Copyright: ©Author(s) 2026.
World J Gastroenterol. May 14, 2026; 32(18): 118267
Published online May 14, 2026. doi: 10.3748/wjg.v32.i18.118267
Published online May 14, 2026. doi: 10.3748/wjg.v32.i18.118267
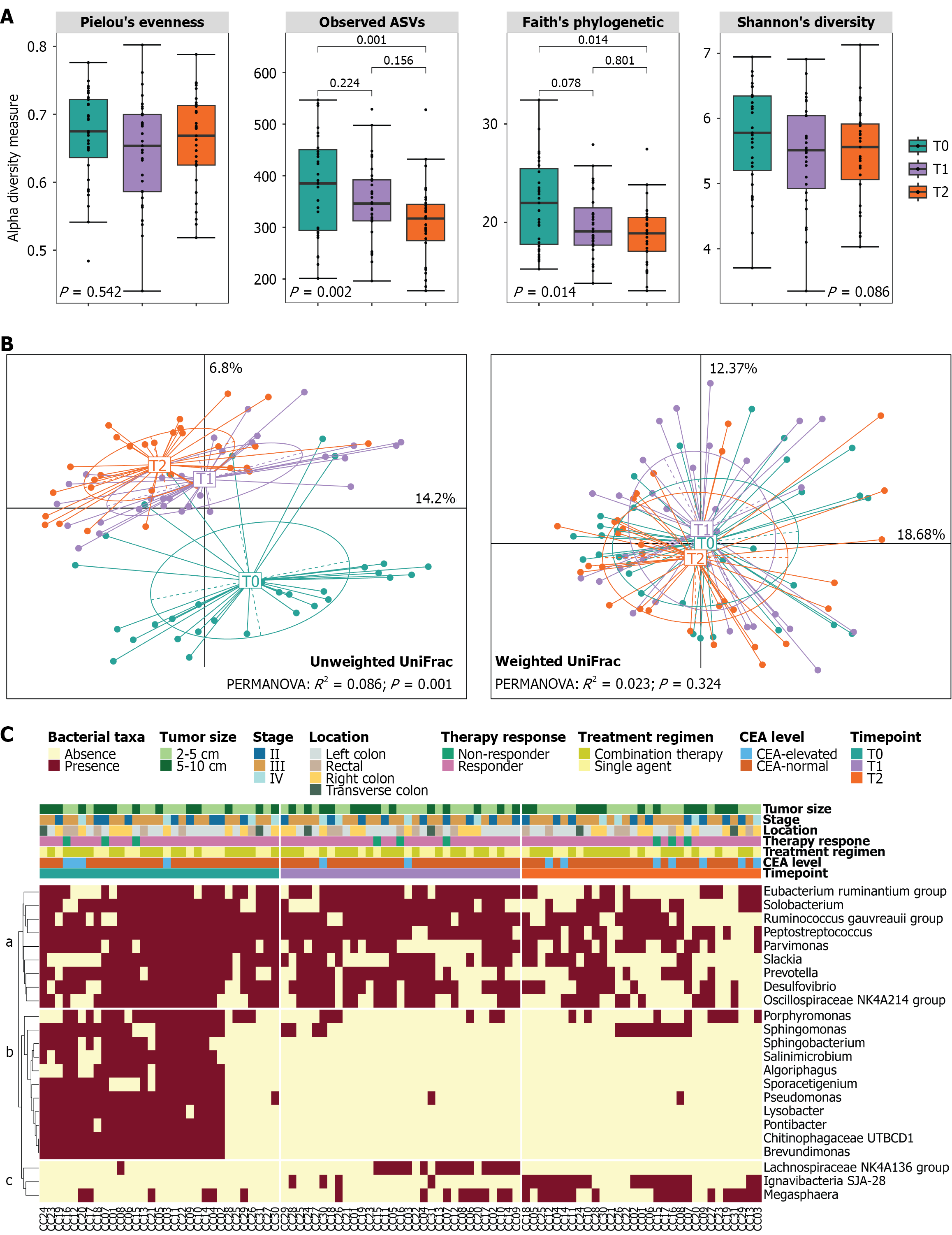
Figure 1 Microbial diversity of the gut microbiota in patients with colorectal cancer at diagnosis (T0), after surgery (T1), and after che
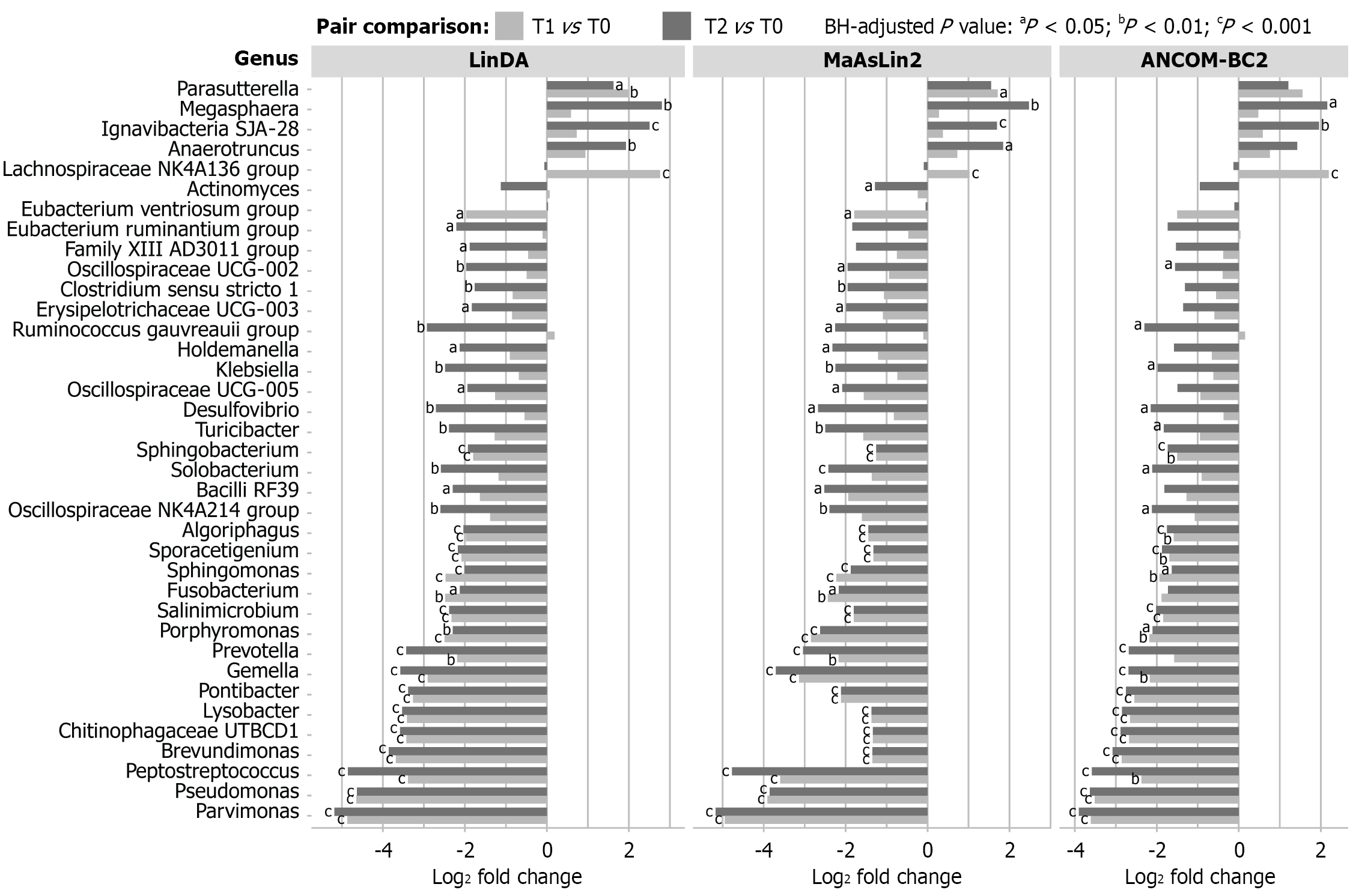
Figure 2 Temporal gut microbiota changes at the genus level during the treatment period.
Differentially abundant bacterial genera at post-treatment timepoints relative to T0 (diagnosis) were identified using LinDA, MaAsLin2, and ANCOM-BC2. Differences between T1 (after surgery) and T0 and between T2 (after chemotherapy) and T0 are shown as log2 fold changes. Only classified genera with significant differences between timepoints are presented. aAdjusted P < 0.05; bAdjusted P < 0.01; cAdjusted P < 0.001.
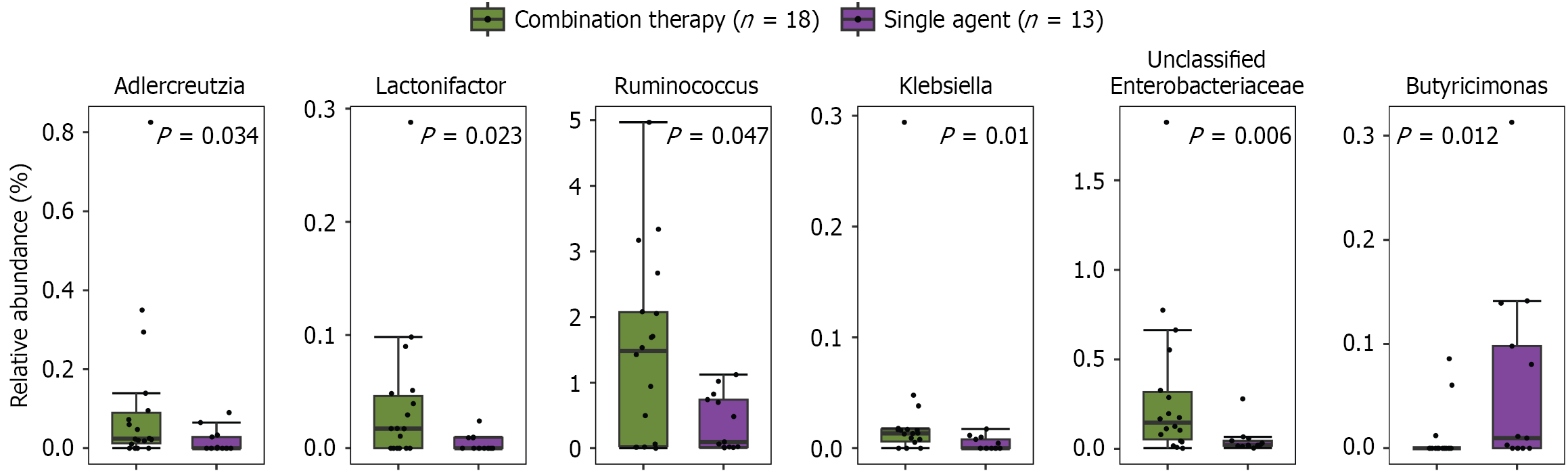
Figure 3 Bacterial genera with significantly different abundance between patient subgroups stratified by treatment regimen after chemotherapy (T2).
Only genera with an uncorrected P value < 0.05 (Mann-Whitney U test) are displayed.
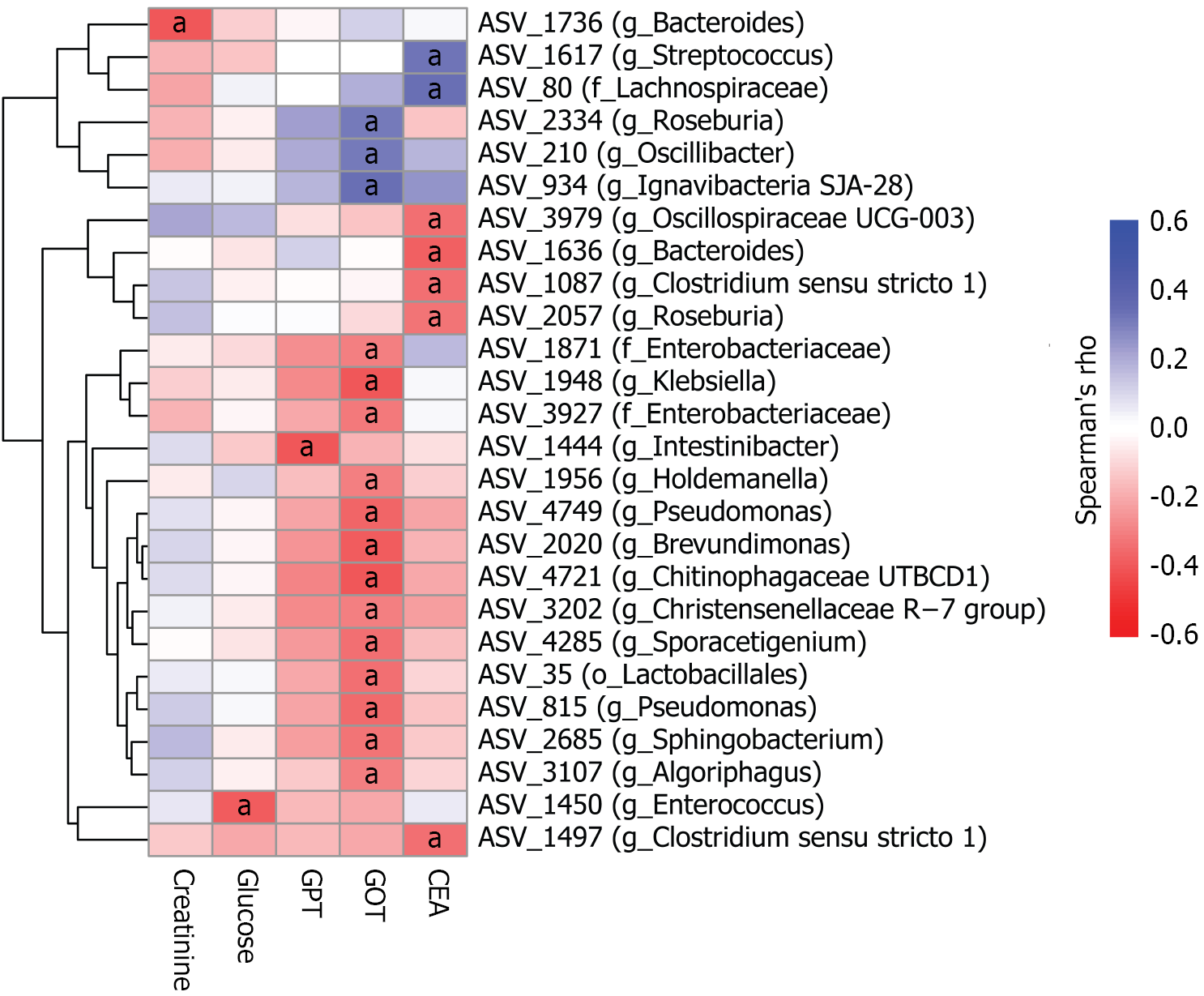
Figure 4 Heatmap showing the Spearman’s rank correlations between amplicon sequence variant abundance and serum biochemical markers.
Spearman’s ρ values were computed for each amplicon sequence variant (ASV) in relation to each biochemical marker. Multiple testing was adjusted using the Benjamini-Hochberg method, and only ASVs or markers with adjusted P values < 0.05 are displayed. Hierarchical clustering of ASVs was performed using the Euclidean distance metric. aAdjusted P < 0.05. ASVs were labeled with genus (g_), or family (f_), or order (o_) when the genus-level classification was unavailable. ASV: Amplicon sequence variant; GOT: Glutamic oxaloacetic transaminase; GPT: Glutamic pyruvic transaminase; CEA: Carcinoembryonic antigen.
- Citation: Le HTT, Le HX, Huyen DT, Tran TA, Quyen DV, Song LH, Tran TV, Thas O, Nhung PTT, Tran TTT. Longitudinal changes in the gut microbiota of Vietnamese patients with colorectal cancer undergoing surgery and chemotherapy. World J Gastroenterol 2026; 32(18): 118267
- URL: https://www.wjgnet.com/1007-9327/full/v32/i18/118267.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i18.118267