocyte antigen genotypes, particularly DQ8 and DQ2.2, may shape the gut microbial landscape in children at risk for CeD, through both bacterial and fungal communities.
- Citation: Salamon D, Krawczyk A, Zapała B, Duplaga M, Kowalska-Duplaga K, Gosiewski T. Gut bacterial and fungal signatures in relation to human leukocyte antigen-DQ2/DQ8 in children with celiac disease and siblings. World J Gastroenterol 2026; 32(10): 116128
- URL: https://www.wjgnet.com/1007-9327/full/v32/i10/116128.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i10.116128
The presence of human leukocyte antigen (HLA)-DQ2 and/or HLA-DQ8 alleles is essential but not sufficient on its own to cause celiac disease (CeD). These alleles are found in approximately 30%-40% of the general population, yet only around 1% of individuals develop the disease. This suggests that additional environmental and biological factors, including the gut microbiota, play key roles in disease onset and progression[1-3].
Dysbiosis, or disruption in the composition or function of the microbiota, comprising bacteria (bacteriobiota), and fungi (mycobiota), has long been observed in CeD patients, including those who have not yet introduced gluten into their diet, suggesting its potential role in disease initiation[4]. Several studies have reported that children with CeD exhibit reduced microbial diversity and alterations in the abundance of specific bacterial taxa, such as increased Proteobacteria and disrupted Firmicutes[5]. In addition, although less well understood, the fungal component of the gut microbiome has also been implicated in CeD (for instance, an increased presence of opportunistic fungi, particularly Candida albicans) and may contribute to mucosal inflammation and immune activation[6]. Importantly, the interaction between host genetics and the gut microbiota is bidirectional: The HLA genotype can influence microbiota composition, while microbial metabolites can, in turn, modulate immune responses in genetically susceptible individuals[7].
This study aimed to characterize and compare the intestinal bacteriobiota and mycobiota profiles of children with newly diagnosed CeD and their unaffected siblings with those of a healthy control group. We also sought to explore correlations between the composition of the microbiota (both bacterial and fungal) and the presence of HLA-DQ2/DQ8 alleles. To our knowledge, this is the first study to jointly analyze these components in this population.
All procedures involving human participants were approved by the Jagiellonian University Bioethics Committee in Krakow, Poland, No. 1072.6120.82.2018. Informed consent was obtained from the patients’ parents or legal guardians (for all patients under 18 years of age) and, additionally, from the patients themselves if they were over 16 years old.
The study included children and adolescents aged 1 years to 18 years and was conducted in the Department of Pediatrics, Gastroenterology and Nutrition, University Children’s Hospital in Krakow, Poland, between 2019 and 2022. This research followed a case-control study design. Participants were divided into three groups. The first one comprised 14 patients with newly diagnosed CeD, meeting the diagnostic criteria established by the European Society for Paediatric Gastroenterology, Hepatology, and Nutrition in 2012[8]. During the implementation of the project, new diagnostic criteria were published; however, this did not affect participant recruitment or study execution. The second group included 16 asymptomatic siblings of the CeD patients. The control group consisted of 19 healthy children in whom CeD and other gastrointestinal diseases were ruled out. Exclusion criteria are presented in the Supplementary material.
Stool samples were collected from all eligible participants and immediately frozen at -80 °C. The complete set of samples was transported under deep-freeze conditions to the Department of Molecular Medical Microbiology, Division of Microbiology, Jagiellonian University Medical College in Krakow where library preparation and sequencing were subsequently performed, ensuring that the relevant analyses could be performed simultaneously on all samples. A detailed description of the following steps and additional materials, and samples is provided in the Supplementary material.
The stool samples were thawed, and 150 mg from each sample was taken for the extraction of fungal and bacterial DNA using the Genomic Mini AX Stool Kit (A and A Biotechnology, Gdańsk, Poland), following our previously published modification, which includes enzymatic lysis with lysozyme, lyticase, mutanolysin, or lysostaphin, as well as mechanical lysis using a homogenizer[9,10]. The extracted DNA was quantified using a NanoDrop spectrophotometer (Thermo Fisher, Waltham, MA, United States) by measuring absorbance at 260 nm and the A260/A280 ratio to assess DNA concentration and purity.
The isolated DNA was subjected to polymerase chain reaction amplification using a T100 Thermal Cycler (BioRad, CA, United States) with primers targeting the 16S for bacteria and internal transcribed spacer (ITS)-1 regions of the fungal rDNA gene to generate ITS libraries. The primer sequences (based on Strati et al[11]) are presented in Supplementary Table 1.
Libraries were prepared following the protocol for Preparing 16S Ribosomal RNA Gene Amplicons for the Illumina MiSeq System (https://support.illumina.com/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf, accessed August 2, 2025). The concentration of the genomic library was determined using the PicoGreen fluorescent dye (Thermo Fisher, Waltham, MA, United States) and combined with a 30% spike-in of PhiX control DNA (Illumina, San Diego, CA, United States). The pooled libraries were then loaded onto a Reagent Kit V3 cartridge (600 cycles; Illumina, San Diego, CA, United States) and sequenced on a MiSeq platform (Illumina, San Diego, CA, United States).
Raw sequencing reads in FASTQ format were processed using the Illumina 16S metagenomics workflow. Taxonomic classification was performed using a high-performance implementation of the ribosomal database project naive Bayesian classifier as described by Wang et al[12]. Open-reference operational taxonomic units (OTUs) were clustered at 97% similarity and preliminarily assigned taxonomy using the Greengenes reference database (version 13.5)[13] to ensure comparability with previous studies based on QIIME workflows (compatible with this database). The 16S rRNA and ITS sequences of microbial genomes were base-called, quality-filtered, and demultiplexed. Following quality filtering and demultiplexing, only high-quality reads (Phred quality ≥ 30) were assigned to biological samples, corresponding to an average of reads per sample. Then genus- and species-level refinement was conducted using the Genome Taxonomy Database[14] for bacterial taxa and the UNITE database[15] for fungal taxa.
Taxonomic abundance tables were generated at multiple taxonomic levels, phylum (L2), class (L3), order (L4), family (L5), genus (L6), and species (L7) using normalized read counts. Relative abundance profiles were filtered to exclude taxa below the prevalence and abundance thresholds to minimize noise. Data integration and the next analysis were performed using the R software package (version 4.5.1).
Post-hoc power analysis was performed using Python (version 3.11) and the statsmodels package (version 0.14.0; https://www.statsmodels.org). A detailed description is provided in the Supplementary material. The descriptive statistics were carried out with IBM SPSS Statistics 28 and were calculated for all variables. Normality of data distribution was assessed using the Shapiro-Wilk test for each variable within the three study groups (CeD, siblings, controls). Variables with normal distribution are presented as mean ± SD, while non-normally distributed variables are presented as median and quartile 1 and quartile 3. Multivariate analysis was performed to determine the statistically significant differences in the means when comparing the three groups’ results. The χ2 and Fisher’s tests were used to analyze the variables on the nominal scale. The Kruskal-Walli’s test was used for quantitative variables, and a post-hoc test (Mann-Whitney U test) was used for each pair of groups with Bonferroni correction to account for multiple comparisons. After Bonferroni correction, the significance level was set at α = 0.017 for three pairwise comparisons. Bioinformatic analyses are detailed in the Supplementary material.
Because the Shapiro-Wilk test indicated that several quantitative variables did not meet the assumption of normality, non-parametric tests were applied for all comparisons between the three study groups. Consequently, all variables in the summary Table 1 are presented as median and quartile 1 and quartile 3 to ensure consistency with the statistical approach. Table 2 present the clinical data (sex and three HLA alleles) for children from all three study groups.
| Variable | CeD, n = 14 | Siblings of CeD, n = 16 | Control group, n = 19 | P value1 | P value1,2 | P value1,3 | P value1,4 |
| BMI (kg/m2) | 15.28 (14.17-16.62) | 16.81 (4.62-18.12) | 19.51 (5.29-21.61) | 0.026a | 0.35 | 0.021a | 0.8 |
| Glycemia (mmol/L) | 4.5 (4.43-4.78) | 4.45 (4.32-4.68) | 4.4 (4.25-4.7) | 0.936 | |||
| Total cholesterol (mmol/L) | 3.95 (3.69-4.4) | 4.42 (3.83-4.79) | 3.83 (3.67-4.21) | 0.005a | 0.24 | 0.65 | 0.004a |
| HDL (mmol/L) | 1.35 (1.24-1.56) | 1.48 (1.32-1.7) | 1.26 (1.1-1.42) | 0.018a | 0.7 | 0.41 | 0.014a |
| LDL (mmol/L) | 2.47 (2.0-2.54) | 2.72 (2.22-2.97) | 2.36 (1.92-2.55) | 0.019a | 0.12 | 1 | 0.017a |
| TGs (mmol/L) | 0.67 (0.58-0.88) | 0.8 (0.72-0.96) | 0.71 (0.66-0.8) | 0.548 | |||
| ALT (U/L) | 24.0 (19.0-25.12) | 23.5 (16.0-26.8) | 15.0 (13.75-27.5) | 0.83 | |||
| Fe (μmol/L) | 12.9 (10.4-19.7) | 17.8 (13.1-21.05) | 16.35 (14.4-21.25) | 0.34 | |||
| Creatinine (μmol/L) | 34.45 (30.72-36.95) | 35.4 (30.55-44.92) | 51.55 (38.92-55.6) | 0.003a | 1 | 0.013a | 0.011a |
| Anti-tTG (U/mL) | 173.9 (77.7-200.0) | 2.0 (2.0-2.03) | 1.9 (1.9-1.9) | < 0.001a | < 0.001a | < 0.001a | < 0.001a |
| Variable | Category of variable | CeD, n = 14 | Siblings of CeD, n = 16 | Control group, n = 19 | P value1 | P value1,2 | P value1,3 | P value1,4 |
| Sex | Female | 7 | 7 | 16 | 0.033a | 0.752 | 0.045a | 0.045a |
| Male | 7 | 9 | 3 | |||||
| DQ2.2 | Positive | 4 (28.6) | 4 (25) | 3 (15.8) | 0.655 | |||
| Negative | 10 (71.4) | 12 (75) | 16 (84.2) | |||||
| DQ2.5 | Positive | 14 (100) | 8 (50) | 3 (15.8) | < 0.001a | 0.031a | < 0.05 | 0.071 |
| Negative | 0 (0.0) | 8 (50) | 16 (84.2) | |||||
| DQ8 | Positive | 1 (7.1) | 1 (6.3) | 6 (31.6) | 0.071 | |||
| Negative | 13 (92.9) | 15 (93.8) | 13 (68.4) |
All CeD patients (n = 14, 100%) carried the DQ2.5 allele, consistent with disease pathogenesis. The prevalence of DQ2.2 and DQ8 was 28.6% and 7.1%, respectively. In siblings, DQ2.5 was present in 50%, DQ2.2 in 25%, and DQ8 in 6.3%. Controls showed 15.8% for DQ2.2 and DQ2.5, and 31.6% for DQ8. Notably, no CeD patient was negative for all alleles, whereas 31.2% (n = 5) of siblings and 52.6% (n = 10) of controls lacked DQ2.2, DQ2.5, and DQ8.The overall power for detecting group differences via ANOVA was 0.086 for small effects (f = 0.10), 0.309 for medium effects (f = 0.25), and 0.678 for large effects (f = 0.40). For very large effects (f = 0.50), power reached 86.8%, indicating adequate sensitivity for strong differences. For alpha diversity, power was 0.678 for large effects (f = 0.40) and 0.868 for very large effects (f = 0.50). For beta diversity, assuming typical microbiome effect sizes (R2 = 0.15, F = 0.4 and R² = 0.20, F = 0.5), power was 0.723 and 0.868, respectively. However, post-hoc analyses for individual parameters revealed high statistical power for anti-tissue transglutaminase (anti-tTG) (η2 = 0.25, power = 0.95), total cholesterol (η2 = 0.25, power = 0.95) and creatinine (η2 = 0.26, power = 0.96), and low-density lipoprotein cholesterol (η2 = 0.17, power = 0.80). Moderate power was observed for body mass index (η2 = 0.16, power = 0.75) and high-density lipoprotein cholesterol (η2 = 0.15, power = 0.73), while glycemia (η2 = 0.019, power = 0.12), triglycerides (η2 = 0.004, power = 0.06), alanine aminotransferase (η2 = 0.028, power = 0.16), and iron (η2 = 0.044, power = 0.24) showed low power (< 0.30).
The χ2 test of independence revealed varying levels of association and statistical power across the analyzed variables. The strongest effect was observed for DQ2.5 (χ2 = 22.884, P < 0.001), with a large effect size (Cramér’s V = 0.683) and near-perfect power (approximately 0.998). Sex differences were statistically significant (χ2 = 7.030, P = 0.030) with moderate effect size (Cramér’s V = 0.379) and power of 0.755. DQ8 showed an intermediate effect (V = 0.329) and moderate power (0.633), while DQ2.2 exhibited a weak effect (Cramér’s V = 0.131) and low power (0.151). These results indicate that strong associations were reliably detected despite the limited sample size.
Alpha and beta diversity assessments were performed at individual taxonomic levels from L2 to L7. However, as only two phyla, Ascomycota and Basidiomycota, were identified at L2, the analysis of mycobiota diversity began at a more taxonomically diverse level, L3. Alpha diversity analysis of bacterial profiles revealed no statistically significant differences (P > 0.05) in sample diversity between the CeD and sibling groups. However, significant differences were observed between CeD children and the control group at both L2 and L7, and at the species level (L7) between siblings and controls. In contrast, mycobiota diversity comparisons between CeD and control groups, and between siblings and controls, showed no significant differences. Notably, when comparing fungal profiles between the CeD and sibling groups, significant differences were found at both L3 and L7 across four diversity indices (Table 3). Beta diversity analysis showed no significant differences between CeD and control groups in either bacterial or fungal profiles (P > 0.05). However, significant dissimilarities were detected in the bacterial profile at L7 (for two indices), and in the fungal profile at both L3 and L7 (for all three indices), when comparing siblings and both CeD and control groups (Table 4).
| Index | Observed | Chao1 | ACE | Shannon | Simpson | Fisher |
| Taxa level1 | Bacteria | |||||
| L2 | 0.52 | 0.52 | 0.67 | 0.211 | 0.121 | 0.532 |
| L7 | 0.139 | 0.204 | 0.22 | 0.699 | 0.63 | 0.551 |
| Taxa level1 | Fungi | |||||
| L3 | < 0.001a | < 0.001a | 0.002a | 0.191 | 0.232 | 0.03a |
| L7 | < 0.001a | < 0.001a | 0.001a | 0.195 | 0.324 | 0.018a |
| Taxa level2 | Bacteria | |||||
| L2 | < 0.001a | < 0.001a | 0.087 | 0.015a | 0.009a | 0.446 |
| L7 | < 0.001a | < 0.001a | < 0.001a | 0.119 | 0.179 | 0.46 |
| Taxa level2 | Fungi | |||||
| L3 | 0.165 | 0.023a | 0.129 | 0.768 | 0.48 | 0.408 |
| L7 | 0.058 | 0.009a | 0.021a | 0.988 | 0.61 | 0.134 |
| Taxa level3 | Bacteria | |||||
| L2 | 0.001a | 0.003a | 0.143 | 0.193 | 0.209 | 0.308 |
| L7 | 0.002a | 0.001a | 0.001a | 0.035a | 0.067 | 0.324 |
| Taxa level3 | Fungi | |||||
| L3 | 0.208 | 0.289 | 0.056 | 0.095 | 0.039a | 0.269 |
| L7 | 0.353 | 0.5 | 0.636 | 0.173 | 0.116 | 0.369 |
| Index | Bray-Curtis | Jensen-Shannon divergence | Jaccard |
| Taxa level1 | Bacteria | ||
| L2 | 0.238 | 0.05a | 3.79 |
| L7 | 0.045a | 0.052 | 1.28 |
| Taxa level1 | Fungi | ||
| L3 | 0.007a | 0.008a | 0.008a |
| L7 | 0.031a | 0.029a | 0.029a |
| Taxa level2 | Bacteria | ||
| L2 | 0.284 | 0.028a | 0.137 |
| L7 | 0.054 | 0.057 | 0.049a |
| Taxa level2 | Fungi | ||
| L3 | 0.851 | 0.889 | 0.805 |
| L7 | 0.732 | 0.662 | 0.681 |
| Taxa level3 | Bacteria | ||
| L2 | 0.664 | < 0.001a | 0.582 |
| L7 | 0.038a | 0.042a | 0.037a |
| Taxa level3 | Fungi | ||
| L3 | 0.007a | 0.004a | 0.012a |
| L7 | 0.02a | 0.005a | 0.44 |
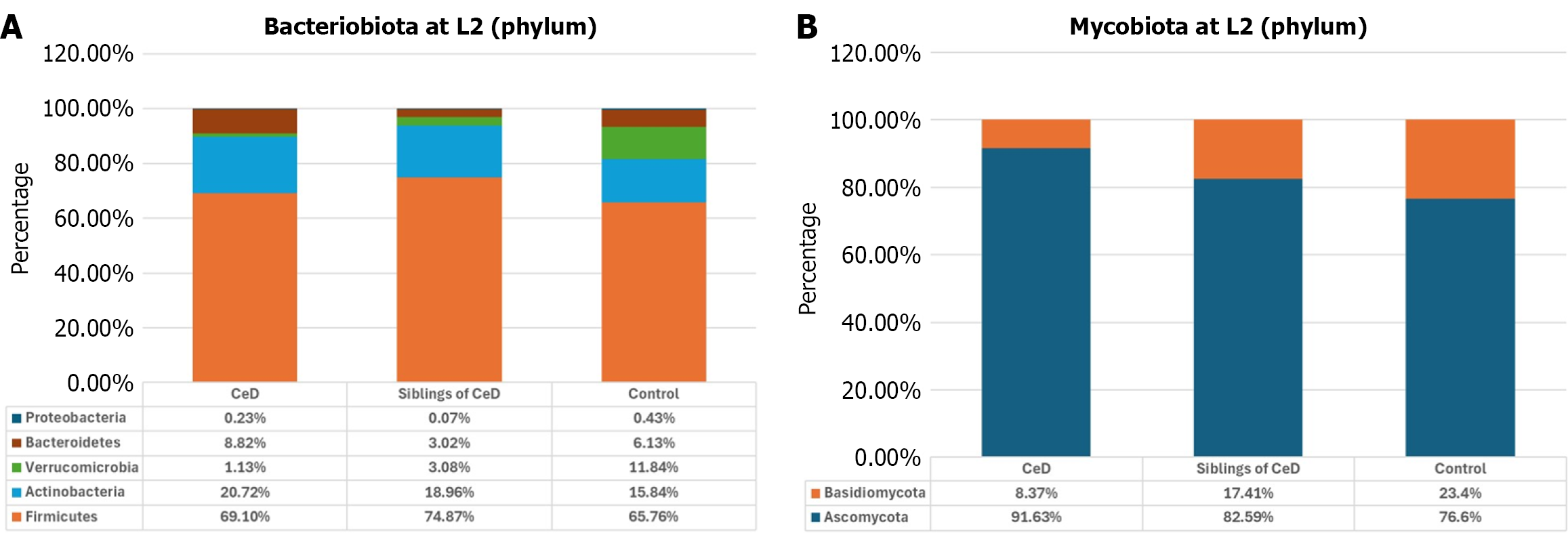
We conducted a systematic assessment of the bacterial and fungal profile of the fecal samples at the L2 taxonomic level to provide an overall picture. Detailed evaluations at the L6 and L7 taxonomic levels were carried out. Due to the large number of significant taxa at L7 (56 bacterial species and 36 fungal species) and the fact that 97% of reads were classified at the species level, the description in this article refers only to selected, statistically significant findings. Taxa with percentages with no significant differences between the groups compared and/or lower than 1% were collectively categorized as “other”. Detailed data on the relative percentages of all bacteria and fungi identified at the L2, L6, and L7 levels, as well as statistically significant differences between the analyzed groups, are presented in Figure 1, Supplementary Figures 1 and 2, and Supplementary Tables 2-7.
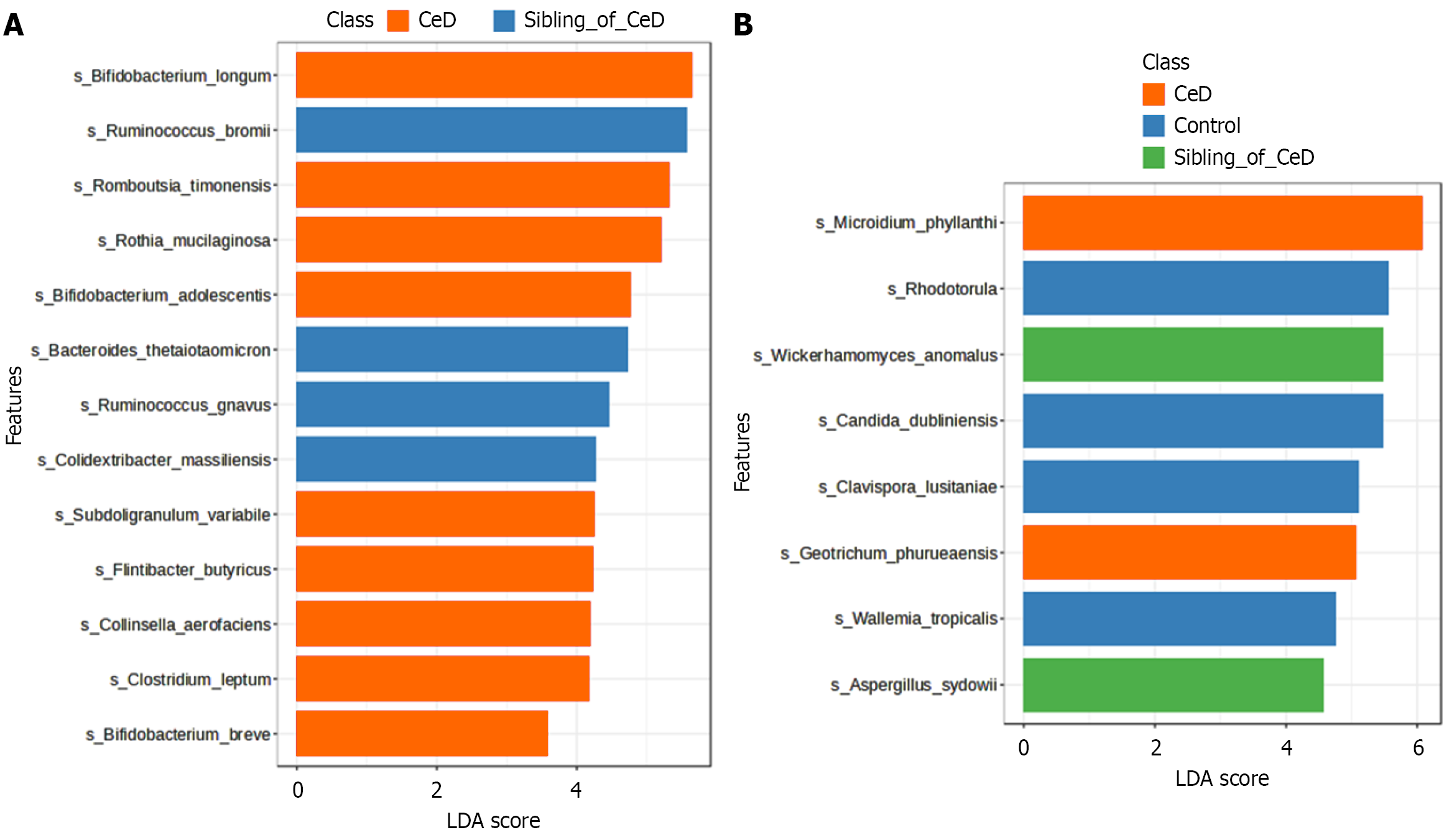
Firmicutes were the dominant bacterial phylum, and Ascomycota was the dominant fungal phylum in all studied groups. At the L7, we identified 19 OTUs which corresponded to bacterial genera and 31 OTUs which corresponded to fungal genera and at the same time exhibited statistically significant (P < 0.05) differences in relative percentages between the studied groups. The linear discriminator analysis revealed statistical significance in the biomarker’s discovery. In the comparisons of children with CeD group vs their siblings, we presented the top 13 indicator bacteria species, whereas in the comparison of groups: CeD children vs their siblings and vs control we showed 8 indicator fungi species (Figure 2).
Analyzing the association of selected bacterial species with the presence of a given HLA DQ allele, we found a moderate (r = 0.4-0.7) negative correlation between the HLA DQ8 allele and Bifidobacterium longum (adjusted P = 0.01) and Romboutsia timonensis (adjusted P = 0.01) and between the HLA DQ 2.2 allele and Subdoligranulum variabile (adjusted P = 0.004). No significant correlation was found between the selected bacterial species and the presence of the HLA DQ 2.5 allele (Supplementary Figure 3). Analysis of the occurrence of the HLA DQ2/DQ8 allele in relation to selected fungal species showed a moderate (r = 0.4-0.7) positive correlation between HLA DQ 2.5 and Microidium phyllanthi (in children with CeD; adjusted P = 0.03 and their siblings; adjusted P = 0.01). No significant correlation was found between the selected fungal species and the presence of the HLA DQ2.2 allele (Supplementary Figure 4).
A relationship between selected bacterial species and fungal species is presented in Supplementary Figure 5. A strong (r = 0.7-0.9) positive relationship was found between Subdoligranulum variabile and other bacterial species (Bacteroides thetaiotaomicron, adjusted P = 0.02; Bifidobacterium adolescentis, adjusted P = 0.02, Bifidobacterium breve, adjusted P = 0.03; Bifidobacterium longum, adjusted P = 0.01; Clostridium leptum, adjusted P = 0.01; Collinsella aerofaciens, adjusted P = 0.03; Colidextribacter massiliensis, adjusted P = 0.01; Flintibacter butyricus, adjusted P = 0.02; Ruminococcus gnavus, adjusted P = 0.02; Romboutsia timonensis, adjusted P = 0.01; Rothia mucilaginosa, adjusted P = 0.02; Ruminococcus bromii, adjusted P = 0.003). A moderate (r = 0.4-0.7) positive correlation was noted between Microidium phyllanthi and Bifidobacterium longum (adjusted P = 0.01), Clostridium leptum (adjusted P = 0.02) and Romboutsia timonensis (adjusted P = 0.04). Additionally, a negative (moderate -r = 0.4-0.7 and weak -r = 0.2-0.4) correlation was found between the fungus Wallemia tropicalis and the selected bacterial species (Bacteroides thetaiotaomicron, adjusted P = 0.01; Bifidobacterium adolescentis, adjusted P = 0.02; Bifidobacterium longum, adjusted P = 0.02; Clostridium leptum, adjusted P = 0.02; Collinsella aerofaciens, adjusted P = 0.01; Colidextribacter massiliensis, adjusted P = 0.04; Flintibacter butyricus, adjusted P = 0.01; Ruminococcus gnavus, adjusted P = 0.02; Romboutsia timonensis, adjusted P = 0.03; Ruminococcus bromii, adjusted P = 0.03; Subdoligranulum variabile, adjusted P = 0.01) (Supplementary Figure 5).
This is the first comprehensive assessment of the microbiome in children with CeD and their first-degree relatives (siblings). In our results, all children with CeD carried at least one of the three alleles tested (DQ2.5, DQ2.2 and DQ8), which confirms the strong relationship between the presence of these alleles and the risk of developing CeD. In 57.9% of children from the control group, none of the alleges were found, which is in line with expectations for the general population reflecting real-world biological conditions[16].
The overall sample size was limited, and the estimated power was low for small and medium effects (8.6% and 30.9%) and moderate for large effects (67.8%), which warrants caution in interpreting and generalizing the results. Nevertheless, study was sufficiently powered to identify large differences in microbial diversity, including alpha and beta diversity metrics. This aligns with previous simulation-based studies, which indicate that 12-15 samples per group are adequate for detecting strong microbiome effects[17,18]. Moreover, detailed power analyses for individual quantitative and categorical variables indicated that the strongest associations were detected reliably despite the small cohort. For example, DQ2.5 showed a large effect size (Cramér’s V = 0.683) with near-perfect power (approximately 0.998), and sex differences had moderate effect size (V = 0.379) and power of 0.755. In turn, the analysis revealed substantial variability in statistical power across biochemical variables, ranging from high (e.g., anti-tTG, η2 = 0.4352, power = 0.99) to very low (e.g., glycemia, η2 = 0.019, power = 0.12). Because anti-tTG reflects disease activity and demonstrated the highest statistical power among the quantitative data, it was selected for the final comparative analysis with bacterial and fungal microbiota to minimize the risk of type II error, which occurs when a true effect is missed due to insufficient statistical power. We observed a significant difference in bacterial alpha diversity between CeD patients and their sibling’s vs the control group. These results are aligned with those reported by Zafeiropoulou et al[5], who found reduced microbial diversity in CeD patients compared to healthy controls, with subtler differences between CeD patients and their siblings. Interestingly, we observed significant differences in alpha diversity between children with CeD and their siblings in mycobiota, an area that remains underexplored in literature. In addition, the beta diversity analysis revealed significant dissimilarities between siblings of CeD patients and both other groups. This is particularly noteworthy because all samples were collected before the gluten-free diet was introduced. Therefore, diet and environmental factors, typically considered key microbiome modulators, were likely comparable across groups. These findings challenge the prevailing view that differences in microbiota composition among sibling’s stem primarily from shared diet or household exposure. In contrast to Roque et al[19], who attributed microbiota differences in siblings to family dietary patterns, our results suggest that intrinsic factors, such as host genetics or early microbial colonization, may be associated with the observed differences, without implying direct causation.
Leonard et al[20] reported that children genetically at risk for CeD displayed unique microbiome trajectories before the clinical onset of disease. Our findings extend this observation to siblings, who, despite being asymptomatic, may harbor microbiota configurations associated with risk or resilience. Notably, the observed dissimilarities were driven by both bacterial and fungal taxa, underscoring the need to include mycobiome data in disease-risk profiling, a field still in its infancy. Collectively, these results raise the possibility that early microbial divergence in CeD siblings reflects not just passive exposure to shared environments, but active host-microbe interactions shaped by host genotype or immunological priming.
The application of linear discriminator analysis in our study revealed key microbial signatures potentially linked to early disease processes, familial predisposition, and environmental influences. The study showed an increased abundance of Bifidobacterium in children with CeD, which seems inconsistent with many previous studies. However, such a phenomenon may be the body’s response to inflammation or damage to the intestinal mucosa in CeD. These bacteria are known for their anti-inflammatory properties and may play a role in modulating the immune response[21].
Fungal biomarkers demonstrated distinct group-specific profiles. Children with CeD were enriched in Microdium phyllanthi and Geotrichum phruensis, fungi rarely reported in gut microbiome studies, warranting further investigation. Their presence may be related to altered mucosal immunity or barrier dysfunction. In siblings, enrichment of Candida dubliniensis, Clavispora lusitaniae, Wallemia tropicalis, and Aspergillus sydowii suggests a unique mycobiota signature, potentially reflecting early microbial shifts in genetically predisposed hosts. Candida dubliniensis has previously been associated with dysbiosis and inflammatory conditions. In particular, its presence correlates positively with inflammatory markers in Crohn’s disease, such as faecal calprotectin and neutrophil gelatinase-related lipocalin, suggesting its potential role in the severity of intestinal inflammation[20,22]. In turn, species of the genus Wallemia, including Wallemia tropicalis, were detected in metagenomic analyses of the gut microbiota, especially in conditions of impaired microbial balance. A review on gut dysbiosis and its immunological consequences emphasised that the presence of rare fungi such as Wallemia spp. may be a marker of impaired microbial balance, especially in the context of autoimmune and metabolic diseases[23]. The analysis of our data, which shows a negative correlation between Wallemia tropicalis and anti-inflammatory bacteria, is consistent with the hypothesis that the presence of this fungus may indicate an environment that promotes inflammation and weakens the intestinal barrier. Furthermore, our results are consistent with the observations of El Mouzan et al[24], who showed significant differences in the composition of intestinal fungi between children with CeD and healthy controls, including increased Candida albicans and decreased alpha diversity. Like their study, we also observed signi
After applying the Benjamini-Hochberg correction for multiple comparisons, we observed multiple significant associations between specific HLA DQ alleles and the composition of the gut microbiota. Both species Bifidobacterium longum and Romboutsia timonensis (a moderate negative correlation with presence of HLA DQ8) are considered beneficial members of the gut microbiota. They are implicated with Bifidobacterium longum in immune modulation, intestinal barrier reinforcement, and short-chain fatty acid production, particularly butyrate[25]. The reduced abundance of Romboutsia timonensis with potential anti-inflammatory effects may similarly reflect immunogenetic pressures associated with CeD susceptibility[26]. Furthermore, Subdoligranulum variabile, a known butyrate producer and contributor to gut homeostasis[27], was negatively associated with HLA DQ2.2, suggesting that even alleles less strongly associated with CeD may influence colonization by key microbial species involved in epithelial integrity and immune balance.
Interestingly, no significant associations were detected between bacterial taxa and HLA DQ2.5, despite this allele’s central role in CeD pathogenesis. This may indicate that DQ2.5 contributes more strongly to antigen presentation and T cell activation rather than directly shaping microbial composition[22]. These findings align with previous reports suggesting that HLA genotype can influence early-life microbiota trajectories, though the directionality and mechanisms remain under investigation[7].
In the fungal microbiota, a moderate positive correlation was observed between Microidium phyllanthi and HLA DQ2.5 in both CeD patients and their siblings. Although data on this fungal species are scarce, its consistent association with DQ2.5-positive individuals may point to immunogenetically influenced fungal persistence or proliferation. These findings warrant further exploration of host genetic influences on fungal ecology in the gut, which remains an underexplored aspect of the microbiome.
Bacteria-fungi analysis further revealed positive correlations between Microidium phyllanthi and bacterial taxa such as Bifidobacterium longum and Romboutsia timonensis, suggesting complex ecological interactions. These findings highlight the need for integrated studies combining genetics, microbiota, and immune profiling to elucidate mechanisms underlying CeD susceptibility.
This study has some limitations. First, the overall sample size was relatively small, resulting in low statistical power to detect small to medium effects. This limitation was primarily due to the coronavirus disease 2019 pandemic, which significantly impacted patient recruitment and reduced the number of available samples compared to initial expectations. Nevertheless, for microbiome diversity metrics, power estimates based on assumed large and very large effects indicated that the study was adequately powered to detect strong differences in alpha and beta diversity. Moreover, additional analyses confirmed that key findings, including anti-tTG levels differences, and strong HLA associations, were detected with sufficient power. These results support the reliability of the main conclusions, although weaker associations should be interpreted cautiously.
The exclusive reliance on stool samples, excluding duodenal biopsy material, may also be considered a limitation of our study. However, because all samples were collected before the introduction of a gluten-free diet, environmental and dietary exposures were consistent across groups. This supports the conclusion that microbial differences, particularly in siblings, may reflect intrinsic host-microbe interactions shaped by genetic background. Furthermore, the fact that DQ2.5 is strongly associated with CeD but shows few microbiota correlations, while DQ2.2 and DQ8 display the opposite trend suggests allele-specific differences in how genetics and microbiota interact. This warrants further investigation in larger cohorts.
It should also be mentioned that in our study, neither siblings nor health controls reported gastrointestinal or extraintestinal symptoms suggestive of non-celiac gluten sensitivity (NCGS); therefore, we did not perform diagnostic procedures in this regard. NCGS remains a clinical diagnosis based on symptom response to gluten withdrawal and recurrence during blinded challenge, as outlined in the randomized, double-blind, placebo-controlled trial by Francavilla et al[28]. There are currently no validated biomarkers or standardized screening tests for NCGS, and its evaluation is only indicated in symptomatic individuals after exclusion of CeD and wheat allergy.
In summary, our findings suggest that host HLA genotypes, particularly DQ8 and DQ2.2, may be associated with differences in the abundance of selected bacterial and fungal taxa in children at risk for CeD, rather than broadly shaping overall microbial composition. While DQ2.5 plays a central role in disease pathogenesis, it appears to have less direct influence on microbial composition. This study underscores the value of assessing both bacteriobiota and mycobiota in identifying early microbial markers of CeD susceptibility. The distinct fungal signatures observed in siblings could serve as early indicators of risk and warrant further investigation. Nevertheless, these findings remain preliminary and require validation in larger cohorts with longitudinal follow-up to clarify causal relationships and inform potential microbiome-based strategies for prevention or early intervention.
The authors would like to acknowledge Jolana Kopec and Danuta Rojek-Zakrzewska for assistance in DNA isolation and preparation of DNA libraries for next-generation sequencing.
| 1. | Sollid LM, Jabri B. Triggers and drivers of autoimmunity: lessons from coeliac disease. Nat Rev Immunol. 2013;13:294-302. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 282] [Cited by in RCA: 256] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 2. | Catassi C, Verdu EF, Bai JC, Lionetti E. Coeliac disease. Lancet. 2022;399:2413-2426. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 417] [Cited by in RCA: 318] [Article Influence: 79.5] [Reference Citation Analysis (0)] |
| 3. | Lupu VV, Sasaran MO, Jechel E, Starcea IM, Ioniuc I, Mocanu A, Rosu ST, Munteanu V, Nedelcu AH, Danielescu C, Salaru DL, Knieling A, Lupu A. Celiac disease - a pluripathological model in pediatric practice. Front Immunol. 2024;15:1390755. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 4. | Olivares M, Neef A, Castillejo G, Palma GD, Varea V, Capilla A, Palau F, Nova E, Marcos A, Polanco I, Ribes-Koninckx C, Ortigosa L, Izquierdo L, Sanz Y. The HLA-DQ2 genotype selects for early intestinal microbiota composition in infants at high risk of developing coeliac disease. Gut. 2015;64:406-417. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 253] [Cited by in RCA: 229] [Article Influence: 20.8] [Reference Citation Analysis (4)] |
| 5. | Zafeiropoulou K, Nichols B, Mackinder M, Biskou O, Rizou E, Karanikolou A, Clark C, Buchanan E, Cardigan T, Duncan H, Wands D, Russell J, Hansen R, Russell RK, McGrogan P, Edwards CA, Ijaz UZ, Gerasimidis K. Alterations in Intestinal Microbiota of Children With Celiac Disease at the Time of Diagnosis and on a Gluten-free Diet. Gastroenterology. 2020;159:2039-2051.e20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 60] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 6. | Richard ML, Sokol H. The gut mycobiota: insights into analysis, environmental interactions and role in gastrointestinal diseases. Nat Rev Gastroenterol Hepatol. 2019;16:331-345. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 157] [Cited by in RCA: 171] [Article Influence: 24.4] [Reference Citation Analysis (0)] |
| 7. | Olivares M, Walker AW, Capilla A, Benítez-Páez A, Palau F, Parkhill J, Castillejo G, Sanz Y. Gut microbiota trajectory in early life may predict development of celiac disease. Microbiome. 2018;6:36. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 82] [Cited by in RCA: 110] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 8. | Husby S, Koletzko S, Korponay-Szabó IR, Mearin ML, Phillips A, Shamir R, Troncone R, Giersiepen K, Branski D, Catassi C, Lelgeman M, Mäki M, Ribes-Koninckx C, Ventura A, Zimmer KP; ESPGHAN Working Group on Coeliac Disease Diagnosis; ESPGHAN Gastroenterology Committee; European Society for Pediatric Gastroenterology, Hepatology, and Nutrition. European Society for Pediatric Gastroenterology, Hepatology, and Nutrition guidelines for the diagnosis of coeliac disease. J Pediatr Gastroenterol Nutr. 2012;54:136-160. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2111] [Cited by in RCA: 1859] [Article Influence: 132.8] [Reference Citation Analysis (3)] |
| 9. | Gosiewski T, Szała L, Pietrzyk A, Brzychczy-Włoch M, Heczko PB, Bulanda M. Comparison of methods for isolation of bacterial and fungal DNA from human blood. Curr Microbiol. 2014;68:149-155. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 30] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 10. | Kowalska-Duplaga K, Krawczyk A, Sroka-Oleksiak A, Salamon D, Wędrychowicz A, Fyderek K, Gosiewski T. Dependence of Colonization of the Large Intestine by Candida on the Treatment of Crohn's Disease. Pol J Microbiol. 2019;68:121-126. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 11. | Strati F, Cavalieri D, Albanese D, De Felice C, Donati C, Hayek J, Jousson O, Leoncini S, Pindo M, Renzi D, Rizzetto L, Stefanini I, Calabrò A, De Filippo C. Altered gut microbiota in Rett syndrome. Microbiome. 2016;4:41. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 121] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 12. | Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261-5267. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17598] [Cited by in RCA: 13281] [Article Influence: 699.0] [Reference Citation Analysis (11)] |
| 13. | DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol. 2006;72:5069-5072. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8383] [Cited by in RCA: 7520] [Article Influence: 376.0] [Reference Citation Analysis (9)] |
| 14. | Parks DH, Chuvochina M, Chaumeil PA, Rinke C, Mussig AJ, Hugenholtz P. A complete domain-to-species taxonomy for Bacteria and Archaea. Nat Biotechnol. 2020;38:1079-1086. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 488] [Cited by in RCA: 945] [Article Influence: 157.5] [Reference Citation Analysis (0)] |
| 15. | Nilsson RH, Larsson KH, Taylor AFS, Bengtsson-Palme J, Jeppesen TS, Schigel D, Kennedy P, Picard K, Glöckner FO, Tedersoo L, Saar I, Kõljalg U, Abarenkov K. The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 2019;47:D259-D264. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1675] [Cited by in RCA: 1825] [Article Influence: 260.7] [Reference Citation Analysis (0)] |
| 16. | Cecilio LA, Bonatto MW. The prevalence of HLA DQ2 and DQ8 in patients with celiac disease, in family and in general population. Arq Bras Cir Dig. 2015;28:183-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 31] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 17. | Anderson MJ. Permutational Multivariate Analysis of Variance (PERMANOVA). In: Wiley StatsRef: Statistics Reference Online. NJ, United States: Willey, 2017. [DOI] [Full Text] |
| 18. | Kelly BJ, Gross R, Bittinger K, Sherrill-Mix S, Lewis JD, Collman RG, Bushman FD, Li H. Power and sample-size estimation for microbiome studies using pairwise distances and PERMANOVA. Bioinformatics. 2015;31:2461-2468. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 333] [Article Influence: 30.3] [Reference Citation Analysis (0)] |
| 19. | Roque A, Zanker J, Brígido S, Tomaz MB, Gonçalves A, Barbeiro S, Benítez-Páez A, Pereira SG. Dietary patterns drive loss of fiber-foraging species in the celiac disease patients gut microbiota compared to first-degree relatives. Gut Pathog. 2024;16:58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 20. | Leonard MM, Valitutti F, Karathia H, Pujolassos M, Kenyon V, Fanelli B, Troisi J, Subramanian P, Camhi S, Colucci A, Serena G, Cucchiara S, Trovato CM, Malamisura B, Francavilla R, Elli L, Hasan NA, Zomorrodi AR, Colwell R, Fasano A; CD-GEMM Team. Microbiome signatures of progression toward celiac disease onset in at-risk children in a longitudinal prospective cohort study. Proc Natl Acad Sci U S A. 2021;118:e2020322118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 135] [Cited by in RCA: 117] [Article Influence: 23.4] [Reference Citation Analysis (11)] |
| 21. | Olshan KL, Leonard MM, Serena G, Zomorrodi AR, Fasano A. Gut microbiota in Celiac Disease: microbes, metabolites, pathways and therapeutics. Expert Rev Clin Immunol. 2020;16:1075-1092. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (1)] |
| 22. | Iversen R, Sollid LM. The Immunobiology and Pathogenesis of Celiac Disease. Annu Rev Pathol. 2023;18:47-70. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 167] [Cited by in RCA: 138] [Article Influence: 46.0] [Reference Citation Analysis (0)] |
| 23. | Zajc J, Gunde-Cimerman N. The Genus Wallemia-From Contamination of Food to Health Threat. Microorganisms. 2018;6:46. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 60] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 24. | El Mouzan M, Al-Hussaini A, Fanelli B, Assiri A, AlSaleem B, Al Mofarreh M, Al Sarkhy A, Alasmi M. Fungal Dysbiosis in Children with Celiac Disease. Dig Dis Sci. 2022;67:216-223. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 25. | Baba Y, Tsuge D, Aoki R. Enhancement of carbohydrate metabolism by probiotic and prebiotic intake promotes short-chain fatty acid production in the gut microbiome: a randomized, double-blind, placebo-controlled crossover trial. Biosci Biotechnol Biochem. 2025;89:1191-1202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 10] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 26. | Ricaboni D, Mailhe M, Khelaifia S, Raoult D, Million M. Romboutsia timonensis, a new species isolated from human gut. New Microbes New Infect. 2016;12:6-7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 117] [Cited by in RCA: 87] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 27. | Van Hul M, Le Roy T, Prifti E, Dao MC, Paquot A, Zucker JD, Delzenne NM, Muccioli G, Clément K, Cani PD. From correlation to causality: the case of Subdoligranulum. Gut Microbes. 2020;12:1-13. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 163] [Article Influence: 27.2] [Reference Citation Analysis (4)] |
| 28. | Francavilla R, Cristofori F, Verzillo L, Gentile A, Castellaneta S, Polloni C, Giorgio V, Verduci E, DʼAngelo E, Dellatte S, Indrio F. Randomized Double-Blind Placebo-Controlled Crossover Trial for the Diagnosis of Non-Celiac Gluten Sensitivity in Children. Am J Gastroenterol. 2018;113:421-430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 51] [Article Influence: 6.4] [Reference Citation Analysis (4)] |
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/