Published online May 28, 2023. doi: 10.3748/wjg.v29.i20.3145
Peer-review started: December 12, 2022
First decision: March 9, 2023
Revised: April 10, 2023
Accepted: April 27, 2023
Article in press: April 27, 2023
Published online: May 28, 2023
Processing time: 164 Days and 20.8 Hours
Cancer detection is a global research focus, and novel, rapid, and label-free techniques are being developed for routine clinical practice. This has led to the development of new tools and techniques from the bench side to routine clinical practice. In this study, we present a method that uses Raman spectroscopy (RS) to detect cancer in unstained formalin-fixed, resected specimens of the esophagus and stomach. Our method can record a clear Raman-scattered light spectrum in these specimens, confirming that the Raman-scattered light spectrum changes because of the histological differences in the mucosal tissue.
To evaluate the use of Raman-scattered light spectrum for detecting endoscop-ically resected specimens of esophageal squamous cell carcinoma (SCC) and gastric adenocarcinoma (AC).
We created a Raman device that is suitable for observing living tissues, and attempted to acquire Raman-scattered light spectra in endoscopically resected specimens of six esophageal tissues and 12 gastric tissues. We evaluated formalin-fixed tissues using this technique and captured shifts at multiple locations based on feasibility, ranging from six to 19 locations 200 microns apart in the vertical and horizontal directions. Furthermore, a correlation between the obtained Raman scattered light spectra and histopathological diagnosis was performed.
We successfully obtained Raman scattered light spectra from all six esophageal and 12 gastric specimens. After data capture, the tissue specimens were sent for histopathological analysis for further processing because RS is a label-free methodology that does not cause tissue destruction or alterations. Based on data analysis of molecular-level substrates, we established cut-off values for the diagnosis of esophageal SCC and gastric AC. By analyzing specific Raman shifts, we developed an algorithm to identify the range of esophageal SCC and gastric AC with an accuracy close to that of histopathological diagnoses.
Our technique provides qualitative information for real-time morphological diagnosis. However, further in vivo evaluations require an excitation light source with low human toxicity and large amounts of data for validation.
Core Tip: Cancer diagnosis is a critical step in patient management, and involves a combination of diagnostic modalities or a single investigation. Diagnostic techniques that provide comprehensive data on the disease process can be particularly valuable, and Raman spectroscopy (RS) is one such modality that offers detailed molecular-level information. In this study, we utilized RS to rapidly detect cancer in resected esophageal and stomach specimens, providing information beyond morphology. By providing detailed molecular-level data, RS can provide a more comprehensive understanding of disease processes and aid in accurate diagnosis.
- Citation: Ito H, Uragami N, Miyazaki T, Shimamura Y, Ikeda H, Nishikawa Y, Onimaru M, Matsuo K, Isozaki M, Yang W, Issha K, Kimura S, Kawamura M, Yokoyama N, Kushima M, Inoue H. Determination of esophageal squamous cell carcinoma and gastric adenocarcinoma on raw tissue using Raman spectroscopy. World J Gastroenterol 2023; 29(20): 3145-3156
- URL: https://www.wjgnet.com/1007-9327/full/v29/i20/3145.htm
- DOI: https://dx.doi.org/10.3748/wjg.v29.i20.3145
Gastrointestinal cancers are typically evaluated using various investigative modalities, like radiological imaging, endoscopy, and histopathology[1,2]. Although tissue diagnosis through histopathological evaluation remains the gold standard for diagnosis, it is primarily based on morphological inter
Endoscopic biopsy is the preferred diagnostic method for gastrointestinal cancer[6]. Recent technological advances in endoscopy have enabled reliable diagnoses based on endoscopic findings alone, which have been further improved by the incorporation of artificial intelligence[5-7]. However, these methodologies rely primarily on morphological assessments and do not consider molecular biological information. Tumor tissues contain abnormal molecules and proteins that are absent in normal tissues. The addition of molecular biological information to existing highly evolved morphological diagnostic methods may become an epoch-making technique with higher diagnostic accuracy[5,8,9].
Hence, we evaluated the biological samples using RS to obtain molecular information[8,10]. RS is a nondestructive inspection method that identifies a substance by measuring the type and amount of molecules contained in the substance by analyzing the wavelength of the reflected light obtained by irradiating the target substance with light, such as a laser[11-13]. Furthermore, RS can be used to evaluate substances with all types of properties, including solids[14-16], liquids[9,11], and gases[17], without sample pretreatment. Therefore, RS is an excellent inspection method for evaluating substances within a short time.
Therefore, attempts have been made to evaluate tissue samples using RS. Bergholt et al[4] presented a diagnostic technique for esophageal disease in the gastrointestinal tract using RS and published diagnostic techniques for gastric cancer. Moreover, Duraipandian et al[3] reported a diagnostic technique for gastric cancer using RS. However, a standard method for biological evaluations using RS has not yet been established.
A major challenge in using RS to evaluate biological samples is interference of autofluorescence[18]. To address this issue, we developed a novel micro-Raman device with unique features that enables effective evaluation of biological samples using RS.
We selected a near-infrared laser, which is not easily affected by autofluorescence, as the excitation light source. Because the Raman-scattered light intensity is inversely proportional to the square of the excitation light wavelength, the scattered light intensity of a long-wavelength near-infrared laser is low[18]. We developed a highly sensitive circuit to detect this weakly scattered light. We attempted to record the Raman-scattered light wavelengths from living esophageal tissues using a micro-Raman device with these characteristics.
Sixteen patients aged 80 years or younger who underwent endoscopic submucosal dissection of the esophagus or stomach at the Digestive Disease Center of Showa University Koto Toyosu Hospital were recruited for this study. Written consent was obtained from all participants. Eleven participants were male and five were female, with an age range of 38-80 years. Eighteen specimens (esophagus, n = 6; stomach, n = 12) were analyzed (Table 1). This study was approved by the in-hospital clinical research review board (approval number: 18T5009) and conducted in accordance with the Declaration of Helsinki.
| Patient number | Patient | Clinical diagnosis | Number of lesions | Treatment date |
| 1 | 50-year-old, female | Esophageal cancer | 1 | December 20, 2018 |
| 2 | 65-year-old, male | Esophageal cancer | 1 | December 20, 2018 |
| 3 | 61-year-old, male | Esophageal cancer | 1 | January 11, 2019 |
| 4 | 61-year-old, male | Esophageal cancer | 2 | January 18, 2019 |
| 5 | 78-year-old, female | Esophageal cancer | 1 | April 18, 2019 |
| 6 | 72-year-old, female | Stomach cancer | 1 | January 8, 2019 |
| 7 | 63-year-old, male | Stomach cancer | 1 | January 18, 2019 |
| 8 | 56-year-old, male | Stomach cancer | 1 | January 24, 2019 |
| 9 | 58-year-old, male | Stomach cancer | 1 | February 3, 2019 |
| 10 | 79-year-old, male | Stomach cancer | 1 | February 4, 2019 |
| 11 | 77-year-old, male | Stomach cancer | 1 | February 17, 2019 |
| 12 | 80-year-old, male | Stomach cancer | 2 | February 28, 2019 |
| 13 | 65-year-old, male | Stomach cancer | 1 | February 28, 2019 |
| 14 | 68-year-old, male | Stomach cancer | 1 | April 8, 2019 |
| 15 | 66-year-old, female | Stomach adenoma | 1 | May 19, 2019 |
| 16 | 38-year-old, female | Stomach submucosal tumor | 1 | April 9, 2019 |
Esophageal and stomach tissue samples (including mucosal and submucosal tissues) were collected immediately after endoscopic resection. The tissue was attached to a black rubber plate with a metal pin, and a 0.02 mm thick aluminum foil was sandwiched between the rubber plate and the esophageal tissue to prevent Raman scattered light from the rubber plate from being included in the measurements.
A linear or grid-like measurement grid was set up to include endoscopically diagnosed lesions in the mucosa and the surrounding normal mucosa. Multiple measurement points were selected at 5 mm or 10 mm intervals depending on the shape of the clinically determined lesion. Raman scattered light wavelengths were recorded from 19 or 6 locations at intervals of 200 μm in the vertical and horizontal directions for each selected point depending on the feasibility. To prevent the sample from drying out, distilled water was sprayed on the sample as necessary during the measurement.
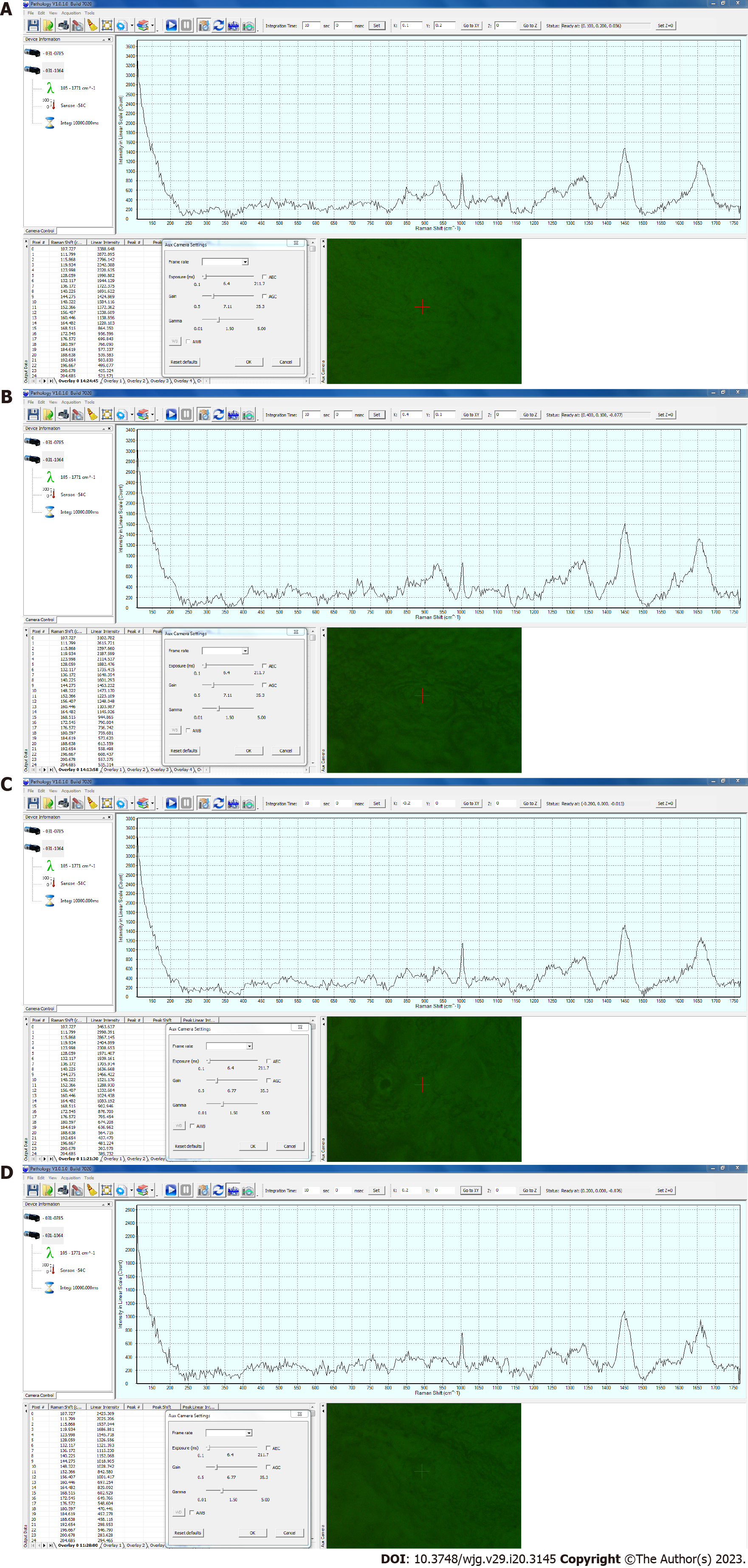
A RS device (BaySpec Inc., San Jose, CA, United States) was used, with a computer-controlled stage, an objective lens of 20 × magnification, a correction collar for near-infrared microscopy (LCPLN20XIR, Olympus Corporation, Tokyo, Japan), and an excitation laser wavelength of 1064 nm. The measurements were captured with a laser power of 200 mW and an exposure time of 10 s per point. Pathologic System Software Version 1.0.1.0 (BaySpec, Inc., San Jose CA, United States) was used and baseline correction was performed without smoothing the waveform. Examples of the Raman spectra of the esophageal and gastric mucosa are shown in Figure 1.
Fifteen types of Raman shifts corresponding to the constituent molecules of living tissues (Table 2) were selected for the analysis. From the recorded Raman-scattered light waveforms, the scattered light intensities of 15 types of Raman shifts were extracted, and the optimum combination matching the range of squamous cell carcinoma (SCC) of the esophagus and gastric adenocarcinoma (AC) by histopathological diagnosis was determined.
| Number | Assigned peak | Shift start (cm-1) | Shift end (cm-1) |
| RS 1 | Phenylalanine C-C twist | 611 | 631 |
| RS 2 | Cholesterol | 700 | 720 |
| RS 3 | Tryptophane ring breath | 751 | 771 |
| RS 4 | Tyrosine ring breath | 830 | 850 |
| RS 5 | Phenylalanine ring breath | 993 | 1013 |
| RS 6 | Skeletal C-C | 1060 | 1080 |
| RS 7 | Nucleotide O-P-O | 1091 | 1111 |
| RS 8 | Skeletal C-C stretch | 1123 | 1143 |
| RS 9 | Amide III beta-sheet | 1244 | 1264 |
| RS 10 | Amide III delta (CH)2 | 1275 | 1295 |
| RS 11 | Amide III alpha-helix | 1322 | 1342 |
| RS 12 | CH2 stretch | 1408 | 1428 |
| RS 13 | CH3 deformation | 1448 | 1468 |
| RS 14 | Phenylalanine C=C | 1596 | 1616 |
| RS 15 | Amide I alpha-helix | 1647 | 1667 |
The study involved 16 patients, and 18 specimens were resected for histopathological analysis. The results showed that five of the specimens were SCC of the esophagus, 10 were well-differentiated tubular AC of the stomach, one was a gastric adenoma, and one was a gastric carcinoid tumor (NET G1) (Table 3).
| Sample number | Patient | Pathological diagnosis (UICC TNM classification) |
| Eso-1 | 50-year-old, female | SCC, pT1a (LPM) |
| Eso-2 | 65-year-old, male | SCC, pT1a (LPM) |
| Eso-3 | 61-year-old, male | SCC, pT1a (MM) |
| Eso-4 | 61-year-old, male | SCC, pTis |
| Eso-5 | 61-year-old, male | SCC, pTis |
| Eso-6 | 78-year-old, female | SCC, pT1b (SM) |
| Sto-1 | 72-year-old, female | Tub1, pT1a |
| Sto-2 | 63-year-old, male | Tub1>tub2, pT1b2 |
| Sto-3 | 56-year-old, male | Tub1, pT1b1 |
| Sto-4 | 58-year-old, male | Tub1, pT1b1 |
| Sto-5 | 79-year-old, male | Tub1, pT1b1 |
| Sto-6 | 77-year-old, male | Tub1, pT1a |
| Sto-7 | 80-year-old, male | Tub1, pT1b1 |
| Sto-8 | 80-year-old, male | Tub1, pT1a |
| Sto-9 | 65-year-old, male | Tub1, pT1a |
| Sto-10 | 68-year-old, male | Tub1, pT1a |
| Sto-11 | 66-year-old, female | Tubular adenoma |
| Sto-12 | 38-year-old, female | Carcinoid tumor (NET G1), pT1b |
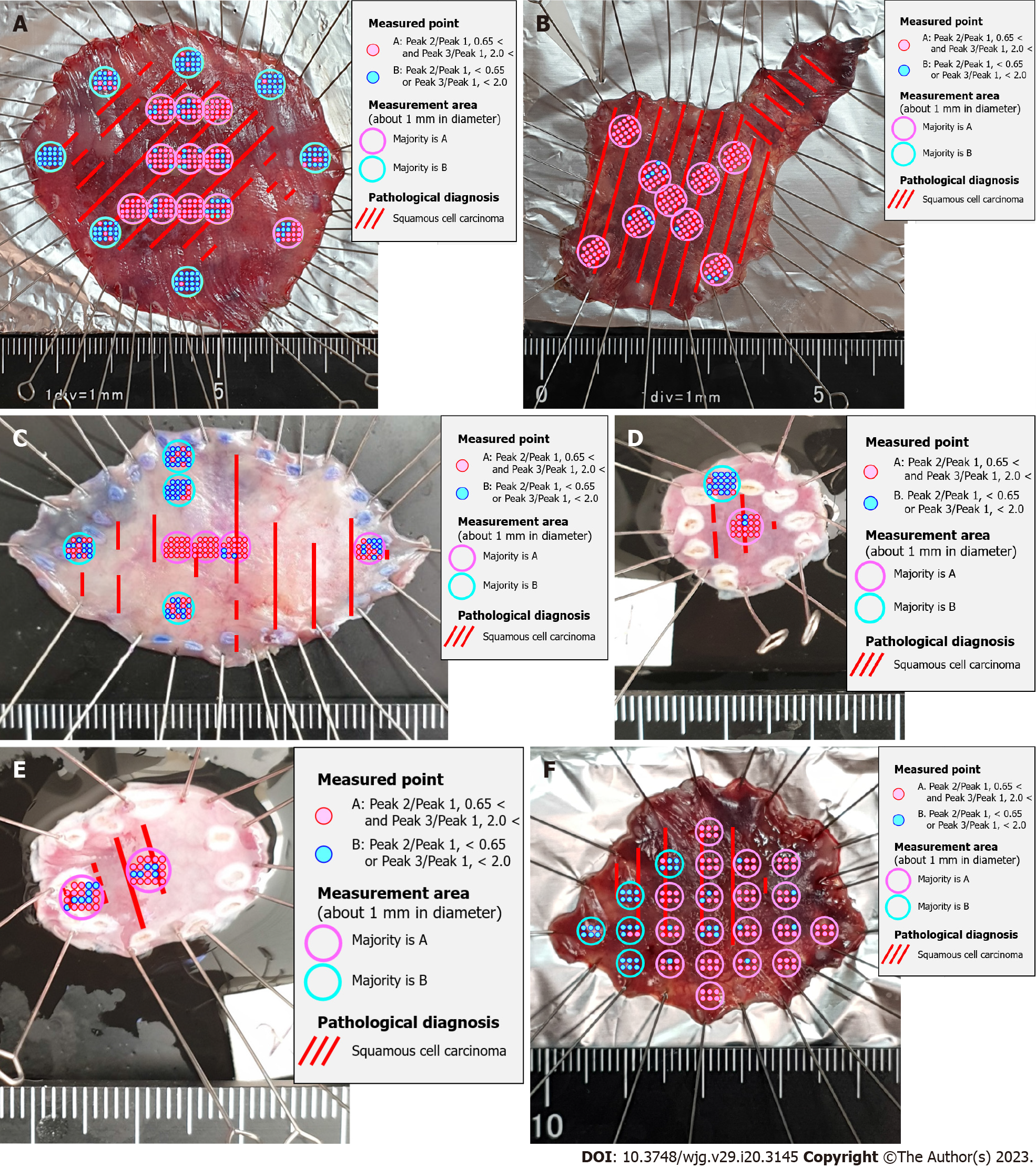
Raman spectra were recorded for all six esophageal and 12 gastric specimens. All combinations of the scattered light intensities of the 15 Raman shifts were calculated to determine the combination that best matched the range of cancer by histopathological diagnosis. In esophageal squamous cell cancer, the scattered light intensity of Skeletal C-C stretch was Peak 1, the scattered light intensity of Phenylalanine C-C twist was Peak 2, the scattered light intensity of CH3 deformation was Peak 3 (Table 4), and Peak 2/Peak 1 was defined as Cut-off 1. Peak 3/Peak 1 was defined as Cut-off 2 (Table 5). The site with more than half of the measurement points having Cut-off 1 greater than 0.65 and Cut-off 2 greater than 2.0 was determined to be the best match for histopathological SCC of the esophagus (Figure 2).
| Peak 1 | Peak 2 | Peak 3 | |
| Esophagus | RS 8 | RS 1 | RS 13 |
| Stomach | RS12 | RS 5 | RS 15 |
| Cut-off 1 (peak 2/peak 1) | Cut-off 2 (peak 3/peak 1) | |
| Esophagus | 0.65 | 2.0 |
| Stomach | 1.85 | 2.0 |
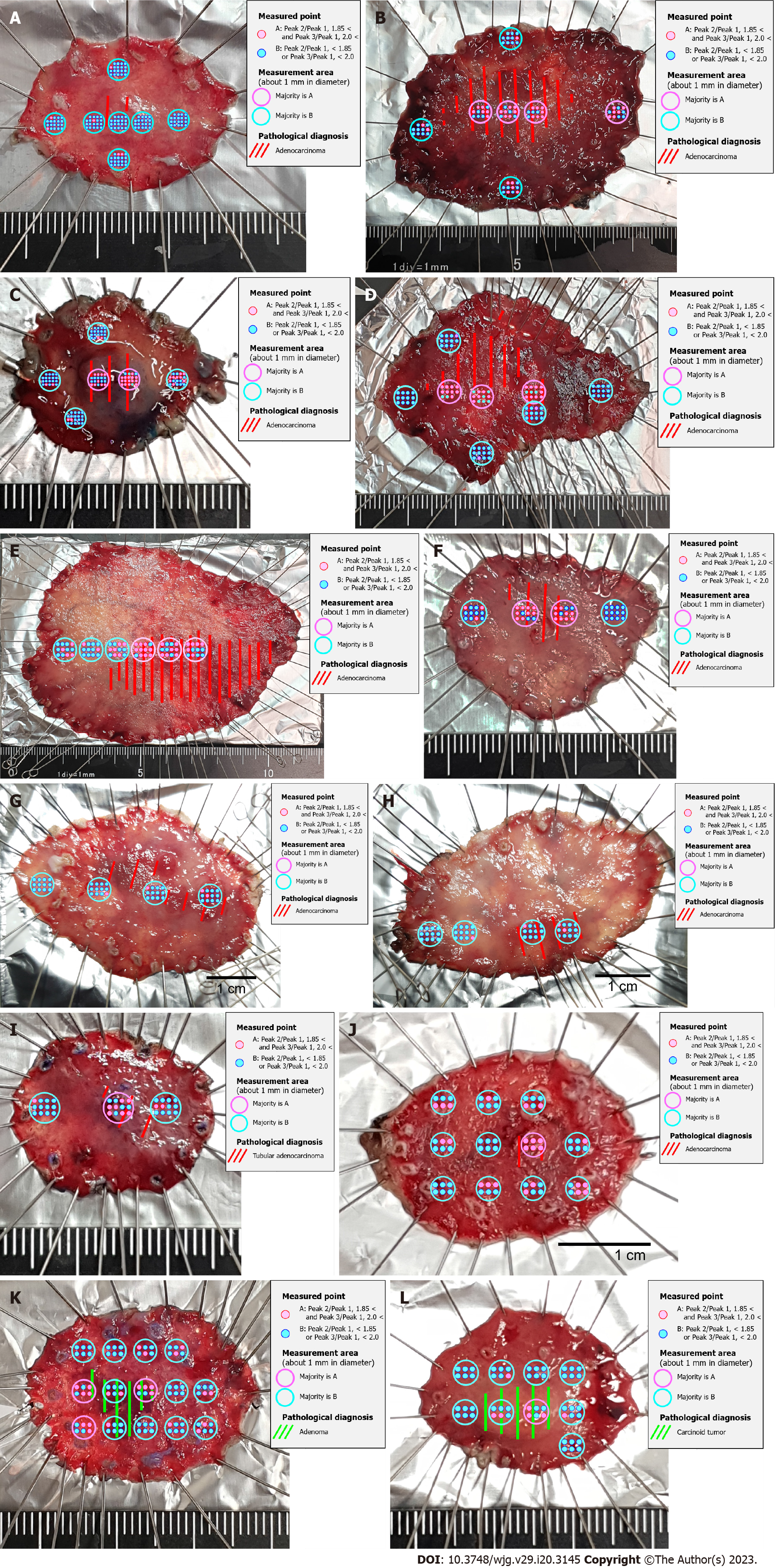
For gastric AC, the scattered light intensity of the CH2 stretch was Peak 1, Phenylalanine ring breadth was Peak 2, and Amide I alpha helix was Peak 3 (Table 4). The ratio of Peak 2/Peak 1 was defined as Cut-off 1 and Peak 3/Peak 1 was defined as Cut-off 2 (Table 5). The site with more than half of the measurement points with Cut-off 1 greater than 1.85 and Cut-off 2 greater than 2.0 was determined to be the best match for the histopathological diagnosis of gastric AC (Figure 3).
Patients with gastrointestinal cancers affecting the esophagus and stomach have symptoms, such as difficulty in swallowing, heartburn, and fullness[6]. Contrast examination or endoscopy is often performed in patients with a clinical evaluation suggestive of esophageal or gastric cancer[6]. Histopathological evaluation of the tissues obtained via endoscopic biopsy can be used to confirm the diagnosis[3,4,6]. In recent years, the accuracy of endoscopic finding-based diagnoses has dramatically improved. One of the reasons for this is the better quality of images obtained through high-resolution camera optics[3,6].
It is highly possible that the ability to diagnose cancer will improve further if it is feasible to depict what cannot be delineated using conventional equipment by making the image richer in features. Another reason is the use of image classification algorithms based on artificial intelligence[13,16,19]. By recognizing and patterning endoscopic images using artificial intelligence, the existence of lesions is clarified, and diagnosis by doctors is supported. Since the diagnosis using artificial intelligence evaluates endoscopic images morphologically, it is possible to prevent the lesion from being overlooked[13,16]. However, its diagnostic ability is not higher than that of an experienced doctor. Another drawback is that diagnostic ability is greatly affected by the quality of the endoscopic image[5,9,14].
As mentioned above, the accuracy of endoscopic diagnosis of lesions in the gastrointestinal tract has greatly improved; however, it is still at the morphological interpretation level[6,7]. Histopathological diagnosis, which is the current definitive diagnostic method, is a form of morphological evaluation; moreover, qualitative information that assists in morphological evaluation is required to improve diagnostic accuracy beyond the existing practice. RS can be used to add qualitative information to the morphological information[2,20]. Short et al[1] published a technique for diagnosing lung cancer using bronchoscopy combined with RS. Lui et al[21] reported a method for diagnosing skin cancer using RS. Krishna et al[22] reported a method for diagnosing oral cancer using RS. Jermyn et al[20] reported that RS can be used to identify lesion areas during brain surgery. Furthermore, Bergholt et al[6] presented a diagnostic technique for esophageal disease in the gastrointestinal tract using RS, and published diagnostic techniques for gastric cancer[4]. Duraipandian et al[3] reported a diagnostic technique for gastric cancer using RS. Molckovsky et al[2] reported a diagnostic technique for colon cancer using RS. However, a standard method for analyzing living organisms and biological samples using RS is yet to be developed.
In this study, we selected 15 Raman shifts corresponding to constituent molecules in living tissues. The analysis results of the scattered light intensity of Raman shift of Skeletal C-C stretch, Phenylalanine C-C twist, and CH3 deformation in SCC of the esophagus, and Raman shift of CH2 stretch, Phenylalanine ring breadth, and Amide I alpha-helix in gastric AC. The results of the analysis of the scattered light intensity of the shift were in good agreement with the range of morphological cancer diagnoses offered by histopathological evaluation. SCC of the esophagus and gastric AC are distinct types of cancers with different biological characteristics. Therefore, a significant Raman shift is expected to differ in the analysis conducted using RS. The technique presented in this study has the potential to provide qualitative evaluation in addition to the morphological evaluation currently used. In particular, it may offer valuable information regarding lesions that are challenging to assess using endoscopic or histopathological diagnoses alone.
However, at some sites, the results of this technique did not match the histopathological diagnoses. The limitations of this study include the small number of samples analyzed (n = 18), which included both the esophagus and stomach. Hence, there is no doubt that more samples must be analyzed in the future to confirm the accuracy of this technique. Using this technique, molecular information not solely dependent on morphological interpretation can be obtained. Furthermore, it may indicate the degree of mucosal abnormality from normal to cancerous. Evaluation of precancerous tissues may enable highly accurate preventive medicine. We aim to further advance this research and establish a method for predicting the current and future states of the gastrointestinal mucosa with high accuracy by clarifying the correlation between the Raman-scattered light intensity and the tissue state at each Raman shift.
Based on the results, it was concluded that RS could accurately identify esophageal SCC and gastric AC by analyzing the scattered light intensities of the 15 types of Raman shifts. The optimum combinations of Raman shifts were identified as Peak 1, Peak 2, and Peak 3 for both cancer types, with specific cut-off values for each peak. Although it is currently used in endoscopic treatment, the qualitative diagnosis of lesions, risk of residual lesions, etc., can be confirmed in a short time by analyzing excised specimens, and treatment can be provided as necessary.
Cancer diagnosis plays an important role in patient management, many researchers focused on the cancer detection, and novel, rapid, and label-free techniques are being developed for routine clinical practice.
To address the issue of using Raman spectroscopy (RS) to evaluate biological samples is interference of autofluorescence.
Our study is to evaluate the use of Raman-scattered light spectrum for detecting endoscopically resected specimens of esophageal squamous cell carcinoma (SCC) and gastric adenocarcinoma (AC).
We created a Raman device, which is suitable for observing living tissues, and we attempted to acquire Raman-scattered light spectra in endoscopically resected specimens of six esophageal tissues and 12 gastric tissues. Furthermore, we performed a correlation between the obtained Raman scattered light spectra and histopathological diagnosis.
We obtained Raman scattered light spectra from all six esophageal and 12 gastric specimens successfully, and developed an algorithm to identify the range of esophageal SCC and gastric AC with an accuracy close to that of histopathological diagnoses.
In this study, we utilized RS to rapidly detect cancer in resected esophageal and stomach specimens, providing information beyond morphology.
By providing detailed molecular level data, RS can provide a more comprehensive understanding of disease processes and aid in accurate diagnosis.
| 1. | Short MA, Lam S, McWilliams A, Zhao J, Lui H, Zeng H. Development and preliminary results of an endoscopic Raman probe for potential in vivo diagnosis of lung cancers. Opt Lett. 2008;33:711-713. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 86] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 2. | Molckovsky A, Song LM, Shim MG, Marcon NE, Wilson BC. Diagnostic potential of near-infrared Raman spectroscopy in the colon: differentiating adenomatous from hyperplastic polyps. Gastrointest Endosc. 2003;57:396-402. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 219] [Cited by in RCA: 164] [Article Influence: 7.1] [Reference Citation Analysis (3)] |
| 3. | Duraipandian S, Sylvest Bergholt M, Zheng W, Yu Ho K, Teh M, Guan Yeoh K, Bok Yan So J, Shabbir A, Huang Z. Real-time Raman spectroscopy for in vivo, online gastric cancer diagnosis during clinical endoscopic examination. J Biomed Opt. 2012;17:081418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 89] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 4. | Bergholt MS, Zheng W, Lin K, Ho KY, Teh M, Yeoh KG, So JB, Huang Z. Raman endoscopy for in vivo differentiation between benign and malignant ulcers in the stomach. Analyst. 2010;135:3162-3168. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 55] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 5. | Chakraborty A, Ghosh A, Barui A. Advances in surface-enhanced Raman spectroscopy for cancer diagnosis and staging. J Raman Spectrosc. 2020;51:7-36. [DOI] [Full Text] |
| 6. |
Bergholt MS, Zheng W, Ho KY, Yeoh KG, Teh M, So JBY, Huang Z.
Real-time depth-resolved Raman endoscopy for |
| 7. | Ito H, Uragami N, Miyazaki T, Yang W, Issha K, Matsuo K, Kimura S, Arai Y, Tokunaga H, Okada S, Kawamura M, Yokoyama N, Kushima M, Inoue H, Fukagai T, Kamijo Y. Highly accurate colorectal cancer prediction model based on Raman spectroscopy using patient serum. World J Gastrointest Oncol. 2020;12:1311-1324. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 9] [Cited by in RCA: 14] [Article Influence: 2.3] [Reference Citation Analysis (4)] |
| 8. | Wang K, Qiu Y, Wu C, Wen ZN, Li Y. Surface-enhanced Raman spectroscopy and multivariate analysis for the diagnosis of oral squamous cell carcinoma. J Raman Spectrosc. 2023;54:355-362. [DOI] [Full Text] |
| 9. | Lei J, Yang D, Li R, Dai Z, Zhang C, Yu Z, Wu S, Pang L, Liang S, Zhang Y. Label-free surface-enhanced Raman spectroscopy for diagnosis and analysis of serum samples with different types lung cancer. Spectrochim Acta A Mol Biomol Spectrosc. 2021;261:120021. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 29] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 10. | Yue X, Wang Y, Guo D, He G, Sui C, Qu Z, Zhao Y, Liu X. Surface-enhanced Raman spectroscopy is an accurate technique in diagnosis of breast cancer: A meta-analysis. J Raman Spectrosc. 2022;53:2058-2067. [DOI] [Full Text] |
| 11. | Falamas A, Faur CI, Baciut M, Rotaru H, Chirila M, Cinta Pinzaru S, Hedesiu M. Raman spectroscopic characterization of saliva for the discrimination of oral squamous cell carcinoma. Anal Lett. 2020;54. [RCA] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 3] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 12. | Zhang H, Cheng C, Gao R, Yan Z, Zhu Z, Yang B, Chen C, Lv X, Li H, Huang Z. Rapid identification of cervical adenocarcinoma and cervical squamous cell carcinoma tissue based on Raman spectroscopy combined with multiple machine learning algorithms. Photodiagnosis Photodyn Ther. 2021;33:102104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 13. | Zhang L, Wu Y, Zheng B, Su L, Chen Y, Ma S, Hu Q, Zou X, Yao L, Yang Y, Chen L, Mao Y, Ji M. Rapid histology of laryngeal squamous cell carcinoma with deep-learning based stimulated Raman scattering microscopy. Theranostics. 2019;9:2541-2554. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 102] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 14. | Zhan Q, Li Y, Yuan Y, Liu J. The accuracy of Raman spectroscopy in the detection and diagnosis of oral cancer: A systematic review and meta-analysis. J Raman Spectrosc. 2020;51:2377-2397. [RCA] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 7] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 15. | Zheng C, Qing S, Wang J, Lü G, Li H, Lü X, Ma C, Tang J, Yue X. Diagnosis of cervical squamous cell carcinoma and cervical adenocarcinoma based on Raman spectroscopy and support vector machine. Photodiagnosis Photodyn Ther. 2019;27:156-161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 32] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 16. | Yu M, Yan H, Xia J, Zhu L, Zhang T, Zhu Z, Lou X, Sun G, Dong M. Deep convolutional neural networks for tongue squamous cell carcinoma classification using Raman spectroscopy. Photodiagnosis Photodyn Ther. 2019;26:430-435. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 41] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 17. | Bögözi T, Popp J, Frosch T. Fiber-enhanced Raman multi-gas spectroscopy: what is the potential of its application to breath analysis? Bioanalysis. 2015;7:281-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 18. | Wachsmann-Hogiu S, Weeks T, Huser T. Chemical analysis in vivo and in vitro by Raman spectroscopy--from single cells to humans. Curr Opin Biotechnol. 2009;20:63-73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 117] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 19. | Xia J, Zhu L, Yu M, Zhang T, Zhu Z, Lou X, Sun G. Dong M. Analysis and classification of oral tongue squamous cell carcinoma based on Raman spectroscopy and convolutional neural networks. J Mod Opt. 2020;67:481-489. [DOI] [Full Text] |
| 20. | Jermyn M, Mok K, Mercier J, Desroches J, Pichette J, Saint-Arnaud K, Bernstein L, Guiot MC, Petrecca K, Leblond F. Intraoperative brain cancer detection with Raman spectroscopy in humans. Sci Transl Med. 2015;7:274ra19. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 352] [Cited by in RCA: 420] [Article Influence: 42.0] [Reference Citation Analysis (0)] |
| 21. | Lui H, Zhao J, McLean D, Zeng H. Real-time Raman spectroscopy for in vivo skin cancer diagnosis. Cancer Res. 2012;72:2491-2500. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 244] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 22. | Krishna H, Majumder SK, Chaturvedi P, Sidramesh M, Gupta PK. In vivo Raman spectroscopy for detection of oral neoplasia: a pilot clinical study. J Biophotonics. 2014;7:690-702. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 43] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
Open-Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/
Provenance and peer review: Invited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Japan
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Gao SG, China S-Editor: Wang JJ L-Editor: A P-Editor: Liu JH