Published online Mar 14, 2026. doi: 10.3748/wjg.v32.i10.115334
Revised: November 10, 2025
Accepted: December 16, 2025
Published online: March 14, 2026
Processing time: 138 Days and 22.1 Hours
Reversal of hepatic fibrosis (HF) represents a potential cure for chronic liver pathologies; however, clinically approved agents targeting this process remain scarce. Emerging evidence from traditional Chinese medicine (TCM) suggests that Hedyotis diffusa (HD), a botanical agent related to TCM principles of liver pa
To elucidate the efficacy and potential mechanism of HD against HF and to ex
Liquid chromatograph mass spectrometer revealed six bioactive components of HD injection (HDI) that enter the blood and liver. Network pharmacology using these components predicted related signaling pathways. A HF mouse model was induced by administration of 10% carbon tetrachloride for 8 weeks to validate the efficacy of HDI. Integrated Gene Expression Omnibus (GEO) mining and liver proteomics revealed the antifibrotic mecha
Network pharmacology analysis suggests that HDI may ameliorate HF through the modulation of circadian rhythm, urea metabolism, and hypoxia inducible factor-1 (HIF-1) signaling. GEO data mining and hepatic pro
HD demonstrates unequivocal efficacy in combating HF, potentially by modulating HIF-1 and the urea cycle through its influence on circadian rhythm genes, with NR1D1 as a prominent representative target.
Core Tip: This study reveals a novel antifibrotic mechanism of Hedyotis diffusa, demonstrating that its bioactive components alleviate hepatic fibrosis by restoring expression of the circadian rhythm gene NR1D1 expression. NR1D1 acts as an upstream regulator, inhibiting hypoxia inducible factor-1 signaling and restoring urea cycle function, thereby reducing hepatic ammonia accumulation and interrupting the vicious cycle of ammonia-induced hepatic stellate cell activation. These findings position circadian rhythm-metabolism interplay as a promising therapeutic axis for liver fibrosis treatment.
- Citation: Xia SW, Liu H, Yang KY, Gao YJ, Zhang MR, Zhou JW, Kong DS, Wu HY, Zhang F, Chen L. Integrative study reveals NR1D1 mediates Hedyotis diffusa’s antifibrosis via hypoxia inducible factor-1/ammonia axis. World J Gastroenterol 2026; 32(10): 115334
- URL: https://www.wjgnet.com/1007-9327/full/v32/i10/115334.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i10.115334
Chronic liver disease is a global public health issue, causing approximately 2 million deaths annually and posing a serious threat to human health[1]. Hepatic fibrosis is an essential stage in the progression of various chronic liver diseases to cirrhosis and hepatocellular carcinoma (HCC)[2]. Reversing hepatic fibrosis may cure most chronic liver diseases. The occurrence and development of hepatic fibrosis involve complex and multifactorial pathological processes[3], including abnormal deposition of extracellular matrix, dysregulation of cell signaling pathways, and changes in the hepatic micro
Many studies have indicated a close correlation between circadian rhythms and organismal energy metabolism[9-11]. Given the role of the liver as the metabolic hub of the body, its function may be subject to circadian regulation. Studies have shown that the circadian rhythm genes Cry and NR1D1/NR1D2 can regulate liver lipid metabolism and glucose production[12,13]. Additionally, a study involving 4740 male participants revealed that with increasing years of night shift work, the serum alanine aminotransferase (ALT) levels of night shift workers were higher than those of daytime workers, suggesting an association between circadian rhythm disruption and liver metabolic dysfunction[10]. These studies suggest that strategies aimed at regulating circadian rhythms are emerging as a novel field in liver disease re
The intricate metabolic regulatory network involved in the hepatic fibrosis process encompasses functional alterations in multiple cell types. Studies have confirmed that during advanced stages of cirrhosis, hepatocyte urea cycle dysfunction and impaired ammonia detoxification ultimately lead to hyperammonemia, which is closely associated with the pro
The characteristic metabolic disruption in the liver makes it difficult to achieve effective intervention in hepatic fibrosis through single targets or pathways[16,17]. The multicomponent, multitarget characteristics of traditional Chinese medicine (TCM) formulas may offer advantages in the treatment of complex metabolic diseases[18]. According to the TCM pathogenesis theory, the main pathogenic factor of hepatic fibrosis is ‘damp-heat evil toxins’, while an important aspect of disease pathology and pathogenesis is ‘blood stasis obstruction’[19,20]. Hepatic fibrosis is proposed to represent a dynamic process from ‘damp-heat’ to ‘stasis’, and from meridian disease to collateral disease. Hedyotis diffusa (HD), as a traditional Chinese herb known for its heat-clearing and detoxifying properties, exhibits significant anti-inflammatory, antioxidant, anticancer, and detoxifying effects, making it widely used in the treatment of various inflammatory diseases and cancers. Additionally, the aqueous extract of HD (HDW) is recognized as an excellent liver-protective agent and a key component in many TCMs for liver diseases. Studies have shown that HDW extract effectively exerts liver-protective effects in a zebrafish model of isoniazid-induced liver injury[21]. while HD decoction demonstrates notable anti-inflammatory effects in an in vivo model of lipopolysaccharide and galactosamine-induced liver inflammation[22]. However, although the roles of HD and its extracts in liver protection and anti-inflammation have been preliminarily validated, the specific mechanisms underlying their effects on liver fibrosis remain incompletely understood, particularly regarding the regulation of key signaling pathways such as circadian rhythm and metabolic reprogramming. Further in-depth research in this area will help reveal the potential value of HD in the treatment of liver fibrosis, providing a theoretical basis for developing more effective strategies for liver disease therapy.
The main purpose of this study was to clarify the anti-hepatic fibrosis efficacy of HD and to elucidate its potential mechanisms of action. To investigate the anti-hepatic fibrosis effects of HD, we employed network pharmacology to identify its potential therapeutic targets, revealing multiple pathway interventions involving circadian rhythm regulation, urea cycle metabolism, and hypoxia inducible factor-1 (HIF-1) signaling. Subsequent validation through Gene Expression Omnibus (GEO) database mining and hepatic proteomic profiling in fibrotic mice demonstrated strong correlations between disease progression and dysregulation of these pathways. Experimental investigations revealed that HD inter
HD injection (HDI) (Z36020632) was purchased from JiangXi Herbi-sky Co., Ltd; carbon tetrachloride (CCl4) (MFCD00000785) was purchased from McLean Reagent Co., Ltd; reverse transcription kit (AU341-02) was purchased from TransGen Biotech; SYBR Green Master Mix (Q111-02) was purchased from Vazyme Biotech; the primary antibody α-smooth muscle actin (SMA) (14395-1-AP), BMAL1 (14268-1-AP) and NR1D1 (14506-1-AP) were purchased from Proteintech Group, HIF-1α (GB111339-100) and β-actin (GB15001-100) were purchased from Servicebio; silymarin (22888-70-6) was purchased from Yuanye Biotech; the mitochondrial tracker Mito-Tracker Red CMXRos was purchased from Beyotime (Nanjing, Jiangsu Province, China).
Animal experiments (approval No. SYLL-2023-004) were conducted in compliance with the regulations set forth by the Institutional and Local Committee on the Care and Use of Animals of Jiangsu Vocational College of Medicine. Animals received humane care according to the guidelines established by the National Institutes of Health (United States). Experimental protocols adhered to the principles of the 3Rs and upheld the highest ethical and animal welfare standards. Male ICR mice (6-week-old, weighing 18-20 g) sourced from Nanjing Qinglongshan Animal Co., LTD. were acclimatized for one week under standard conditions: Room temperature maintained at 20 ± 2 °C, relative humidity at 40% ± 5%, with a 12-hour light/dark cycle (lights on at 7:00 and off at 19:00).
Animal experiments were mainly divided into three parts. The first part primarily aimed to elucidate the therapeutic efficacy of HDI against hepatic fibrosis and its potential mechanisms of action. The grouping and treatment methods of mice were as follows: Vehicle group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g olive oil for 8 weeks); CCl4-induced hepatic fibrosis group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks); CCl4 + silymarin group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injection of silymarin 100 mg/kg was given for the last 4 weeks); CCl4 + HDI low dose group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injections of HDI 5 mL/kg was given for the last 4 weeks); CCl4 + HDI medium dose group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injections of HDI 10 mL/kg was given for the last 4 weeks); CCl4 + HDI high dose group (n = 6, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injections of HDI 20 mL/kg was given for the last 4 weeks[23,24]).
The second part of the animal experiments focused on validating the upstream and downstream relationships of NR1D1 with the HIF-1 signaling pathway as well as the urea cycle and ammonia metabolism. The setting and operation of the part mice groups were as follows: Adeno-associated viruses (AAV) 9-NR1D1 knockdown (KD), AAV9-NR1D1 overexpression (OE), and AAV9-scrambles were constructed and purchased from Genomeditech Co. Ltd (Shanghai, China). Vehicle group (n = 6, AAV9-scrambles tail vein injection); NR1D1 KD group (n = 6, AAV9-NR1D1 KD tail vein injection); NR1D1 OE group (n = 6, AAV9-NR1D1 OE tail vein injection).
The purpose of the third part was to validate NR1D1 as an important target of HD for the treatment of liver fibrosis. The setting and operation of the part mice groups were as follows: AAV9-NR1D1 KD and AAV9-scrambles were constructed and purchased from Genomeditech Co. Ltd (Shanghai, China). Vehicle group (n = 6, AAV9-scrambles tail vein injection,1 week later injected intraperitoneally three times a week with 0.5 μL/g olive oil for 8 weeks); CCl4-induced liver fibrosis group (n = 6, AAV9-scrambles tail vein injection, 1 week later, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks); CCl4 + HDI group (n = 6, AAV9-scrambles tail vein injection, 1 week later, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injections of HDI 10 mL/kg was given for the last 4 weeks), NR1D1 KD + CCl4 + HDI group (n = 6, AAV9-NR1D1 KD tail vein injection, 1 week later, injected intraperitoneally three times a week with 0.5 μL/g 10% CCl4 olive oil solution, for 8 weeks; daily intraperitoneal injections of HDI 10 mL/kg was given for the last 4 weeks). 10% CCl4 olive oil solution was CCl4/olive 1:9 (v/v); zeitgeber time (ZT) was set as following rules: Set 7:00 as ZT0, 24 hours a cycle.
Human HSC-LX2 cells were obtained from the Cell Bank of Chinese Academy of Sciences (Shanghai, China). Cells were cultured in Dulbecco’s modified eagle medium (Invitrogen, Grand Island, NY, United States) with 10% fetal bovine serum, 1% antibiotics, and grown in a 5% carbon dioxide humidified atmosphere at 37 °C. Ammonium chloride (NH4Cl) was provided by Aladdin Biotech. Co. Ltd. (Shanghai, China). Briefly, HSC-LX2 cells at 60% confluence were co-cultured with NH4Cl at a final concentration of 5 mmol/L[25] in medium without serum and antibiotics for 24 hours.
Liver tissues were obtained from mice treated with HDI. A total of 200 mg of liver tissue was homogenized in 200 μL of ultrapure water for 10 minutes. The homogenate was then centrifuged at 12000 rpm for 10 minutes, and 100 μL of the supernatant was collected. Then, 200 μL of acetonitrile was added, followed by vortexing for 30 seconds and shaking for 3 minutes. After centrifugation at 12000 rpm for 10 minutes, the supernatant was transferred to another 1.5 mL centrifuge tube as much as possible and dried under nitrogen at 40 °C. The residue was reconstituted in 100 μL of 50% methanol, vortexed for 15 seconds, shaken for 3 minutes, and centrifuged at 12000 rpm for 10 minutes twice. The supernatant was collected after each centrifugation for analysis. The chromatographic conditions are shown in Table 1. Positive and negative ion modes were analyzed separately using an electro spray ionization-equipped mass detector. Time of flight (TOF)-mass spectrometer (MS) parameters were optimized as follows: TOF-MS scan mode covered the molecular weight range of m/z 100-1000, with a cumulative time of 0.2499 seconds. The ion source temperature was set to 550 °C, gas 1 at 50 psi, gas 2 at 60 psi, and the curtain gas at 20 psi. Declustering potential was set at 80 V, collision energy at 10 eV. Ion spray voltage floating was 5500 V for positive mode and 4500 V for negative mode. Data collection utilized information-dependent acquisition, dynamic background subtraction, and high-sensitivity mode. Instrument operation and data acquisition were controlled via Analyst® TF 1.6 software from AB SCIEX.
| Time (minute) | Analytical flow (μL/minute) | A (%) | B (%) |
| 0 | 300 | 95 | 5 |
| 1.0 | 300 | 95 | 5 |
| 15.00 | 300 | 60 | 40 |
| 20.00 | 300 | 40 | 60 |
| 25.00 | 300 | 5 | 95 |
| 28.00 | 300 | 5 | 95 |
| 28.1 | 300 | 95 | 5 |
| 32 | 300 | 95 | 5 |
We employed network pharmacology to systematically predict the potential targets and therapeutic mechanisms of HDI against hepatic fibrosis. The specific procedures were as follows. First, the chemical structures of the six primary bioactive components of HDI were obtained from the PubChem database, and their potential protein targets were predicted using the Pharm Mapper server. Targets with a fit score > 0 were selected and standardized via the UniProt database to retain only human targets. Disease-associated targets were retrieved by searching the keyword “liver fibrosis” in the GeneCards, National Center for Biotechnology Information (NCBI) gene database, and Online Mendelian Inheritance in Man (OMIM) databases. After the duplicates were merged and removed, 2651 unique targets related to hepatic fibrosis were identified. The drug targets and disease targets were intersected using Venny 2.1 software, resulting in 101 common targets. These common targets were then imported into the STRING database (v11.0) to construct a protein-protein interaction network, with the organism set to “Homo sapiens” and a minimum interaction confidence threshold of > 0.4. The resulting network, comprising 101 nodes and 836 edges, was imported into Cytoscape software (v3.8.0) for topological analysis using the NetworkAnalyzer tool. The top 30 hub targets were identified based on their degree values. To elucidate the complex relationships among components, diseases, and targets, a multidimensional network diagram was constructed using Cytoscape, and the importance of each component was evaluated based on its degree value. Finally, Gene Ontology (GO) functional annotation and Kyoto Encyclopedia of Genes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses for the common targets were performed using the clusterProfiler package in R software (v4.0.3). Terms with an adjusted P value of less than 0.05 were considered statistically significant for identifying key biological processes and signaling pathways.
The GEO gene set GSE990877 and GSE33650 were underwent analysis, focusing on gene expression in nonalcoholic steatohepatitis (NASH)-HCC tumor and precarcinoma tissues. KEGG and GO analyses were conducted using Sangerbox (http://www.sangerbox.com/tool). The analysis parameters included a minimum gene set of 5 and a maximum gene set of 5000, with statistical significance defined as a P value < 0.05 and false discovery rate < 0.25.
Liver samples were used to extract total RNA using Trizol reagent (Sigma, Saint Louis, MO, United States). The RNA was then reverse transcribed into complementary DNA using the TransScript all-in-one first-strand complementary DNA synthesis super mix for quantitative polymerase chain reaction (qPCR) kits from TransGen Biotech Co. Ltd. (Beijing, China), following the provided protocols. Real-time PCR was conducted using the SYBR Green Master Mix from Vazyme Biotech Co. Ltd. (Nanjing, Jiangsu Province, China), following the manufacturer’s instructions. The fold changes in messenger RNA levels of target genes were normalized to the invariant control glyceraldehyde phosphate dehydrogenase. Primers were obtained from GenScript Co. Ltd. (Nanjing, Jiangsu Province, China) and are listed in Supplementary Table 1.
Whole cell protein extracts were prepared from mouse liver tissues using radio immunoprecipitation assay (RIPA) buffer. Protein detection and blot visualization using Image J were performed according to standard methods. β-actin was used as invariant controls for equal loading of total proteins. Representative blots were shown.
Blood samples were collected from the intraocular fossa vein of mice, and serum liver biochemical indexes [ALT, aspartate aminotransferase (AST), alkaline phosphatase, lactate dehydrogenase, total bilirubin, laminin (LN), hyaluronidase (HA), procollagen-III, type IV collagen] were detected by Hitachi 7020 chemical analyzer.
The experimental procedure was adapted from previously published literature[26]. In brief, mouse liver tissues were fixed in 10% neutral buffered formalin, embedded in paraffin, and sectioned (5 μm thickness). Histological evaluation of liver tissue was performed using hematoxylin-eosin (HE) staining, while collagen deposition was examined using Masson and Sirius red staining. Images were captured blindly from five random fields under a microscope (ZEISS Axio Vert. A1, Germany), and representative views were presented.
Primary antibodies against α-SMA were applied to liver tissue sections (5 μm thickness), followed by immunohistochemical analysis using standard methods. Images were captured blindly from five random fields under a microscope (ZEISS Axio Vert. A1, Germany). Representative views were presented.
Liver tissue samples underwent lysis using RIPA solution (composed of 1% Triton X-100, 1% deoxycholate, and 0.1% sodium dodecyl sulfate, with protease inhibitor added prior to use) and were quantified via the bicinchoninic acid method. Protein samples (200 μg) were subjected to digestion using the filter-aided sample preparation technique. Following digestion, peptides were desalted using Ziptip C18 cartridges (Millipore, MA, United States) and quantified using a NanoDrop spectrophotometer (Thermo Scientific, United States). For label-free proteomics analysis, three independent biological replicates from each experimental group were analyzed using an on-line nano liquid chromatograph system (EASY-nLC 1200, Thermo Scientific) coupled to an Orbitrap Exploris 480 MS equipped with high-field asymmetric waveform ion mobility spectrometry. Chromatographic separation was achieved using an analytical column. Mass spectrometry was conducted in data-dependent analysis mode with dynamic exclusion set to 30 seconds. Full-scan MS spectra (m/z 350-1500) were acquired with a resolution of 60000 (m/z 200), followed by fragmentation of the most intense ions within a 1 second cycle time using high-energy collisional dissociation with a normalized collision energy of 30.0. MS/MS scans were acquired with a resolution of 15000 (m/z 200). The resulting ‘proteingroups.txt’ table was filtered for contaminants and reverse hits, and the number of unique peptides (≥ 1) was determined using Perseus version 1.5.3.2. Log2 transformation of label free quantitation intensity values and two-sample t-tests were performed to identify differentially expressed proteins (DEPs) among the experimental groups. Proteins with an average |log2 fold change| ≥ 0.585 and a P value < 0.05 were considered to be DEPs.
Ammonia level of liver tissues and LX2 cells were detected by ELISA kit purchased from YIFEI XUE Biotechnology.
Immunofluorescence staining with LX2 cells was performed according to our previously reported methods. Diamidino-phenyl-indole was used to stain nucleus. Images were blindly taken at five random fields under a microscope (ZEISS Axio Vert. A1, Germany). Representative views were shown
Mitochondrial fission or fusion in LX2 was labeled with mito-tracker, according to previous report[27]. When the cells reached a certain density, the cell culture medium was removed and the prepared mito-tracker red CMXRos (Beyotime, China) working solution was added and incubated at 37 °C for 15-30 minutes. Then, the working solution was removed and fresh cell culture medium pre-incubated at 37 °C was added. Confocal microscopy (Leica, TCS SP8, Germany) was used for detection.
The data were expressed as mean ± SD and analyzed using GraphPad Prism 9.0 (San Diego, CA, United States). Student’s t-test was employed to assess significant differences between two groups for normally distributed data, while one-way analysis of variance followed by post hoc Tukey’s test was utilized for comparisons among multiple groups. For non-normally distributed data, the Mann-Whitney U test was applied for comparisons between two groups, and the Kruskal-Wallis H test followed by post hoc Steel-Dwass test was used for comparisons among multiple groups. A P value < 0.05 was considered statistically significant.
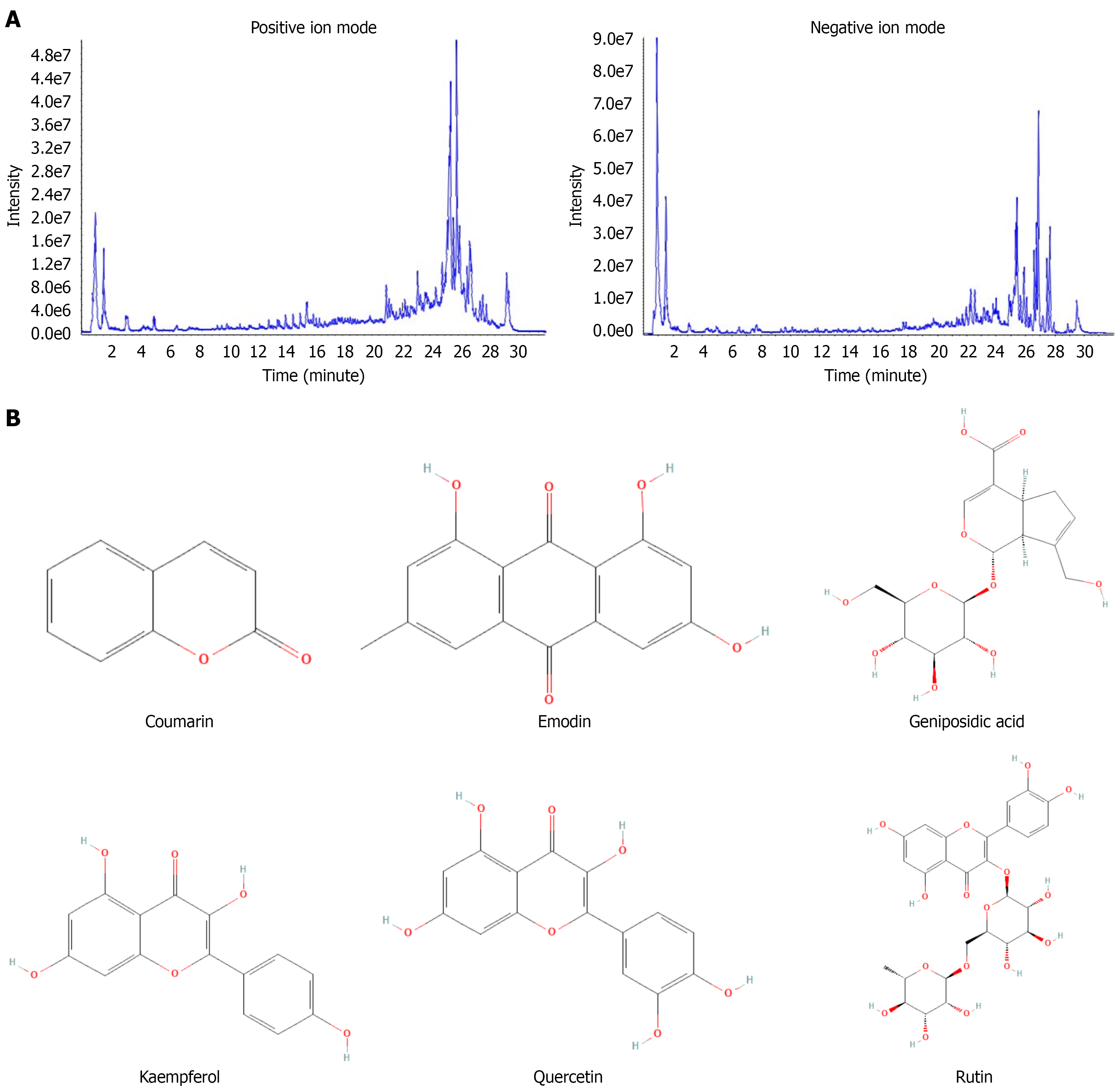
To identify the bioactive constituents of HDI that exert hepatic effects, liver tissues were collected from hepatic fibrosis model mice following HDI treatment and subjected to component analysis using high-performance liquid chromatography coupled with quadrupole TOF mass spectrometry-MS. The mass spectrometry profiles of reference standards are shown in Figure 1A, with the chemical structures of identified hepatic components illustrated in Figure 1B. A total of six primary bioactive constituents absorbed into the liver were identified through HDI intervention, as detailed in Table 2. These findings suggest these components may critically contribute to HDI’s antifibrotic efficacy.
| RT (minute) | Compound | Formula | Adduct | MS2 | Source |
| 1.52 | Coumarin | C9H6O2 | [M+H] + | 147.0438; 119.0492; 91.0541; 65.0387 | Liver |
| 9.88 | Geniposidic acid | C16H22O10 | [M+H] + | 375.1299; 357.1190; 285.0976; 243.0874; 172.0860; 97.0280 | Liver |
| 10.21 | Quercetin | C15H10O7 | [M+H] + | 303.0756; 303.0507; 257.0453; 229.0482; 153.0179 | Liver |
| 11.07 | Kaempferol | C15H10O6 | [M+H] + | 287.0545; 213.0531; 165.0182; 153.0187; 121.0275 | Liver |
| 11.08 | Rutin | C27H30O16 | [M-H] - | 609.1481; 285.0412; 284.0343; 255.0312; 227.0356; 206.8698 | Liver |
| 17.92 | Emodin | C15H10O5 | [M-H] - | 269.0453; 159.0437; 135.0449; 133.0291; 65.0041; 63.0241 | Liver |
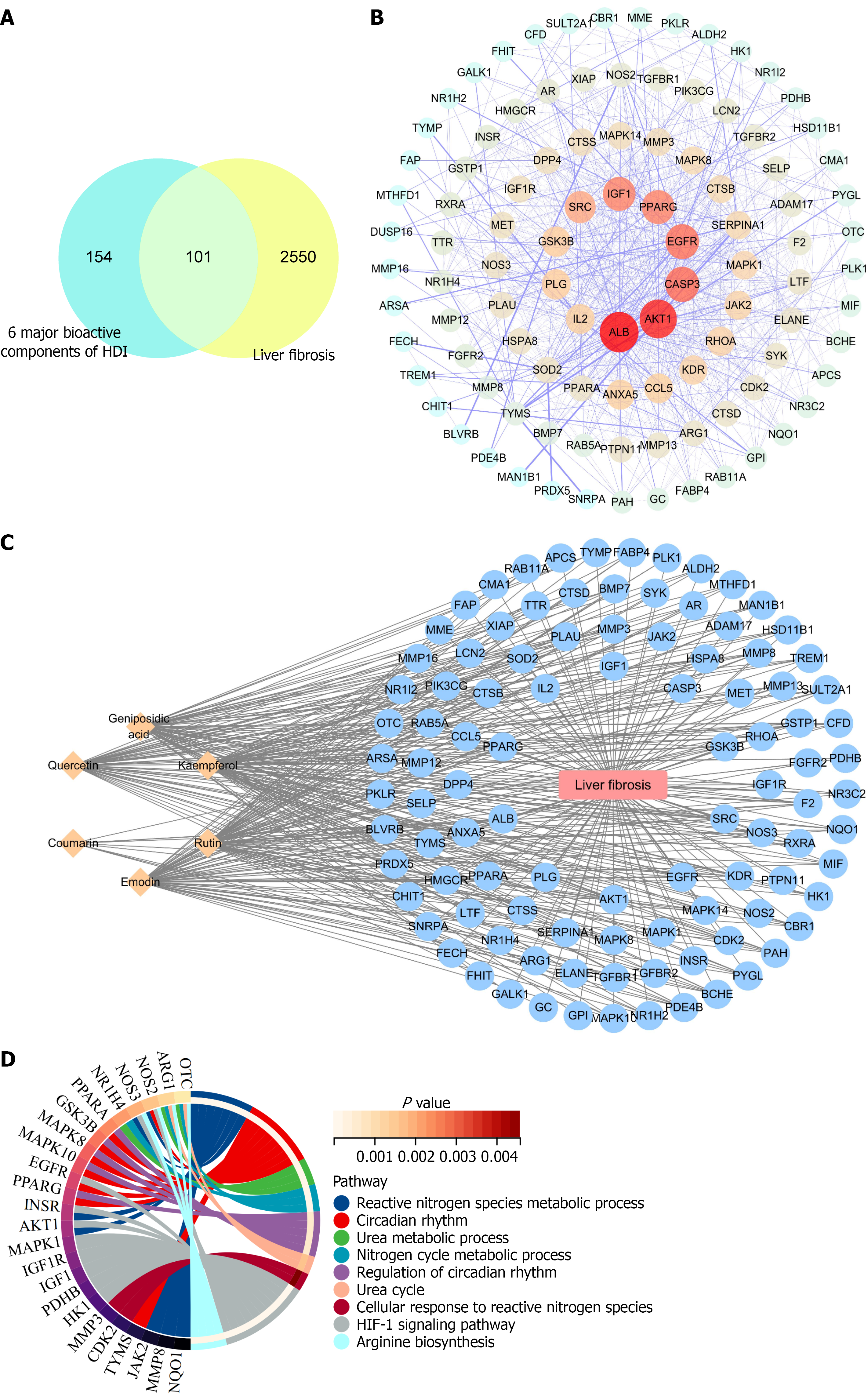
Network pharmacology is widely applied to explore the effects and potential mechanisms of multi-component drugs[28]. It is proposed that hepatic fibrosis is a dynamic process from ‘damp-heat’ to ‘stasis’, from meridian disease to collateral disease. Based on this disease theory, we believe that the HDI is a potential therapeutic drug for liver fibrosis. Therefore, we analyzed the potential therapeutic effects and mechanisms of HDI on liver fibrosis through network pharmacology. The six major bioactive components of HDI were used to analyzed with liver fibrosis disease targets which obtained from the GeneCards, NCBI, and OMIM databases. The screened drug targets and disease targets were input into Venny 2.1 software to generate a Venn diagram, identifying 101 overlapping targets (Figure 2A), which were subsequently selected as potential therapeutic targets for pathway enrichment analysis. The protein-protein interaction network was imported into Cytoscape 3.8.0 for topological analysis using the NetworkAnalyzer tool. Based on degree ranking, the top 30 hub targets were visualized using R4.0.3 (Figure 2B). To elucidate the intricate interactions among bioactive components, diseases, and associated targets, a component-disease-target network diagram was constructed (Figure 2C). KEGG pathway enrichment analysis of the shared drug-disease targets revealed that HDI’s hepatic components may exert therapeutic effects by modulating circadian rhythm signaling, urea metabolic process, and the HIF-1 signaling pathway (Figure 2D).
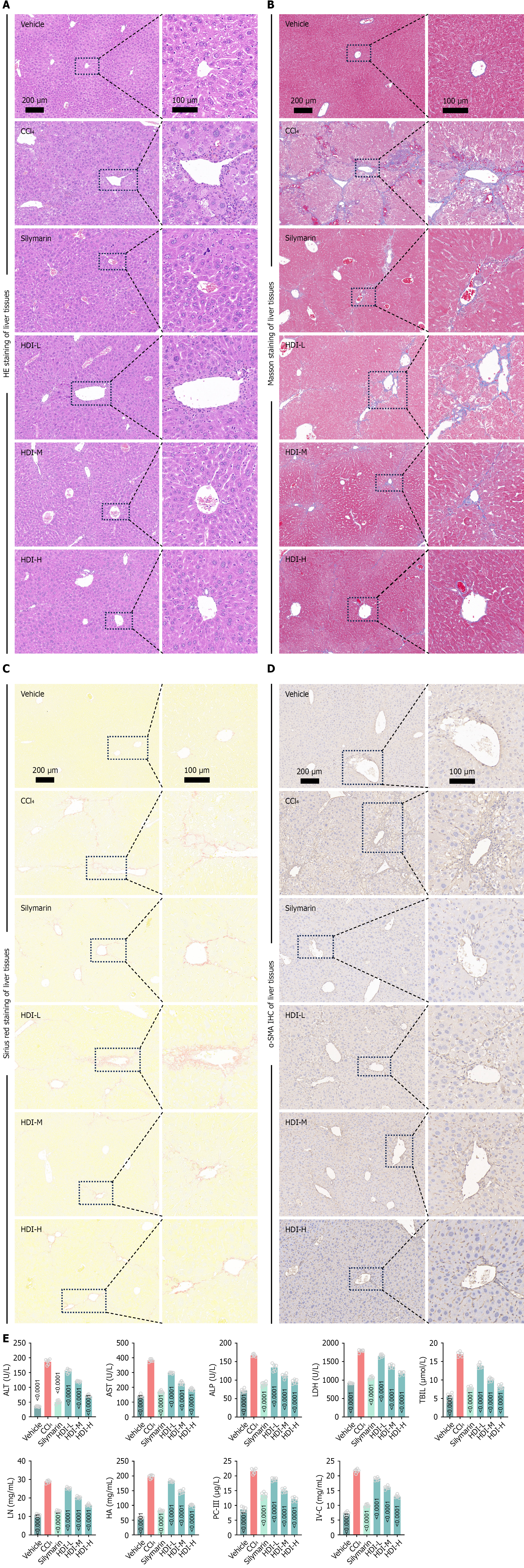
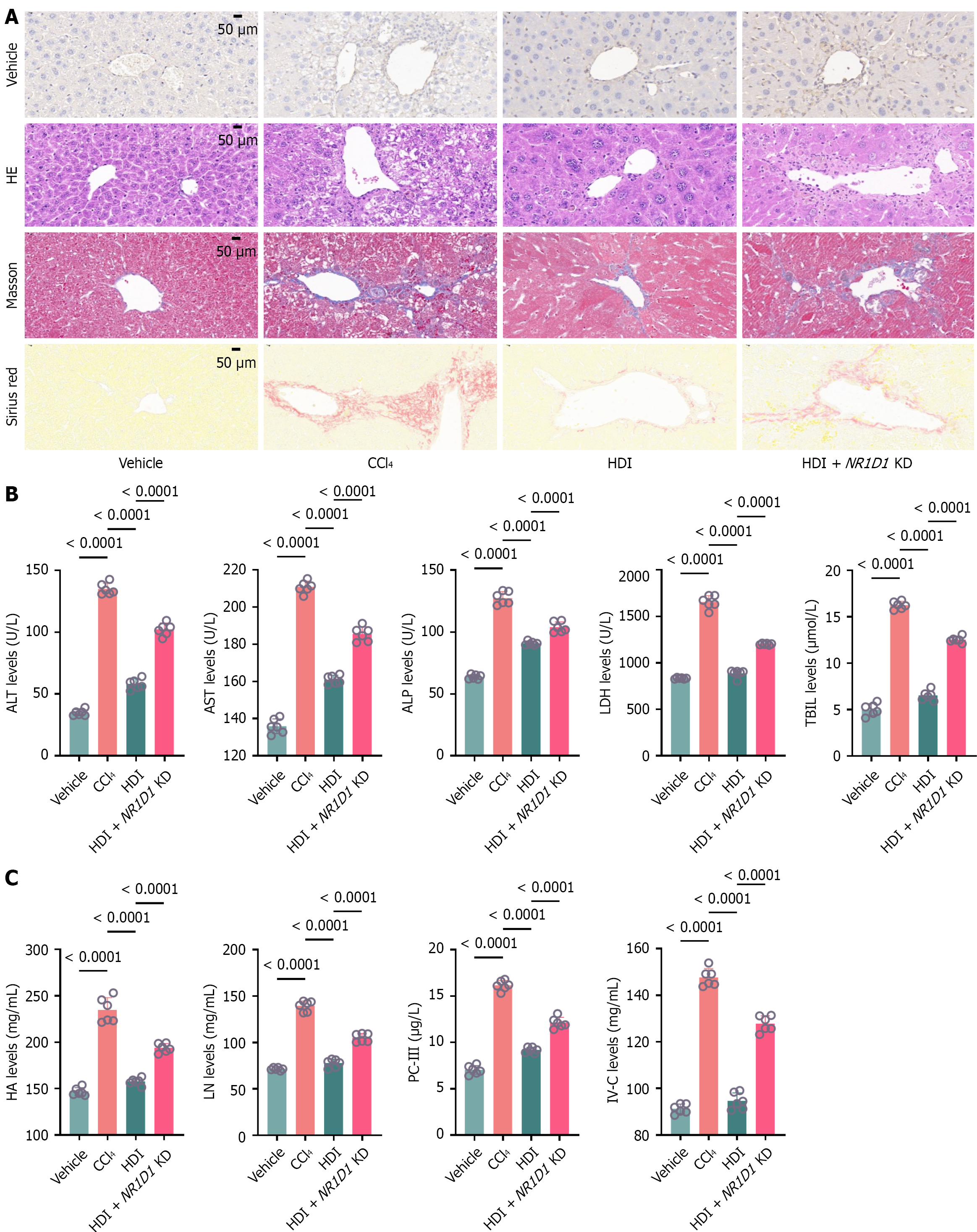
The above findings gave us a clue that HDI can treat liver fibrosis, and further we verified the anti-hepatic fibrosis effect of HDI in mice. We verified the antifibrotic effects of HDI using a CCl4-induced liver fibrosis mouse model. Initially, we assessed liver tissue damage and collagen deposition in mice treated with HDI through HE, Masson, and Sirius red staining. Additionally, we examined the levels of the HSC activation marker α-SMA by immunohistochemistry (Figure 3A-D). The results showed that, compared to the model group, HDI significantly ameliorated liver fibrosis damage and inhibited HSC activation. Further, serum biochemical tests for liver injury and fibrosis indicators demonstrated that HDI reduced liver injury and fibrosis markers at the systemic level in mice (Figure 3E), exhibiting significant antifibrotic effects. The in vivo experiments in mice indicate that HDI indeed has notable antifibrotic properties. However, the specific mechanisms of its action require further investigation.
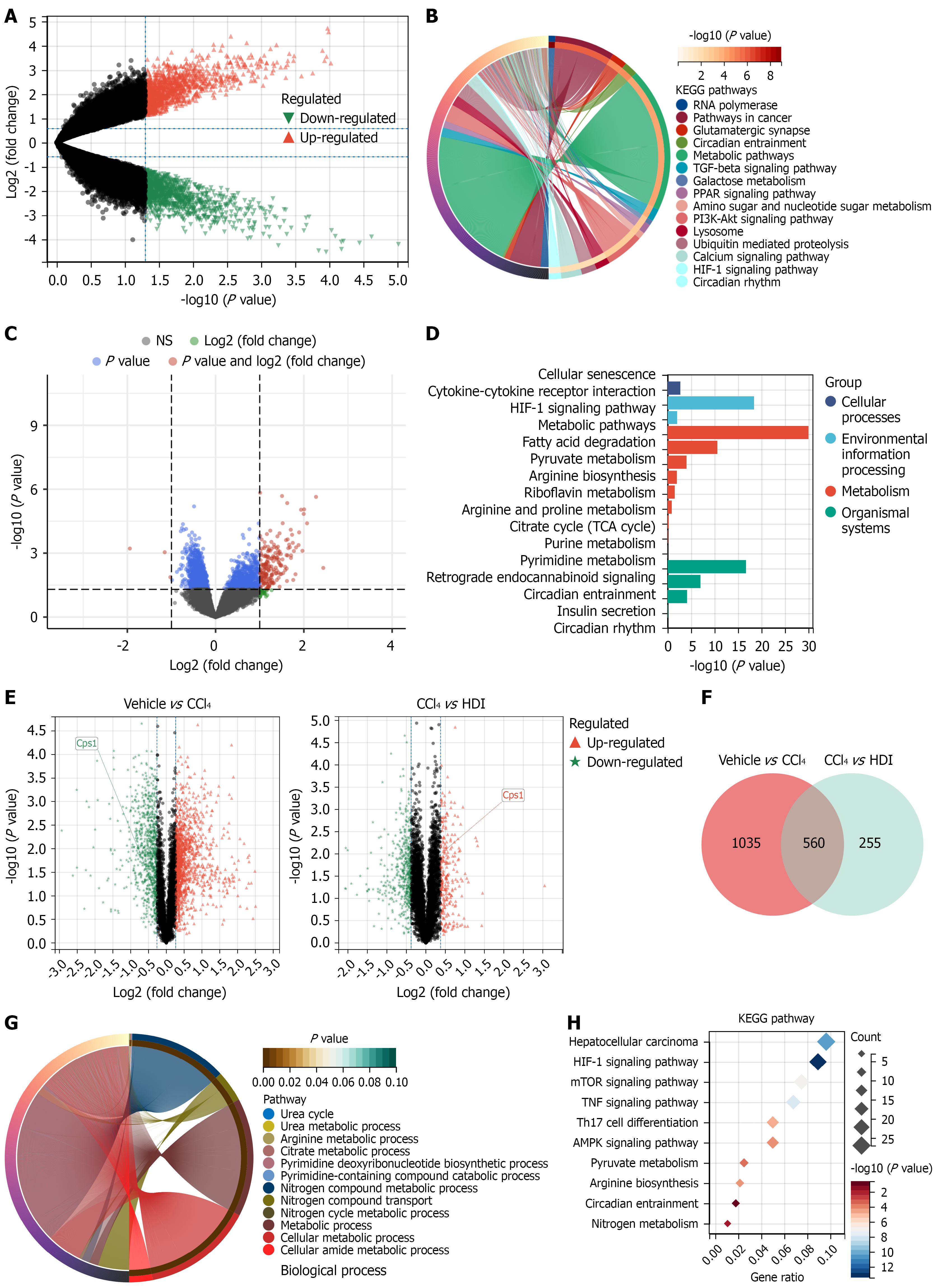
Liver fibrosis plays a crucial role in the progression of chronic liver diseases, with its continuous advancement being a major factor in the progression from general liver disease to cirrhosis and even liver cancer[29]. To investigate the clinical relevance of circadian rhythm and HIF-1 signaling in human liver disease progression a direction suggested by our preliminary network pharmacology analysis we first analyzed human GEO datasets. Analysis of the GSE990887 and GSE33650 datasets, which include samples from patients with NASH and HCC, revealed significant enrichment of genes associated with circadian rhythm and HIF-1 signaling pathways during the pathogenesis of human liver disease (Figure 4A-D). These findings in human samples provide a strong clinical rationale for further investigating the roles of these pathways in our experimental models. In this context, we performed proteomic analysis on the livers of mice from our CCl4-induced fibrosis model that underwent HDI treatment. The results of the analysis revealed that 560 proteins were significantly differentially expressed after HDI treatment (Figure 4E and F). KEGG and GO enrichment analyses of these HDI-regulated proteins revealed that HDI modulated circadian rhythm, the HIF-1 signaling pathway and urea metabolism in the mouse model (Figure 4G and H). These results, which are consistent with the network pharmacology prediction and supported by the human GEO data, strongly suggest that the anti-hepatic fibrosis mechanism of HD is related to these three important biological signals.
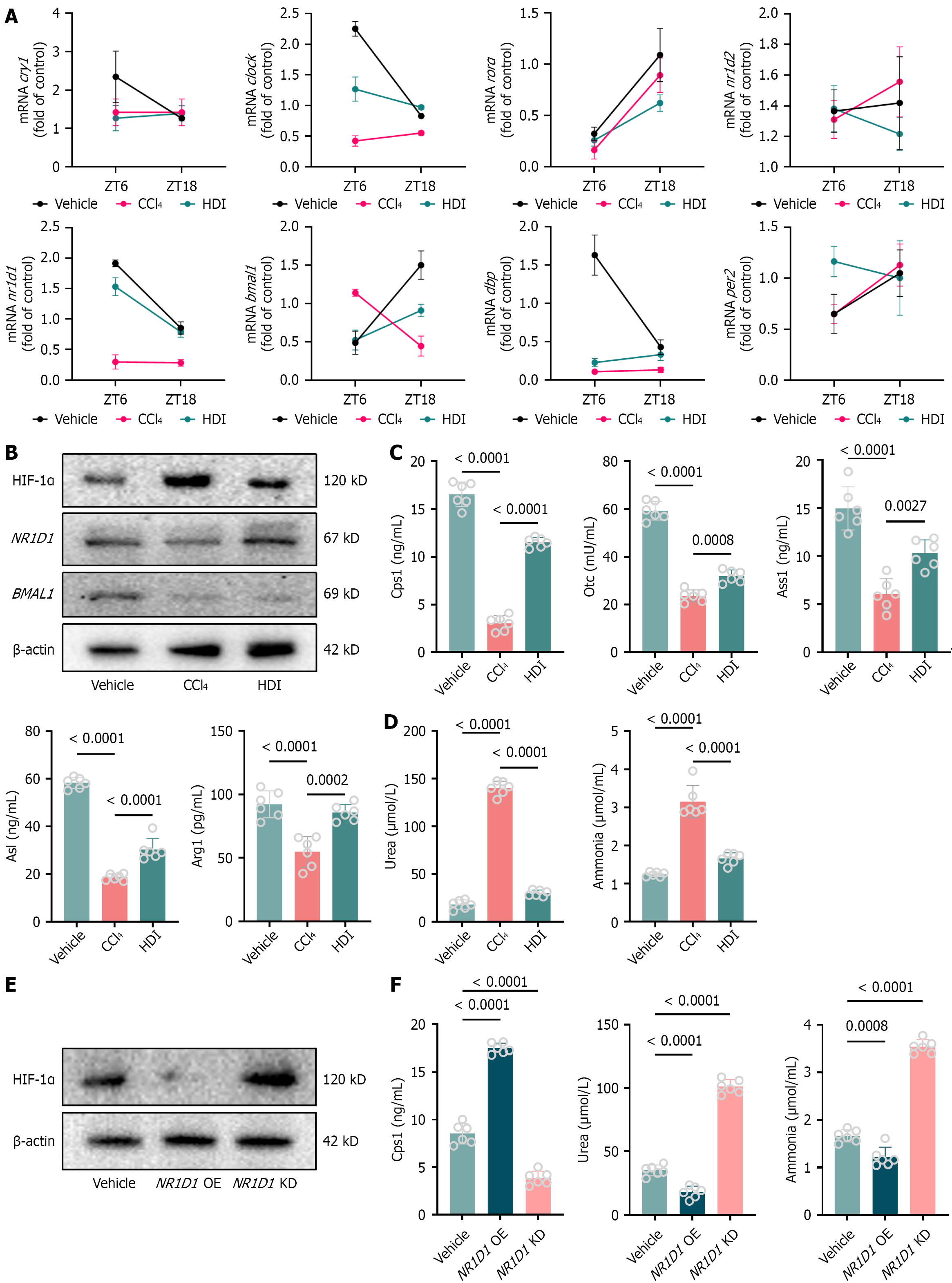
Based on the predictions from network pharmacology and the results of proteomic analysis, both highlighting the significance of the circadian rhythm pathway, we focused on analyzing the expression of core clock genes at circadian time points (ZT6 and ZT18). Given the central role of NR1D1 in metabolic regulation and its potential as a therapeutic target, we paid particular attention to it. PCR analysis revealed disrupted circadian gene expression profiles in fibrotic livers, with HDI most prominently restoring rhythmic oscillations of NR1D1 and BMAL1 (Figure 5A). Subsequent Western blot analysis demonstrated that HDI significantly upregulated NR1D1 protein levels while suppressing HIF-1α expression (Figure 5B). ELISA profiling of urea cycle enzymes showed widespread suppression in fibrotic models, which was reversed by HDI intervention, particularly evidenced by marked restoration of carbamoyl phosphate synthetase 1 (CPS1) activity (Figure 5C). Given the critical link between urea cycle dysfunction and ammonia dysregulation in fibrosis progression, we quantified hepatic urea/ammonia levels, finding that HDI significantly attenuated fibrosis-induced urea retention and hyperammonemia (Figure 5D). These findings collectively demonstrate that HDI ameliorates hepatic fibrosis through NR1D1-mediated coordination of urea cycle restoration and ammonia homeostasis.
To further elucidate NR1D1’s regulatory role in HIF-1 signaling , urea cycle, and ammonia metabolism, we established NR1D1 OE and KD murine models via AAV9-mediated hepatic delivery. Western blot analysis demonstrated an inverse correlation between NR1D1 expression and HIF-1α protein levels. ELISA quantification revealed that NR1D1 upregulation positively correlated with hepatic CPS1 activity while negatively associated with urea retention and hyperammonemia (Figure 5E and F), establishing NR1D1 as a master upstream regulator of these pathways. To verify NR1D1’s indispensability for HDI’s antifibrotic efficacy, we subjected NR1D1 KD mice to CCl4-induced fibrogenesis with concur
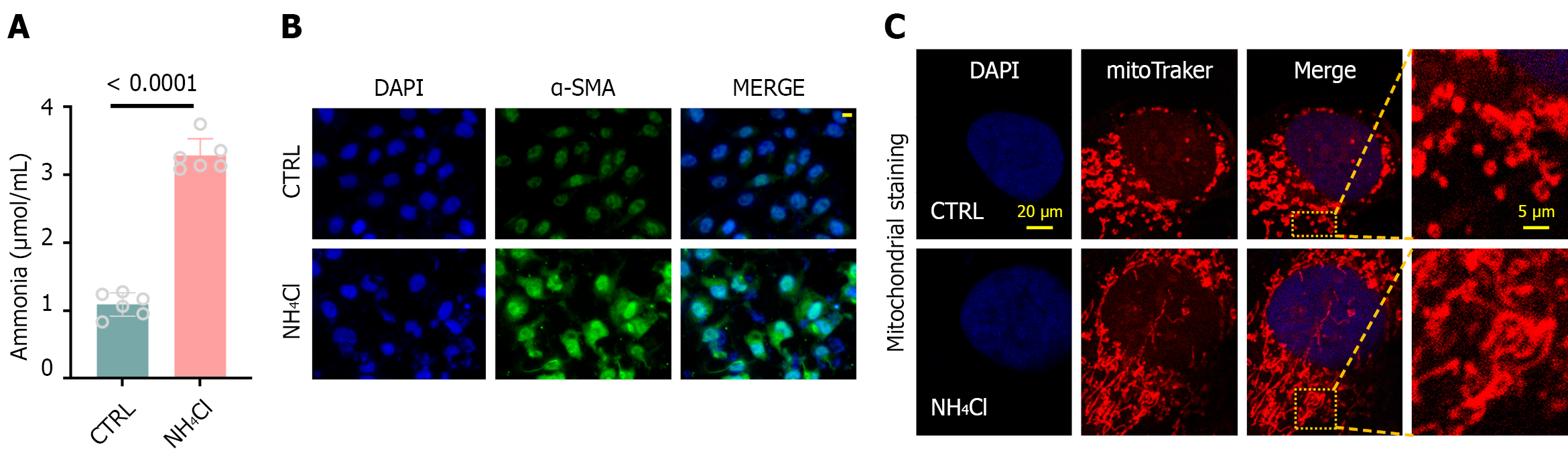
Building upon previous observations of ammonia accumulation in fibrotic livers, we conducted in vitro validation using LX2 cells a human HSC line. Ammonia overload was induced by co-culturing LX2 cells with 5 mmol/L NH4Cl, which significantly elevated intracellular ammonia levels as confirmed by ELISA (Figure 7A). Immunofluorescence analysis demonstrated marked upregulation of α-SMA expression under ammonia stress (Figure 7B), indicative of enhanced HSC activation. Given the critical role of mitochondrial dynamics in cellular activation states, we performed MitoTracker staining coupled with confocal microscopy, revealing ammonia-induced mitochondrial hyperfusion in LX2 cells (Figure 7C). In our previous study report[26], we have demonstrated that increased mitochondrial fusion in HSCs is an important phenotype of their abnormal activation. Therefore, we further concluded that hepatocellular injury in the disease state of liver fibrosis leads to impaired urea cycling, which in turn leads to abnormally high ammonia levels in hepatic tissues and promotes the continued activation of HSC, thus creating a vicious feedback loop. And HDI breaks this vicious cycle through the rhythm gene NR1D1 and plays an important role in ameliorating liver fibrosis.
The progression of liver fibrosis is typically accompanied by long-term liver damage caused by infectious (viral), chemical (mainly alcoholic), metabolic (nonalcoholic fatty liver disease), cholestatic, or autoimmune factors. Ultimately, fibrosis can lead to clinically significant cirrhosis and liver failure. Over the years, researchers have made steady progress in understanding the pathogenesis and clinical prognosis of liver fibrosis[5,30]. Although these studies have provided a substantial theoretical foundation for the development of antifibrotic drugs, no drugs have been approved to date[1]. Therefore, the ongoing exploration of the mechanisms of liver fibrosis and the search for effective medications remain key areas of research[6]. Recent studies have shown that TCM has unique advantages and efficacy in inhibiting the progression of liver fibrosis and promoting its reversal[31]. Researchers have identified many TCM compounds and individual herbs with significant antifibrotic effects and have elucidated their potential mechanisms at the cellular and molecular levels. However, theoretical innovation in the antifibrotic effects of TCM remains insufficient.
The circadian clock is an endogenous timing system that synchronizes the physiological activities of mammals with the day night cycle[10,32]. In recent years, studies have increasingly revealed that the circadian rhythm affects the entire digestive system, particularly the physiological functions of the liver, which appear to be tightly controlled by the circadian rhythm. Liver-specific rhythm gene knockout can lead to a significant disruption of the diurnal transcription processes of functional genes, thereby inducing liver diseases[13]. Current research has almost unequivocally established the essential relationship between liver function and circadian rhythms[9]. However, the relationship between liver fibrosis and circadian rhythm disturbances has not yet been extensively studied or reported.
Based on the aforementioned background, this study selected the heat-clearing and detoxifying TCM HD as the research subject, aiming to elucidate its pharmacological mechanisms. For the first time, by integrating network pharmacology, proteomics, GEO data mining, and in vivo and in vitro experimental validation, we systematically elucidated a novel multi-target antifibrotic mechanism. This mechanism involves HD improving ammonia metabolism and inhibiting the sustained activation of HSCs by regulating the core circadian gene NR1D1 to coordinate the HIF-1 signaling pathway and urea cycle function.
The establishment of this mechanism began with the screening of key targets. From numerous candidate targets, we prioritized NR1D1 for in-depth mechanistic investigation based primarily on the following three reasons: (1) Significant oscillation amplitude: Our qPCR data showed that during fibrosis progression, the oscillation of NR1D1 was the most severely disrupted (Figure 5A), suggesting it might be the most critically dysregulated target; (2) Core role in metabolic regulation: Authoritative studies have clearly established NR1D1 as a core negative regulator of hepatic lipid and glucose metabolism[12,13], which highly aligns with the essence of metabolic disruption in liver fibrosis; and (3) High drug ability: As a nuclear receptor, NR1D1 is a highly druggable target, making it more amenable to targeting and modulation by small-molecule compounds (such as components of Chinese herbal medicine)[33].
Finally, this study successfully established NR1D1 as the upstream master regulator in HD’s antifibrotic mechanism. Through AAV9-mediated gene intervention experiments, we confirmed that NR1D1 can simultaneously suppress HIF-1α protein expression, upregulate the activity of the urea cycle rate-limiting enzyme CPS1, and reduce tissue ammonia load (Figure 5E and F). More importantly, NR1D1 KD significantly attenuated the antifibrotic efficacy of HDI (Figure 6), confirming that HDI’s therapeutic effect is dependent on NR1D1. These findings suggest that circadian genes may act as core systemic regulators in complex metabolic diseases by establishing cross-pathway regulatory networks.
Furthermore, another significant finding of this study is the first revelation of the role played by the vicious cycle mechanism of “urea cycle dysfunction abnormal ammonia metabolism sustained HSC activation” in driving the progression of liver fibrosis. Although the association between hyperammonemia and the prognosis of advanced cirrhosis is recognized, the pathological significance of local ammonia metabolism dysregulation in the early and middle stages of fibrosis has long been overlooked. In this study, we found that during the fibrotic stage, even before the onset of systemic hyperammonemia, urea cycle function in hepatocytes was already significantly suppressed, leading to elevated local ammonia levels (Figure 5D). Using an NH4Cl-induced LX2 cell activation model, we discovered that ammonia load may promote sustained HSC activation (Figure 7A and B) by inducing mitochondrial fusion (Figure 7C) and altering the energy metabolism pattern of HSCs. Our data indicate that NR1D1 simultaneously regulates both HIF-1α protein levels and urea cycle function. An important and unanswered question is whether crosstalk exists between these two down
Although this study reveals the regulatory mechanism of HDI mediated by NR1D1 through a multi-omics integration strategy, several key aspects remain to be further explored, which also points the way for subsequent research. First, regarding component synergy, this study identified six liver-targeting active components in HDI, and whether their synergistic or dominant effect on NR1D1 regulation is a key unresolved issue. Although the specific contribution of each component has not been parsed, it is noteworthy that independent studies have suggested that several of these com
Second, a primary mechanistic limitation of our current work is the unclear pathway through which NR1D1 regulates HIF-1α stability. To directly elucidate this pathway, future studies should employ co-immunoprecipitation combined with MS to determine whether NR1D1 interacts with E3 ubiquitin ligases (such as Von Hippel-Lindau) involved in HIF-1α degradation, thereby revealing the underlying regulatory mechanism from the perspective of the protein interaction network.
Furthermore, although we hypothesize a vicious cycle hypothesis linking urea cycle dysfunction, ammonia accumulation, and HSC activation, direct experimental evidence for a positive feedback loop between ammonia and HIF-1α is still lacking. To rigorously test this hypothesis, future research could utilize an inducible HIF-1α gene knockout system to determine whether the absence of HIF-1α expression can abolish the suppression of urea cycle enzymes under ammonia stress conditions.
Finally, to bridge our preclinical findings to clinical application, it is crucial to validate, in prospective clinical cohorts, whether potential biomarkers (such as blood ammonia levels or urea cycle intermediate metabolites) can predict the efficacy of HDI. This constitutes a critical step toward precision medicine for treating hepatic fibrosis.
In this study, we systematically revealed the investigated the multitargeted antifibrotic mechanism of HD to overcome the vicious cycle of ‘ammonia metabolism-HSC activation’ through NR1D1-mediated rhythmic reprogramming and the synergistic regulation of the HIF-1 signaling pathway and the function of the urea cycle. These findings provide not only a scientific basis for the clinical application of HD but also a new paradigm for intervening in liver fibrosis from the perspective of biorhythm-metabolism interactions, providing an important theoretical reference for the design of therapeutic strategies for complex metabolic diseases.
| 1. | Younossi ZM, Wong G, Anstee QM, Henry L. The Global Burden of Liver Disease. Clin Gastroenterol Hepatol. 2023;21:1978-1991. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 396] [Cited by in RCA: 331] [Article Influence: 110.3] [Reference Citation Analysis (6)] |
| 2. | Bogomolova A, Balakrishnan A, Ott M, Sharma AD. "The Good, the Bad, and the Ugly" - About Diverse Phenotypes of Hepatic Stellate Cells in the Liver. Cell Mol Gastroenterol Hepatol. 2024;17:607-622. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 3. | Cheng D, Chai J, Wang H, Fu L, Peng S, Ni X. Hepatic macrophages: Key players in the development and progression of liver fibrosis. Liver Int. 2021;41:2279-2294. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 162] [Cited by in RCA: 151] [Article Influence: 30.2] [Reference Citation Analysis (0)] |
| 4. | Gao J, Wei B, de Assuncao TM, Liu Z, Hu X, Ibrahim S, Cooper SA, Cao S, Shah VH, Kostallari E. Hepatic stellate cell autophagy inhibits extracellular vesicle release to attenuate liver fibrosis. J Hepatol. 2020;73:1144-1154. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 255] [Cited by in RCA: 243] [Article Influence: 40.5] [Reference Citation Analysis (0)] |
| 5. | Kamm DR, McCommis KS. Hepatic stellate cells in physiology and pathology. J Physiol. 2022;600:1825-1837. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 260] [Cited by in RCA: 217] [Article Influence: 54.3] [Reference Citation Analysis (0)] |
| 6. | Trivedi P, Wang S, Friedman SL. The Power of Plasticity-Metabolic Regulation of Hepatic Stellate Cells. Cell Metab. 2021;33:242-257. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 357] [Cited by in RCA: 330] [Article Influence: 66.0] [Reference Citation Analysis (0)] |
| 7. | Yang L, Liu Y, Sun Y, Huang C, Li J, Wang Y. New advances of DNA/RNA methylation modification in liver fibrosis. Cell Signal. 2022;92:110224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 8. | Parola M, Pinzani M. Liver fibrosis: Pathophysiology, pathogenetic targets and clinical issues. Mol Aspects Med. 2019;65:37-55. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 998] [Cited by in RCA: 902] [Article Influence: 128.9] [Reference Citation Analysis (2)] |
| 9. | Reitmeier S, Kiessling S, Neuhaus K, Haller D. Comparing Circadian Rhythmicity in the Human Gut Microbiome. STAR Protoc. 2020;1:100148. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 42] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 10. | Wang F, Zhang L, Wu S, Li W, Sun M, Feng W, Ding D, Yeung-Shan Wong S, Zhu P, Evans GJ, Wing YK, Zhang J, Vlaanderen JJ, Vermeulen RCH, Zhang Y, Chan EY, Li Z, Tse LA. Night shift work and abnormal liver function: is non-alcohol fatty liver a necessary mediator? Occup Environ Med. 2019;76:83-89. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 40] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 11. | Cao X, Wang L, Selby CP, Lindsey-Boltz LA, Sancar A. Analysis of mammalian circadian clock protein complexes over a circadian cycle. J Biol Chem. 2023;299:102929. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 12. | Wada T, Yamamoto Y, Takasugi Y, Ishii H, Uchiyama T, Saitoh K, Suzuki M, Uchiyama M, Yoshitane H, Fukada Y, Shimba S. Adiponectin regulates the circadian rhythm of glucose and lipid metabolism. J Endocrinol. 2022;254:121-133. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 13. | Liu L, Liu L, Deng S, Zou L, He Y, Zhu X, Li H, Hu Y, Chu W, Wang X. Circadian Rhythm Alteration of the Core Clock Genes and the Lipid Metabolism Genes Induced by High-Fat Diet (HFD) in the Liver Tissue of the Chinese Soft-Shelled Turtle (Trionyx sinensis). Genes (Basel). 2024;15:157. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 8] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 14. | Katayama K. Clinical significance of the latency period of abnormal ammonia metabolism in chronic liver disease: Proposal of a new concept. Hepatol Res. 2022;52:75-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 15. | De Chiara F, Thomsen KL, Habtesion A, Jones H, Davies N, Gracia-Sancho J, Manicardi N, Hall A, Andreola F, Paish HL, Reed LH, Watson AA, Leslie J, Oakley F, Rombouts K, Mookerjee RP, Mann J, Jalan R. Ammonia Scavenging Prevents Progression of Fibrosis in Experimental Nonalcoholic Fatty Liver Disease. Hepatology. 2020;71:874-892. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 75] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 16. | Yan M, Li H, Xu S, Wu J, Li J, Xiao C, Mo C, Ding BS. Targeting Endothelial Necroptosis Disrupts Profibrotic Endothelial-Hepatic Stellate Cells Crosstalk to Alleviate Liver Fibrosis in Nonalcoholic Steatohepatitis. Int J Mol Sci. 2023;24:11313. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 17. | Cheng Z, Chen Y, Schnabl B, Chu H, Yang L. Bile acid and nonalcoholic steatohepatitis: Molecular insights and therapeutic targets. J Adv Res. 2024;59:173-187. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 23] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 18. | Ding X, He X, Tang B, Lan T. Integrated traditional Chinese and Western medicine in the prevention and treatment of non-alcoholic fatty liver disease: future directions and strategies. Chin Med. 2024;19:21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 25] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 19. | Yuan S, Wang N, Wang JL, Pan J, Xue XY, Zhang YN, Ma T. Gender differences in Damp-Heat Syndrome: A review. Biomed Pharmacother. 2021;143:112128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 20. | Choi T, Jun JH, Park B, Lee JA, You S, Jung J, Lee MS. Concept of blood stasis in Chinese medical textbooks: A systematic review. Eur J Integr Med. 2016;8:158-164. [RCA] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 19] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 21. | Wang X, Zhao J, Zhang R, Liu X, Ma C, Cao G, Wei Y, Yang P. Protective Effect of Hedyotis diffusa Willd. Ethanol Extract on Isoniazid-Induced Liver Injury in the Zebrafish Model. Drug Des Devel Ther. 2022;16:1995-2015. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 22. | Dai M, Wang F, Zou Z, Xiao G, Chen H, Yang H. Metabolic regulations of a decoction of Hedyotis diffusa in acute liver injury of mouse models. Chin Med. 2017;12:35. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 16] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 23. | Wu S, Wang X, Xing W, Li F, Liang M, Li K, He Y, Wang J. An update on animal models of liver fibrosis. Front Med (Lausanne). 2023;10:1160053. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 39] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 24. | Ge MX, Niu WX, Bao YY, Lu ZN, He HW. Sclareol attenuates liver fibrosis through SENP1-mediated VEGFR2 SUMOylation and inhibition of downstream STAT3 signaling. Phytother Res. 2023;37:3898-3912. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 25. | Li Z, Ji X, Wang W, Liu J, Liang X, Wu H, Liu J, Eggert US, Liu Q, Zhang X. Ammonia Induces Autophagy through Dopamine Receptor D3 and MTOR. PLoS One. 2016;11:e0153526. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 29] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 26. | Chen L, Xia S, Wang F, Zhou Y, Wang S, Yang T, Li Y, Xu M, Zhou Y, Kong D, Zhang Z, Shao J, Xu X, Zhang F, Zheng S. m(6)A methylation-induced NR1D1 ablation disrupts the HSC circadian clock and promotes hepatic fibrosis. Pharmacol Res. 2023;189:106704. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 27. | Chen S, Liu S, Wang J, Wu Q, Wang A, Guan H, Zhang Q, Zhang D, Wang X, Song H, Qin J, Zou J, Jiang Z, Ouyang S, Feng XH, Liang T, Xu P. TBK1-Mediated DRP1 Targeting Confers Nucleic Acid Sensing to Reprogram Mitochondrial Dynamics and Physiology. Mol Cell. 2020;80:810-827.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 54] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 28. | Wang Z, Linghu KG, Hu Y, Zuo H, Yi H, Xiong SH, Lu J, Chan G, Yu H, Huang RY. Deciphering the Pharmacological Mechanisms of the Huayu-Qiangshen-Tongbi Formula Through Integrating Network Pharmacology and In Vitro Pharmacological Investigation. Front Pharmacol. 2019;10:1065. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 20] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 29. | Devaraj E, Perumal E, Subramaniyan R, Mustapha N. Liver fibrosis: Extracellular vesicles mediated intercellular communication in perisinusoidal space. Hepatology. 2022;76:275-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 47] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 30. | Hou LS, Zhang YW, Li H, Wang W, Huan ML, Zhou SY, Zhang BL. The regulatory role and mechanism of autophagy in energy metabolism-related hepatic fibrosis. Pharmacol Ther. 2022;234:108117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 28] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 31. | Fu K, Wang C, Ma C, Zhou H, Li Y. The Potential Application of Chinese Medicine in Liver Diseases: A New Opportunity. Front Pharmacol. 2021;12:771459. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 31] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 32. | Durrington HJ, Krakowiak K, Meijer P, Begley N, Maidstone R, Goosey L, Gibbs JE, Blaikley JF, Gregory LG, Lloyd CM, Loudon ASI, Ray DW. Circadian asthma airway responses are gated by REV-ERBα. Eur Respir J. 2020;56:1902407. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 31] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 33. | Zhang-Sun ZY, Xu XZ, Escames G, Lei WR, Zhao L, Zhou YZ, Tian Y, Ren YN, Acuña-Castroviejo D, Yang Y. Targeting NR1D1 in organ injury: challenges and prospects. Mil Med Res. 2023;10:62. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 14] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 34. | Zhou G, Li C, Zhang R, Zhan Y, Lin L, Lang Z, Tao Q, Zheng J. Kaempferol Inhibits Hepatic Stellate Cell Activation by Regulating miR-26b-5p/Jag1 Axis and Notch Pathway. Front Pharmacol. 2022;13:881855. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 35. | Zheng Q, Wu X, Peng S. The immunotherapy mechanism of Hedyotis Diffusae Herba in treating liver cancer: a study based on network pharmacology, bioinformatics, and experimental validation. Naunyn Schmiedebergs Arch Pharmacol. 2025;398:951-965. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/