Published online Jun 15, 1997. doi: 10.3748/wjg.v3.i2.121
Revised: January 31, 1997
Accepted: March 1, 1997
Published online: June 15, 1997
AIM: To investigate the relationship between the loss of heterozygosity (LOH) of microsatellites on the deleted in colorectal carcinoma (DCC) gene and prognosis of colorectal adenocarcinoma.
METHODS: A retrospective study of 58 colorectal adenocarcinoma cases with follow-up data and paired control normal mucosal tissues from 1983 to 1985 from files from the West China University of Medical Sciences Department of Pathology was carried out by PCR microsatellite analysis. Sixteen, 35, and seven cases had well-, moderately, and poorly differentiated tumors, respectively; 11, 30, and 17 cases were staged as Dukes’ A, B, and C, respectively.
RESULTS: LOH of DCC microsatellites was detected in 18 cases (31.0%). The 5-year survival rate between LOH-positive and LOH-negative patients was 44.4% and 77.5%, respectively (P < 0.05). The results suggest that LOH of DCC microsatellites correlate with prognosis but not with differentiation (P > 0.05) and Dukes’ stage (P > 0.05) in colorectal adenocarcinoma.
CONCLUSION: LOH of DCC microsatellites may be a marker of malignancy. Combined with the traditional prognostic indicators, LOH can predict prognosis of colorectal adenocarcinoma.
- Citation: Zhao P, Yu YC, Wang DW, Wang ZP, Xu XZ, Yi PY, Gao YB, Yang GH. Relationship between loss of heterozygosity of deleted in colorectal carcinoma gene microsatellites and prognosis of colorectal adenocarcinoma. World J Gastroenterol 1997; 3(2): 121-122
- URL: https://www.wjgnet.com/1007-9327/full/v3/i2/121.htm
- DOI: https://dx.doi.org/10.3748/wjg.v3.i2.121
The deleted in colorectal carcinoma (DCC) gene is located on chromosome 18q21. In colorectal adenocarcinoma, there is an allelic deletion on the DCC gene and its level of expression is decreased, but few studies on the relationship between loss on the DCC gene and the prognosis of colorectal adenocarcinoma have been published to date. We used a PCR microsatellite method with a pair of primers flanking a dinucleotide microsatellite located within a DCC gene intron to study the relationship between the loss of heterozygosity (LOH) of DCC gene microsatellites and the prognosis of colorectal adenocarcinoma.
Paraffin blocks containing normal and neoplastic tissues from 71 patients with adenocarcinoma of the colon and rectum were selected for the study from files in the West China University of Medical Sciences Department of Pathology. The hematoxylin and eosin (H and E) slides from these patients were examined. Tumors were graded as well, moderately, or poorly differentiated and were assigned to Dukes’ stage A, B, or C based on the standard criteria. All patients had a 5-year follow up after surgery. The PCR microsatellite method was performed as described previously[1,2] but with slight modifications, i.e., ethidium bromide instead of isotope staining.
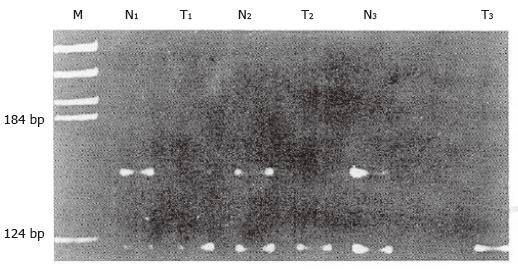
Fifty-eight amplified cases were informative for the marker of heterozygosity; 18 cases (31.0%) showed LOH. Figure 1 shows that compared to the tumor (T1) sample, the paired normal mucosa (N1) sample contained two amplified bands representing two different alleles: One paternal and another maternal, and no allele loss was scored when comparing the normal and tumor DNA. Allele loss was detected from case 2 (N2, T2) and case 3 (N3, T3), showing the loss of the top band as related to that of the normal tissue.
Of the 58 colorectal carcinoma cases, 19 patients died and 39 were still alive within five years, with a 5-year survival rate of 67.2%. The 5-year survival rate in patients with and without LOH of DCC gene microsatellites was 44.4% (8/18) and 77.5% (31/40), respectively (P < 0.05).
The positive rates of well-, moderately, and poorly differentiated colorectal adenocarcinoma were 25.0% (4/16), 31.4% (11/35), and 42.9% (3/7), respectively, in the LOH-positive cases, and 9.1% (1/11), 30.0% (9/30), and 47.1% (8/17) in Dukes’ stage A, B, and C, respectively. No statistically significant correlations were found between LOH and differentiation or Dukes’ stage.
Huang et al[2] reported LOH of DCC gene microsatellites in 33% (7/26) of colorectal adenocarcinoma cases, but they studied a small number of cases and no follow-up data. The frequency of LOH of DCC gene microsatellites in our study was similar to theirs, and our results were also concordant with theirs in terms of the relationship between LOH and differentiation or Dukes’ stage, which could be attributed to the small sample size in both studies. According to our follow-up data, LOH of DCC gene microsatellites might be closely related to the 5-year survival rate of patients with colorectal adenocarcinoma. Thus, we believe that when combined with traditional prognostic indicators, LOH of DCC gene microsatellites might be a new predictive marker of colorectal adenocarcinoma prognosis; however, a large number of cases are needed to confirm it.
| 1. | Risinger JI, Boyd J. Dinucleotide repeat polymorphism in the human DCC gene at chromosome 18q21. Hum Mol Genet. 1992;1:657. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 2. | Huang TH, Quesenberry JT, Martin MB, Loy TS, Diaz-Arias AA. Loss of heterozygosity detected in formalin-fixed, paraffin-embedded tissue of colorectal carcinoma using a microsatellite located within the deleted in colorectal carcinoma gene. Diagn Mol Pathol. 1993;2:90-93. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
Original title:
Presented at The Third International Symposium for Pathologists, October 1994, Guangzhou.
S- Editor: Filipodia L- Editor: Jennifer E- Editor: Liu WX