Published online Apr 1, 2004. doi: 10.3748/wjg.v10.i7.1075
Revised: November 12, 2003
Accepted: December 16, 2003
Published online: April 1, 2004
AIM: To investigate the resistance rate of Helicobacter pylori (H pylori ) to clarithromycin, metronidazole, amoxicillin and tetracycline to guide clinical practice, and to study the mechanism of H pylori resistant to clarithromycin.
METHODS: Thirty H pylori strains were isolated from the mucosa of peptic ulcer, gastric tumor and chronic gastritis patients, then the minimal inhibitory concentration (MIC) to clarithromycin, metronidazole, amoxicillin and tetracycline was evaluated by E-test method. The sequence analysis of PCR fragments was conducted in 23S rRNA gene of H pylori resistant to clarithromycin to get the resistance mechanism of the bacteria.
RESULTS: Among 30 H pylori strains, 7 cases were resistant to clarithromycin, 12 to metronidazole, 2 to tetracycline and no strain was found to be resistant to amoxicillin. The resistance rates were 23.3%, 40%, 6.7% and 0%, respectively. Three new mutation points were found to be related to the clarithromycin resistance in H pylori isolates, which were G2224A, C2245T and T2289C.
CONCLUSION: In northeast China, H pylori shows high resistance to metronidazole, while sensitive to amoxicillin. The mechanism of resistance to clarithromycin may be related to the mutation of G2224A, C2245T and T2289C in the 23S rRNA gene.
- Citation: Hao Q, Li Y, Zhang ZJ, Liu Y, Gao H. New mutation points in 23S rRNA gene associated with Helicobacter pylori resistance to clarithromycin in northeast China. World J Gastroenterol 2004; 10(7): 1075-1077
- URL: https://www.wjgnet.com/1007-9327/full/v10/i7/1075.htm
- DOI: https://dx.doi.org/10.3748/wjg.v10.i7.1075
H pylori plays an important role in the pathogenesis of many digestive diseases. The recurrence of peptic ulcer and the development of the gastric cancer are also due to H pylori infection[1,2]. In the different eradication therapy regimen including PPI and multiple antibiotics, cases of eradication failure due to resistance of H pylori to antibiotics have been reported keeping increasing. The resistance mechanism of H pylori in the former studies showed the mutation from A to G in 2143 and 2144 position of 23S rRNA gene. But in our study we found other three new mutation points in 23S rRNA gene related to the H pylori resistance to clarithromycin.
In the endoscopic examination of the patients with digestive symptoms, we collected such patients with peptic ulcer, chronic gastritis and gastric carcinoma as our subjects. Firstly, one piece of gastric antra mucosa biopsy specimen was obtained from the patients for the purpose of rapid urease enzyme test. Then two pieces of antra mucosa biopsy specimens were obtained from the same patient with H pylori infection diagnosed by the positive rapid urease enzyme test. The biopsy specimens were cut up in sterile plate and then cultured on the Columbia agar base with 100 mL/L no-fiber fresh rabbit blood and 3 mg/L bacitracin at 37 °C for 3-5 d under microaerobic conditions (50 mL/L O2, 100 mL/L CO2, 850 mL/L N2). The organisms were identified as H pylori by Gram stain morphology, colony morphology and positive urease, catalase and oxidase activities. The typical bacteria were subcultured to obtain the pure H pylori isolates.
The pure H pylori suspension of 1 McFrand unit was prepared with sterile 9 g/L natrium chloride, and spread onto the Mueller-Hinton agar base with 100 mL/L fresh rabbit blood. The minimal inhibitory concentration (MIC) of H pylori to different antibiotics was evaluated with E-test strips. Strains were considered resistant to clarithromycin, metronidazole, amoxicillin and tetracycline if the MIC was ≥ 8 μg/mL, 8 μg/mL, 2 μg/mL and 8 μg/mL respectively.
Three clarithromycin resistant H pylori isolates and one sensitive H pylori isolate were chosen, that is, No.13 (MIC 8 mg/L), No. 17(MIC 64 mg/L), No.22 (MIC > 256 mg/L) and No.33 (MIC 0.125 mg/L). The DNA was extracted from the bacteria with the phenol-chloroform extraction method. We designed primers according to the 23S rRNA gene sequence reported by Hiratsuka (GeneBank accession number U27270). The primers were synthesized by Shanghai Sangon corporation and the sequences were as follows: forward primer: 5’-CTG CAT GAA TGG CGT AAC GAG-3’(complementary to 23S rRNA gene sequence from 2 047 to 2 067);and reverse primer: 5’-GAG CGA CCG CCC CGA TCA AAC-3’ (complementary to 23S rRNA gene sequence from 2 327 to 2 347), which will generate a 301 bp product. PCR amplification reaction mixture (20 μL) contained 13.8 μL double distilled H2O, 2.5 μL 10 × PCR buffer, 2 μL dNTPs (2.5 mmol/L), 0.2 μL Ex-Taq polymerase (5u/μL), 1.5 μL primer (1:5 × ) and 0.5 μL DNA sample. PCR cycle conditions were 32 cycles of 94 °C for 40 s, 61.5 °C for 1 min,72 °C for 1 min after 94 °C for 4 min once at first, then followed by a final extension step at 72 °C for 7 min. The purification of PCR products was observed by electrophoresis on an 80 g/L polyacrylamide gel. Then PCR products were sent to Sangon Corporation to conduct the sequence analysis. The amplified 23s rRNA gene sequence of resistant H pylori was compared to that of sensitive ones in order to find out the difference between them.
There were 7 cases among 30 H pylori isolates resistant to clarithromycin (MICs were from 8 mg/L to 256 mg/L), 12 cases of the isolates was resistant to metronidazole (MICs were from 24 mg/L to 256 mg/L), 2 cases resistant to tetracycline (MICs were 16 mg/L and 32 mg/L) and no case resistant to amoxicillin. The resistance rates to the above 4 different antibiotics were 23.3%, 40%, 6.7% and 0%, respectively.
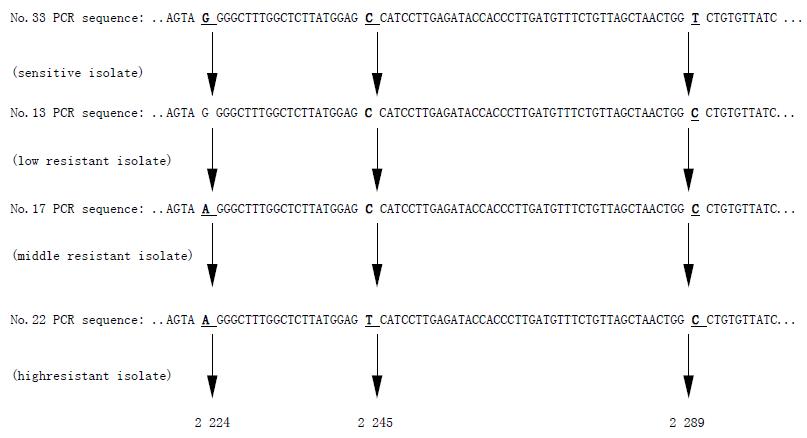
No.33 H pylori isolate was sensitive to clarithromycin, while No.13, No.17, and No.22 isolates were all resistant to clarithromycin. Point mutations appeared at three positions of the intended DNA fragments of clarithromycin resistant H pylori 23s rRNA gene. In comparison with the sequence of sensitive H pylori, No.13 resistant isolate with MIC 8 mg/L had one point mutation from T to C at 2289 (T2289C), No.17 isolate with MIC 64 mg/L had two point mutations which were G to A at 2 224 position (G2224A) and T to C at 2 289 position (T2289C), and No.22 isolate with the highest MIC >256 mg/L had three point mutations, and these were mutations from G to A at 2224 position (G2224A), from C to T at 2 245 position (C2245T) and the mutation from T to C at 2289 position (T2289C). With the increasing resistance of H pylori to clarithromycin, the number of point mutation were increased (Figure 1).
The prevalence of H pylori infection is about one-half of the world’s population[3], and still higher in the developing countries and low socio-economic populations[4-6]. It has been demonstrated that H pylori is an important etiologic factor of digestive diseases. H pylori infection is also correlated with cardio-cerebrovascular and pulmonary disease[7-10]. So eradication of H pylori becomes very important in the cure of the above diseases, especially peptic ulcer. But recently, cases of eradication failure become more and more due to the resistance of H pylori to antibiotics in the triple regimens. Tsuneoka et al[11] reported that H pylori strains from 19 cases (82.6%) out of 23 of failed eradication therapy became resistant to clarithromycin. Reports have shown that about 3-14%[12-14] of H pylori isolates are resistant to clarithromycin, 12-44%[13-16] are resistant to metronidazole. But no H pylori was found primary resistant to amoxicillin. The study showed that tetracycline resistance rate was ranging from 0 to 11%[13,16,17].
In our study, we isolated 30 H pylori strains from patients of peptic ulcer, chronic gastritis and gastric carcinoma and determined their MICs to clarithromycin, metronidazole, amoxicillin and tetracycline by E-test, respectively. Results showed a higher clarithromycin resistance than that in Europe (23.3%, 7/30). It is acknowledged that resistant clarithromycin H pylori strains become predominant because of antibiotics selective pressure. Clarithromycin frequently appearing in H pylori eradication therapy regimens led to the increasing of clarithromycin resistance. While in China, especially in northeast China, clarithromycin is rarely applied in clinic. How did the high resistance to clarithromycin generate? Is it associated with the extensive application of the other macrolide agents, such as erythromycin, azithromycin, and so on? Dose cross-resistance to macrolide exist among H pylori strains? Midolo and Saika et al[18,19] have demonstrated the existence of this phenomenon. But no study on this aspect has been done in China. The high resistance to clarithromycin should be considered in selecting H pylori eradication therapy regimen.
Metronidazole has been extensively used to treat anaerobic and parasitic infections for a long time, especially in the developing countries. It has been demonstrated that previous exposure of H pylori to metronidazole in vivo results in the emergence of resistant strains. Metronidazole resistant rate was reported ranging from 12% to 44%, similar to that in this study (40%). But it was also reported that metronidazole resistance as determined by E-test was significantly higher than that determined by agar dilution. Houben et al[20] found whether resistant to metronidazole could not affect the therapy outcome of OMC (omeprazole, metronidazole, clarithromycin) regimen. While in another study[21], it showed that the eradication rates were higher than the sensitive rates compared with resistant strains.
Almost all the studies in vitro showed high susceptibility of H pylori to amoxicillin. This study had the same results. In 30 isolates H pylori strains, only 1 strain’s MIC to amoxicillin was 4 μg/mL, and other MICs were all very low, between < 0.016-0.75 μg/mL. The influence of gastric acid on bactericidal effects was little against amoxicillin compared with macrolide. Furthermore, its activity in vivo is considerably enhanced when given concomitantly with proton pump inhibitors (PPI)[22]. So amoxicillin is still a selection of optimal drugs for the eradication of H pylori. Tetracycline is seldom applied in clinical practice at present. So studies on it were also rare. Perhaps due to less application, tetracycline resistance was very low, similar to this study. It was reported that the effect of the combination of macrolide agents with tetracycline was favorable[23].
The mechanism of H pylori resistance to clarithromycin was demonstrated to be associated with the mutation of A2143G or A2144G in 23S rRNA gene[24]. Some other mutation points were also reported after that, such as A2142C, A2143C, A2143T, A2115C and G2141[25]. Fontana[26] found that T2717C mutation was related to the H pylori resistance. It was known that the genetic character of H pylori in the different area was different[27,28], so did the genetic character of resistance H pylori strains[29]. Using gene segment analysis directly to determine the mutation position has still not been reported in China yet. We found in our study that the number of mutation points increased with the MIC of the resistant strains. The higher MIC is, the more mutation points are. This result has not been reported in other studies. The further investigation is still required to demonstrate the geographic differences existing in H pylori from different countries.
| 1. | Walsh JH, Peterson WL. The treatment of Helicobacter pylori infection in the management of peptic ulcer disease. N Engl J Med. 1995;333:984-991. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 257] [Cited by in RCA: 247] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 2. | Wang RT, Wang T, Chen K, Wang JY, Zhang JP, Lin SR, Zhu YM, Zhang WM, Cao YX, Zhu CW. Helicobacter pylori infection and gastric cancer: evidence from a retrospective cohort study and nested case-control study in China. World J Gastroenterol. 2002;8:1103-1107. [PubMed] |
| 3. | Dunn BE, Cohen H, Blaser MJ. Helicobacter pylori. Clin Microbiol Rev. 1997;10:720-741. [PubMed] |
| 4. | Bener A, Uduman SA, Ameen A, Alwash R, Pasha MA, Usmani MA, AI-Naili SR, Amiri KM. Prevalence of Helicobacter pylori infection among low socio-economic workers. J Commun Dis. 2002;34:179-184. [PubMed] |
| 5. | Wang KJ, Wang RT. [Meta-analysis on the epidemiology of Helicobacter pylori infection in China]. Zhonghua Liuxingbingxue Zazhi. 2003;24:443-446. [PubMed] |
| 6. | Strnad M, Presecki V, Babus V, Turek S, Dominis M, Kalenić S, Hebrang A, Katicić M. [Epidemiology of Helicobacter pylori infection]. Lijec Vjesn. 2002;124 Suppl 1:5-9. [PubMed] |
| 7. | Mendall MA, Goggin PM, Molineaux N, Levy J, Toosy T, Strachan D, Camm AJ, Northfield TC. Relation of Helicobacter pylori infection and coronary heart disease. Br Heart J. 1994;71:437-439. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 462] [Cited by in RCA: 427] [Article Influence: 13.3] [Reference Citation Analysis (1)] |
| 8. | Pasceri V, Cammarota G, Patti G, Cuoco L, Gasbarrini A, Grillo RL, Fedeli G, Gasbarrini G, Maseri A. Association of virulent Helicobacter pylori strains with ischemic heart disease. Circulation. 1998;97:1675-1679. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 217] [Cited by in RCA: 187] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 9. | Markus HS, Mendall MA. Helicobacter pylori infection: a risk factor for ischaemic cerebrovascular disease and carotid atheroma. J Neurol Neurosurg Psychiatry. 1998;64:104-107. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 84] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 10. | Roussos A, Philippou N, Gourgoulianis KI. Helicobacter pylori infection and respiratory diseases: a review. World J Gastroenterol. 2003;9:5-8. [PubMed] |
| 11. | Tsuneoka H, Takaba M, Nagatomi Y, Mori K, Matsumoto T, Honda T. [Sensitivity of Helicobacter pylori to amoxicillin and clarythromycin with special reference to eradication therapy]. Kansenshogaku Zasshi. 1998;72:335-341. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 12. | Franzin L, Pennazio M, Cabodi D, Paolo Rossini F, Gioannini P. Clarithromycin and amoxicillin susceptibility of Helicobacter pylori strains isolated from adult patients with gastric or duodenal ulcer in Italy. Curr Microbiol. 2000;40:96-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 10] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 13. | Taylor DE, Jiang Q, Fedorak RN. Antibiotic susceptibilities of Helicobacter pylori strains isolated in the Province of Alberta. Can J Gastroenterol. 1998;12:295-298. [PubMed] |
| 14. | Savarino V, Zentilin P, Pivari M, Bisso G, Raffaella Mele M, Bilardi C, Borro P, Dulbecco P, Tessieri L, Mansi C. The impact of antibiotic resistance on the efficacy of three 7-day regimens against Helicobacter pylori. Aliment Pharmacol Ther. 2000;14:893-900. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 38] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 15. | Osato MS, Reddy R, Reddy SG, Penland RL, Graham DY. Comparison of the Etest and the NCCLS-approved agar dilution method to detect metronidazole and clarithromycin resistant Helicobacter pylori. Int J Antimicrob Agents. 2001;17:39-44. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 90] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 16. | Ani AE, Malu AO, Onah JA, Queiroz DM, Kirschner G, Rocha GA. Antimicrobial susceptibility test of Helicobacter pylori isolated from Jos, Nigeria. Trans R Soc Trop Med Hyg. 1999;93:659-661. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 19] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 17. | Vasquez A, Valdez Y, Gilman RH, McDonald JJ, Westblom TU, Berg D, Mayta H, Gutierrez V. Metronidazole and clarithromycin resistance in Helicobacter pylori determined by measuring MICs of antimicrobial agents in color indicator egg yolk agar in a miniwell format. The Gastrointestinal Physiology Working Group of Universidad Peruana Cayetano Heredia and the Johns Hopkins University. J Clin Microbiol. 1996;34:1232-1234. [PubMed] |
| 18. | Saika T, Kobayashi I, Fujioka T, Nasu M, Okamoto R, Inoue M. [A mechanism of clarithromycin resistance in Helicobacter pylori]. Kansenshogaku Zasshi. 1998;72:918-923. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 19. | Midolo PD, Bell JM, Lambert JR, Turnidge JD, Grayson ML. Antimicrobial resistance testing of Helicobacter pylori: a comparison of Etest and disk diffusion methods. Pathology. 1997;29:411-414. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 31] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 20. | Houben MH, Hensen EF, Rauws EA, Hulst RW, Hoff BW, Ende AV, Kate FJ, Tytgat GN. Randomized trial of omeprazole and clarithromycin combined with either metronidazole or amoxycillin in patients with metronidazole-resistant or -susceptible Helicobacter pylori strains. Aliment Pharmacol Ther. 1999;13:883-889. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 22] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 21. | Moayyedi P, Ragunathan PL, Mapstone N, Axon AT, Tompkins DS. Relevance of antibiotic sensitivities in predicting failure of omeprazole, clarithromycin, and tinidazole to eradicate Helicobacter pylori. J Gastroenterol. 1998;33 Suppl 10:62-65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 25] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 22. | Hirschl AM, Rotter ML. Amoxicillin for the treatment of Helicobacter pylori infection. J Gastroenterol. 1996;31 Suppl 9:44-47. [PubMed] |
| 23. | Bamba H, Kondo Y, Wong RM, Sekine S, Matsuzaki F. Minimum inhibitory concentration of various single agents and the effect of their combinations against Helicobacter pylori, as estimated by a fast and simple in vitro assay method. Am J Gastroenterol. 1997;92:659-662. [PubMed] |
| 24. | Stone GG, Shortridge D, Flamm RK, Versalovic J, Beyer J, Idler K, Zulawinski L, Tanaka SK. Identification of a 23S rRNA gene mutation in clarithromycin-resistant Helicobacter pylori. Helicobacter. 1996;1:227-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 68] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 25. | van Doorn LJ, Debets-Ossenkopp YJ, Marais A, Sanna R, Mégraud F, Kusters JG, Quint WG. Rapid detection, by PCR and reverse hybridization, of mutations in the Helicobacter pylori 23S rRNA gene, associated with macrolide resistance. Antimicrob Agents Chemother. 1999;43:1779-1782. [PubMed] |
| 26. | Fontana C, Favaro M, Minelli S, Criscuolo AA, Pietroiusti A, Galante A, Favalli C. New site of modification of 23S rRNA associated with clarithromycin resistance of Helicobacter pylori clinical isolates. Antimicrob Agents Chemother. 2002;46:3765-3769. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 60] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 27. | Mukhopadhyay AK, Kersulyte D, Jeong JY, Datta S, Ito Y, Chowdhury A, Chowdhury S, Santra A, Bhattacharya SK, Azuma T. Distinctiveness of genotypes of Helicobacter pylori in Calcutta, India. J Bacteriol. 2000;182:3219-3227. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 179] [Cited by in RCA: 178] [Article Influence: 6.8] [Reference Citation Analysis (1)] |
| 28. | Yu FJ, Wu DC, Kuo CH, Lu CY, Su YC, Lee YC, Lin SR, Liu CS, Jan CM, Wang WM. Diagnosis of Helicobacter pylori infection by stool antigen test in southern Taiwan. Kaohsiung J Med Sci. 2001;17:344-350. [PubMed] |
| 29. | Meyer JM, Silliman NP, Wang W, Siepman NY, Sugg JE, Morris D, Zhang J, Bhattacharyya H, King EC, Hopkins RJ. Risk factors for Helicobacter pylori resistance in the United States: the surveillance of H. pylori antimicrobial resistance partnership (SHARP) study, 1993-1999. Ann Intern Med. 2002;136:13-24. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 154] [Cited by in RCA: 151] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
Edited by Zhang JZ and Xu FM