Copyright: ©Author(s) 2026.
World J Gastroenterol. Apr 14, 2026; 32(14): 111455
Published online Apr 14, 2026. doi: 10.3748/wjg.v32.i14.111455
Published online Apr 14, 2026. doi: 10.3748/wjg.v32.i14.111455
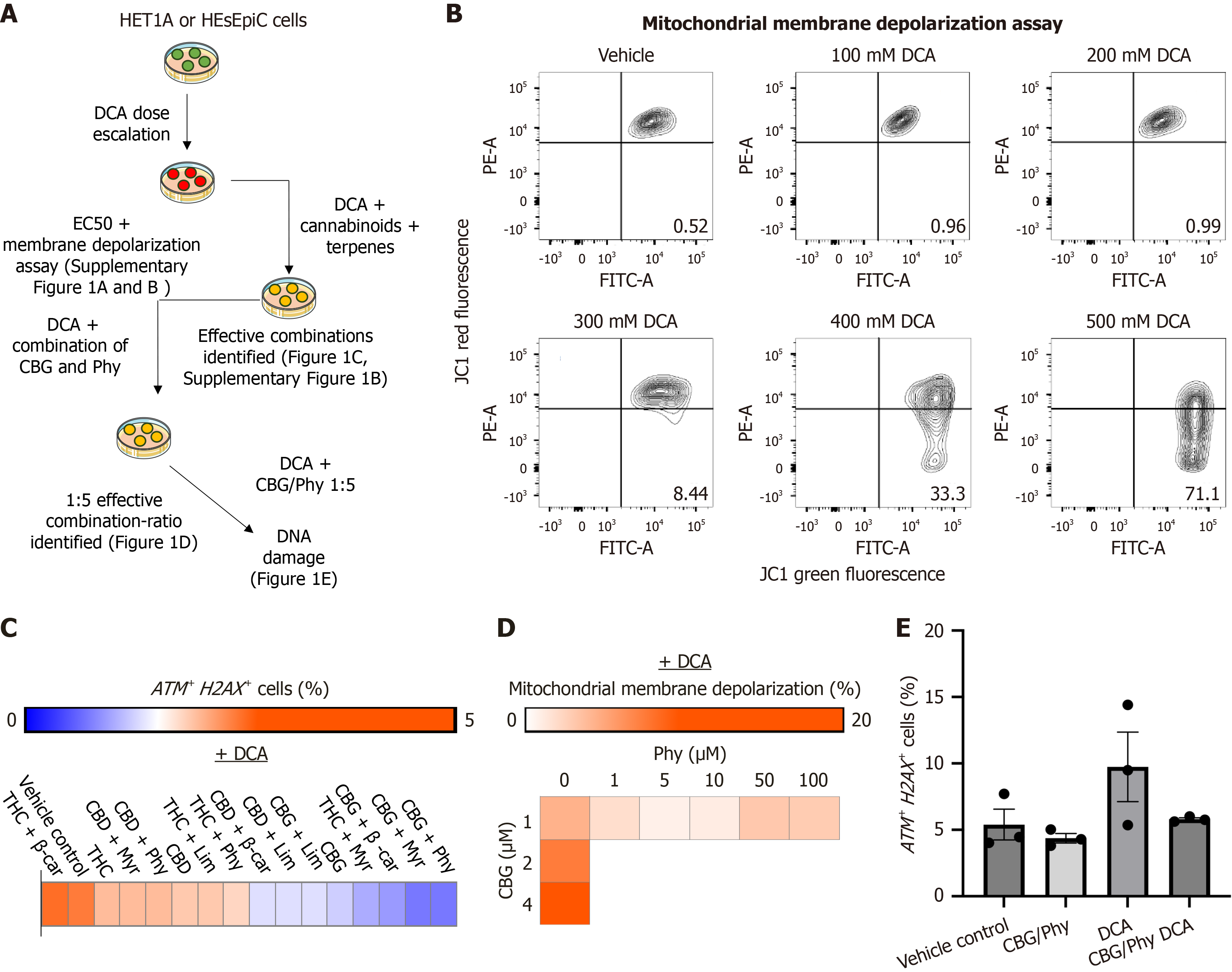
Figure 1 Cannabigerol and phytol combination ratio of 1:5 can reverse deoxycholic acid-induced mitochondrial depolarization and DNA damage in HET1A and human esophageal epithelial cells.
A: Schematic illustration of experimental design; B: Representative flow cytometry analyses of mitochondrial membrane potential in deoxycholic acid (DCA)-treated HET1A cells measured by red- and green-fluorescent changes performed using the MitoProbe JC-1 assay kit. Cells with normal mitochondrial membrane potential are depicted in the right upper corner (red), and the cells with reduced mitochondrial membrane potential are shown in the right lower corner (green); C: The heatmap illustrates the percentage of ATM+ H2AX+ cells in human esophageal epithelial cell cultures pretreated with diverse cannabinoids and terpenes at a 1:5 ratio, while exposed to 100 μmol/L DCA. Data were analyzed using the Muse Multi-color DNA damage kit (Cytek, United States); D: The heatmap illustrates the changes in the mitochondrial membrane potential in HET1A cells treated with varied concentrations and ratios of cannabigerol (CBG) and phytol (Phy) in the presence of 100 μmol/L DCA; E: The bar graph illustrates the percentage of ATM+ H2AX+ HET1A cells after treatment with CBG and Phy at 1:5 ratio in the presence of DCA. Data were analyzed using the Muse Multi-color DNA damage kit (Cytek, United States). HEsEpiC: Human esophageal epithelial cells; DCA: Deoxycholic acid; EC50: Lethal dose 50%; CBG: Cannabigerol; Phy: Phytol; THC: Tetrahydro cannabinoid; β-car: Β-caryophyllene; CBD: Cannabidiol; Myr: Myrcene; Lim: Limonene.
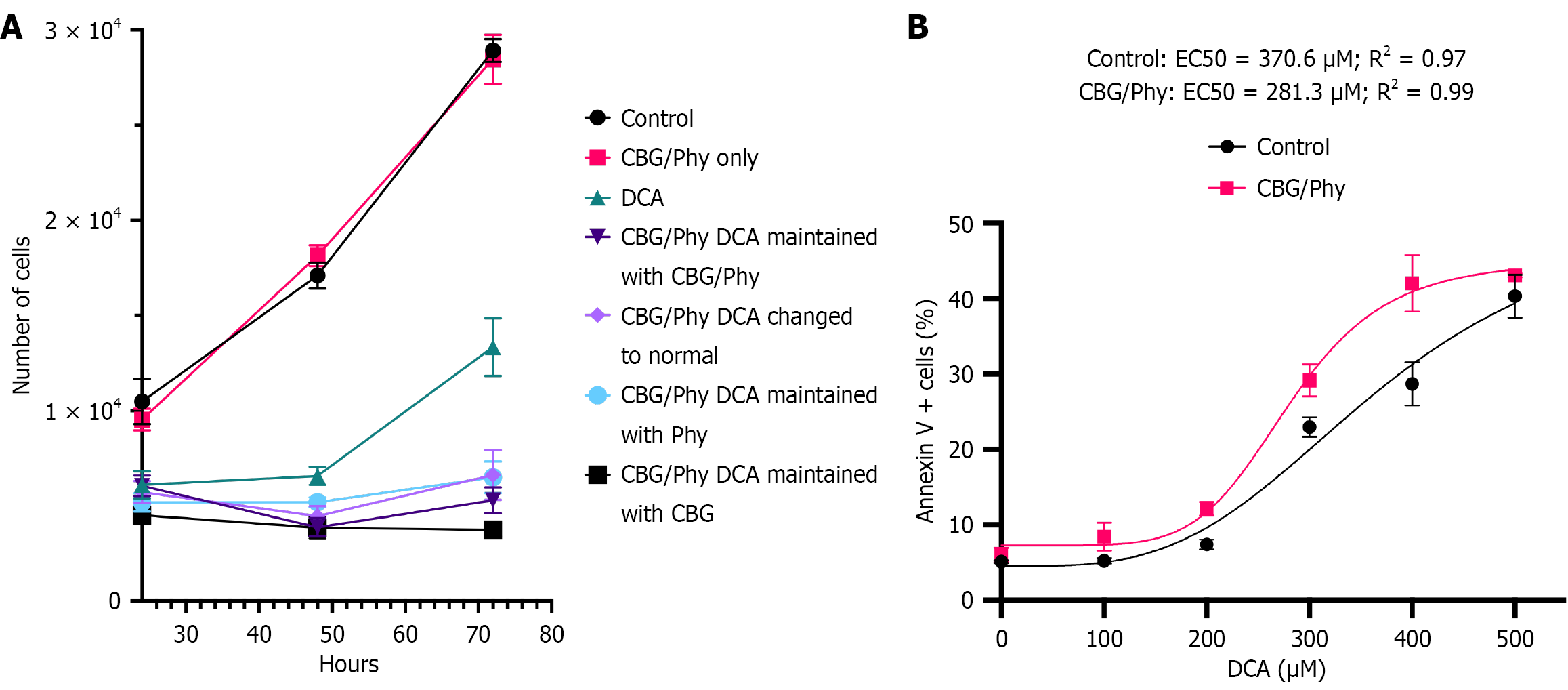
Figure 2 The effect of cannabigerol and phytol on the proliferation and apoptosis of deoxycholic acid-damaged esophageal cells.
A: XTT cell viability assay of HET1A cells subjected to 100 μmol/L deoxycholic acid (DCA) or vehicle control treatment for 24 hours in the presence or absence of cannabigerol (CBG) and phytol (Phy); B: The plot represents a non-linear regression analysis of the percentage of apoptotic cells in the HET1A cell line cultured with or without CBG/Phy pretreatment and exposed to DCA at concentrations ranging from 0 μmol/L to 500 μmol/L. DCA: Deoxycholic acid; EC50: Lethal dose 50%; CBG: Cannabigerol; Phy: Phytol.
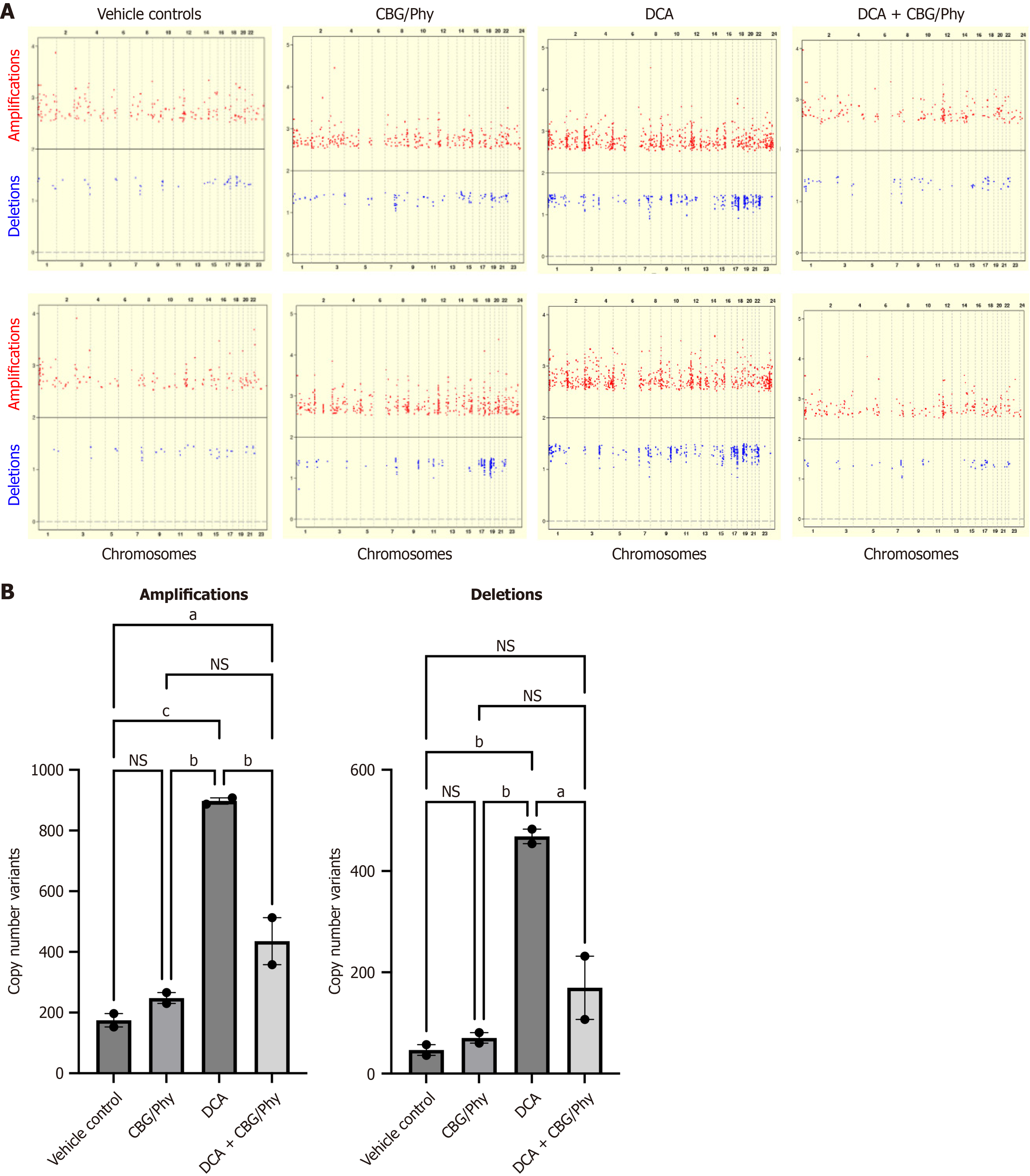
Figure 3 Cannabigerol and phytol at a 1:5 ratio reduce deoxycholic acid-induced genomic instability in normal esophageal epithelial cells.
A: Scatter plots of the whole genome amplifications and deletions in HET1A cells cultured in the presence of either vehicle control, 100 μmol/L deoxycholic acid (DCA), cannabigerol (CBG)/phytol admixture, or a combination of 100 μmol/L DCA and CBG/phytol admixture for 14 days. DNA extracted at day 14 was compared using precision medicine diversity arrays (Affymetrix) to the DNA of untreated cells collected at day 0; B: Bar graphs illustrate the quantitative analyses of the whole genome amplifications (left panel) and deletions (right panel). Data were analyzed using one-way analysis of variance with multiple comparisons. aP < 0.05. bP < 0.01. cP < 0.001. NS: Not significant; DCA: Deoxycholic acid; CBG: Cannabigerol; Phy: Phytol.
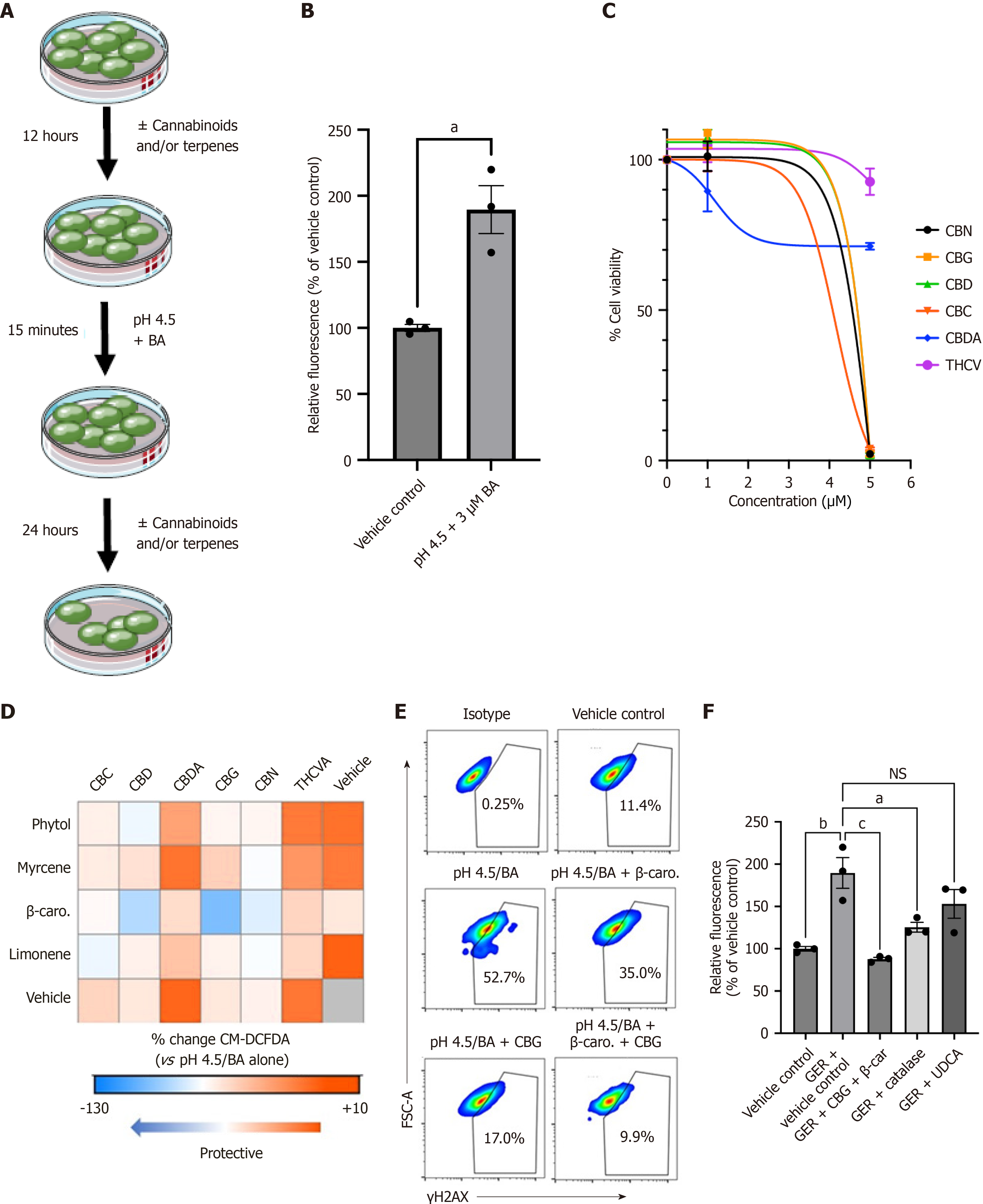
Figure 4 Cannabigerol and β-caryophyllene (1:5) reverse the damaging effect of the acid-containing gastroesophageal refluxate in metaplastic Barrett’s esophageal cells.
A: Experimental design schematic. Barrett’s esophageal cell line CP-A was treated with a vehicle control or cannabinoids in the presence or absence of terpenes for 12 hours, followed by an acute 15-minute exposure to a physiologically relevant low potential of hydrogen and bile acid cocktail, as described in the methods. The cells were either immediately analyzed for reactive oxygen species (ROS) via fluorescent detection of CM-DCFDA or recovered for 24 hours and analyzed for DNA damage using γ-H2AX flow cytometry; B: Bar graph illustrates the comparative analyses of ROS measured by CM-DCFDA fluorescence in CP-A cells following acute exposure to gastroesophageal reflux (GER). The samples were analyzed by flow cytometry, and the results are shown as percent increases relative to the vehicle control. The data were analyzed using an unpaired t-test with Welch’s correction (n = 3); C: Non-linearfit of the dose response cell viability analysis of CP-A cells after exposure to cannabinoids for 24 hours, analyzed using the MTS assay kit; D: Heat map shows changes in ROS induced by GER in CP-A cells pretreated with the cannabinoid and terpene combinations at a 1:5 ratio as determined by CM-DCFDA fluorescence. Data are expressed as the percent change when compared to GER alone; E: Representative flow cytometry analyses of DNA damage as determined by γ-H2AX fluorescence of CP-A cells pretreated with cannabigerol (CBG) and β-caryophyllene (β-car) following exposure to GER; F: Bar graph depicts the comparative analyses of ROS measured by CM-DCFDA fluorescence in CP-A cells exposed to GER and antioxidants (catalase), ursodeoxycholic acid, or the cannabinoid/terpene combination of CBG and β-car at a 1:5 ratio. The samples were analyzed by flow cytometry, and the results are shown as percent increases relative to the vehicle control. The data were analyzed using one-way analysis of variance with Šídák’s multiple comparisons test. aP < 0.05. bP < 0.01. cP < 0.001. NS: Not significant; pH: Potential of hydrogen; BA: Bile acids; CBN: Cannabinol; CBD: Cannabidiol; CBC: Cannabichromene; CBDA: Cannabidiolic acid; THCV: Tetrahydrocannabivarinic acid; FSC-A: Forward scatter area; CBG: Cannabigerol; β-car: Β-caryophyllene; GER: Gastroesophageal reflux; UDCA: Ursodeoxycholic acid.
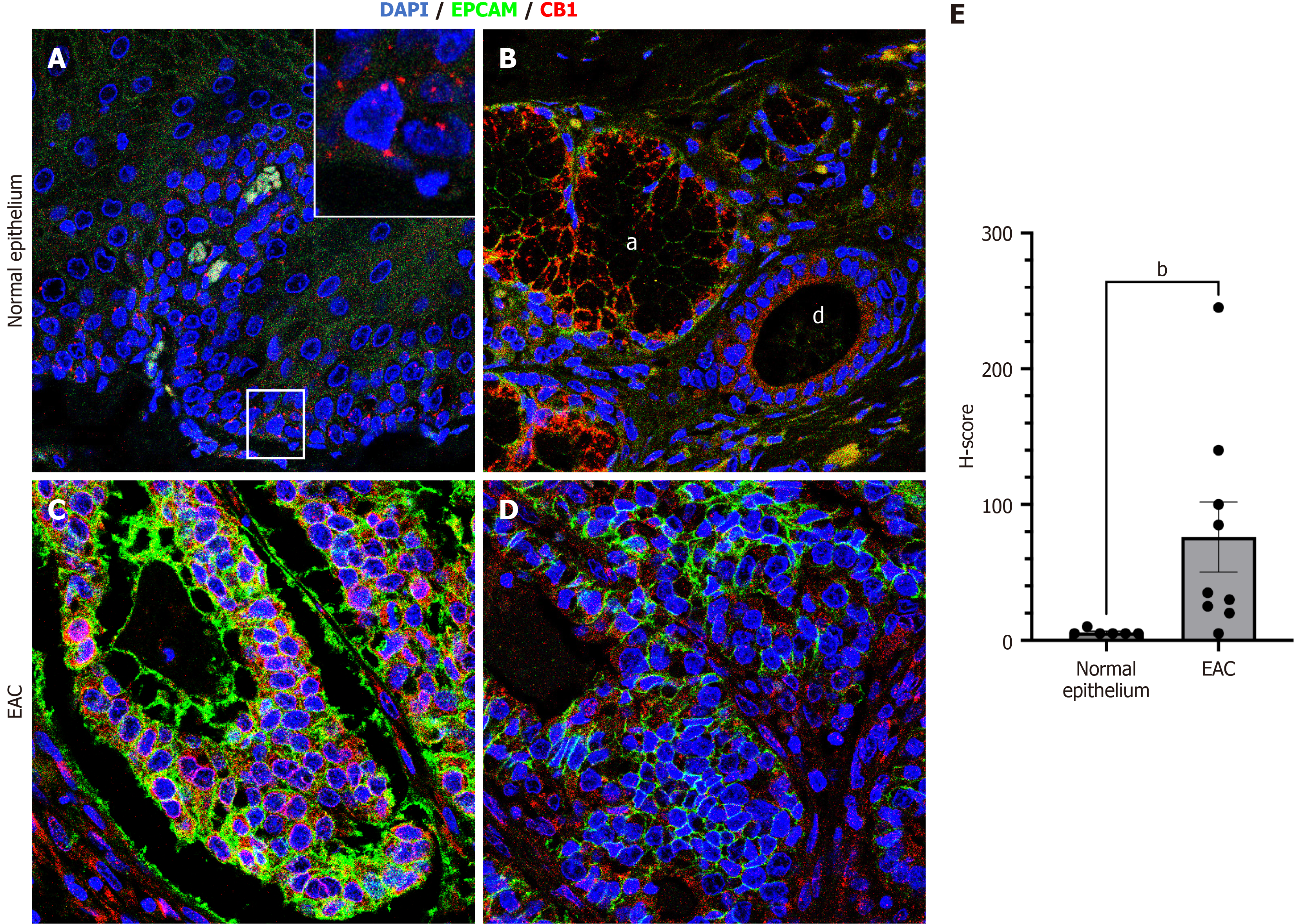
Figure 5 Expression of CB1 in human normal esophageal mucosa and esophageal adenocarcinoma assessed by confocal microscopy.
A: A representative immunofluorescence analysis of CB1 (red) and epithelial cell adhesion molecule (EPCAM) (green) expression in the normal human esophageal mucosa (magnification 63 ×). The inset highlights the CB1 expression in the basal epithelial layer; B: A representative immunofluorescence analysis depicting CB1 and EPCAM co-expression in the ductal epithelium of the submucosal gland (d) and acini of the submucosal gland (a); C and D: Representative immunofluorescence analyses of CB1 and EPCAM co-expression in two distinct human esophageal adenocarcinoma (EAC) samples; E: Bar graph represents a comparative H-score semiquantitative intensity analysis of CB1 expression in human normal esophageal mucosa and EAC. The data were analyzed using the Mann-Whitney U test. bP < 0.01. DAPI: 4’,6-diamidino-2-phenylindole; EPCAM: Epithelial cell adhesion molecule; EAC: Esophageal adenocarcinoma.
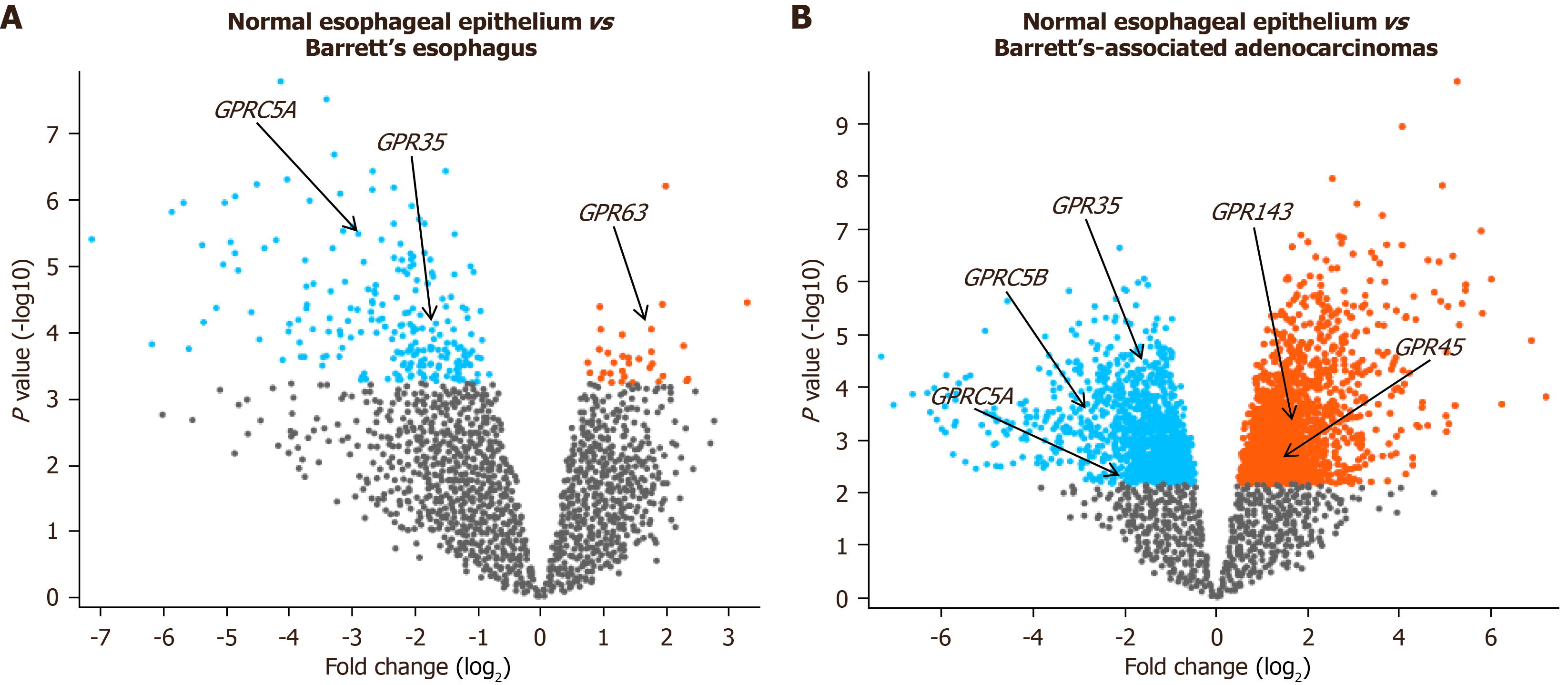
Figure 6 Dysregulation of G-protein coupled receptor gene expression in clinical Barrett’s esophagus and Barrett’s esophagus-associated esophageal adenocarcinoma.
A: The volcano represents differentially expressed genes in clinical Barrett’s esophagus (BE) (n = 8) compared to the normal esophageal epithelium (n = 8). The data were obtained from the GSE database (GSE1420). BE upregulated (blue) and downregulated (red) genes were identified using the GEO2R tool, based on log2 fold change > 0.9 and adjusted P values of < 0.05; B: The volcano represents differentially expressed genes in clinical BE-associated esophageal adenocarcinoma (EAC) (n = 8) compared to the normal esophageal epithelium (n = 8). The data were obtained from the GSE database (GSE1420). EAC upregulated (blue) and downregulated (red) genes were identified using the GEO2R tool, based on log2 fold change > 0.9 and adjusted P values of < 0.05.
- Citation: Goldman A, Gonzalez G, Karpova SA, Buon L, Shammas MA, Mashimo H, Frank MH, Frank NY. Optimal cannabinoid-terpene combination ratios suppress mutagenicity of gastric reflux in normal and metaplastic esophageal cells. World J Gastroenterol 2026; 32(14): 111455
- URL: https://www.wjgnet.com/1007-9327/full/v32/i14/111455.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i14.111455