Copyright: ©Author(s) 2026.
World J Gastroenterol. Apr 7, 2026; 32(13): 113985
Published online Apr 7, 2026. doi: 10.3748/wjg.v32.i13.113985
Published online Apr 7, 2026. doi: 10.3748/wjg.v32.i13.113985
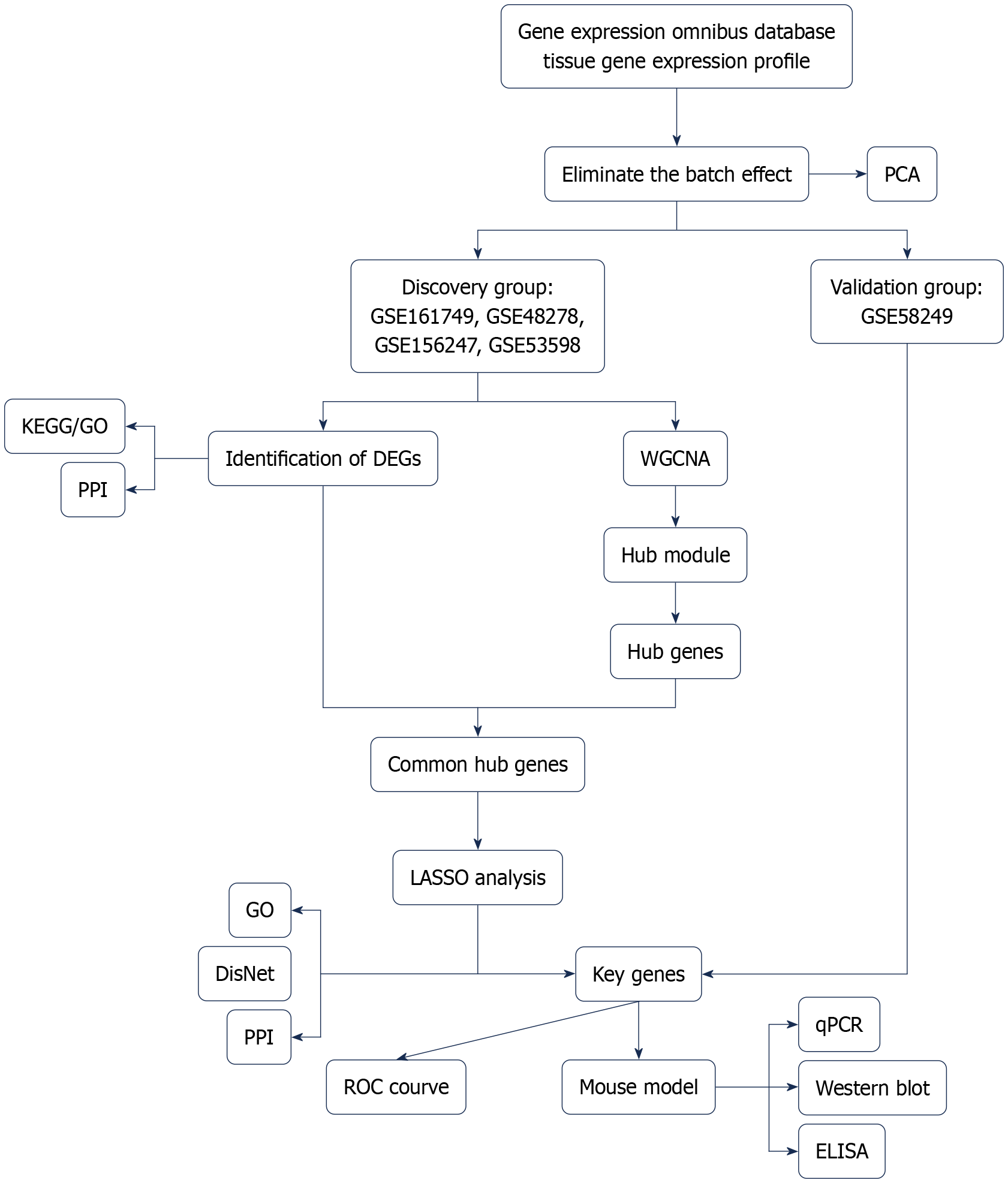
Figure 1 Flowchart for identifying biomarkers related to exercise in the population with metabolic dysfunction-associated steatotic liver disease, including data extraction, procession and analysis.
PCA: Principal component analysis; DEGs: Differentially expressed genes; WGCNA: Weighted gene co-expression network analysis; KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: Gene Ontology; PPI: Protein-protein interaction; LASSO: Least absolute shrinkage and selection operator; ROC: Receiver operating characteristic; ELISA: Enzyme-linked immunosorbent assay; qPCR: Quantitative polymerase chain reaction.
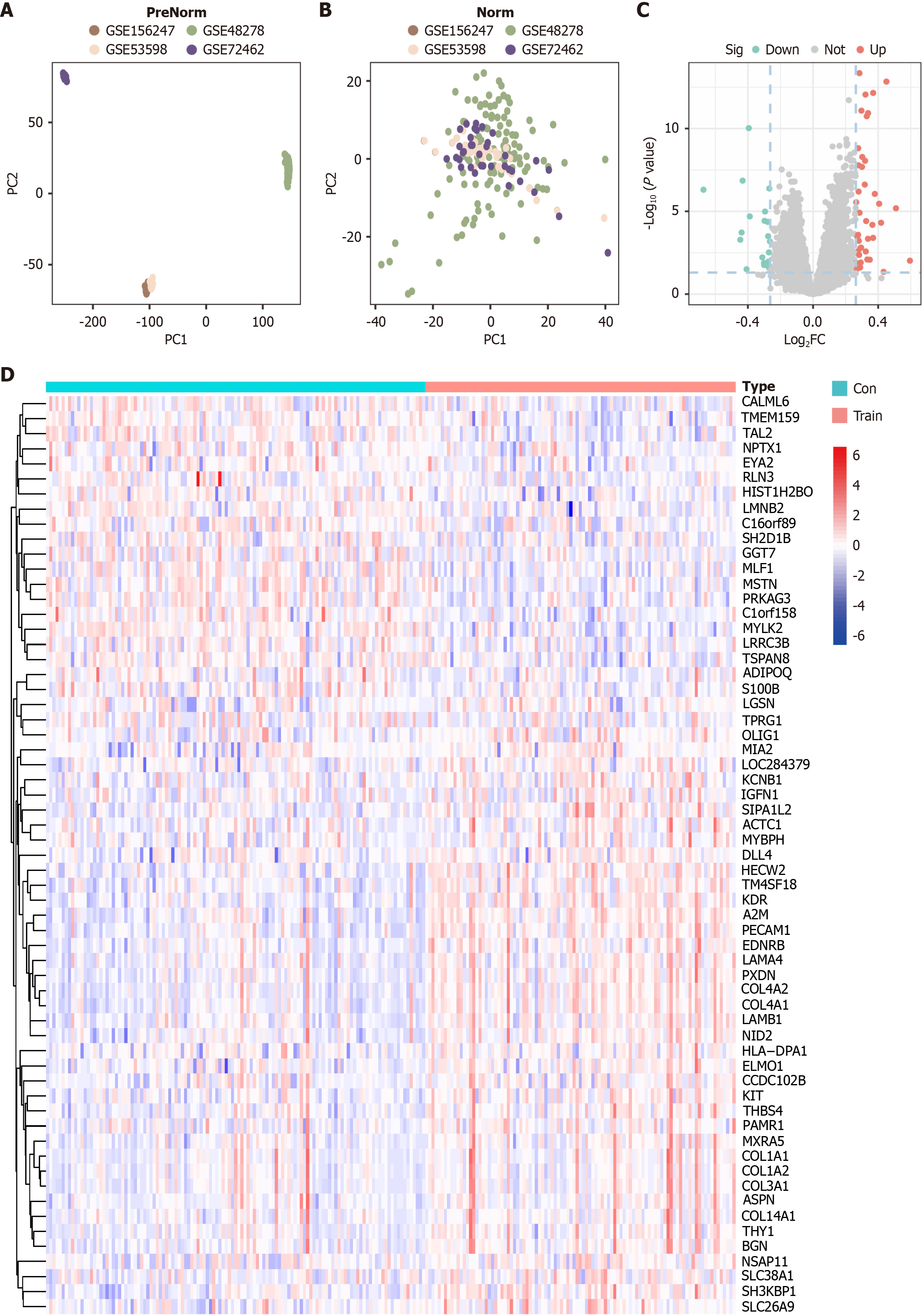
Figure 2 Identification of differentially expressed genes in muscle tissue samples.
A and B: Principal component analysis of different gene expression omnibus datasets. The gene expression profiles were visualized using scatter plots based on the first two principal components with (B) and without (A) batch effect correction; C: Volcano plots of the differentially expressed genes (DEGs); D: Heatmap showing DEGs between pre-exercise and post-exercise samples. PC1: Principal components 1; PC2: Principal components 2; FC: Fold change; Sig: Significance; Down: Down-regulation; Not: Not significant; Up: Up-regulation; Con: Pre-exercise; Train: Post-exercise.
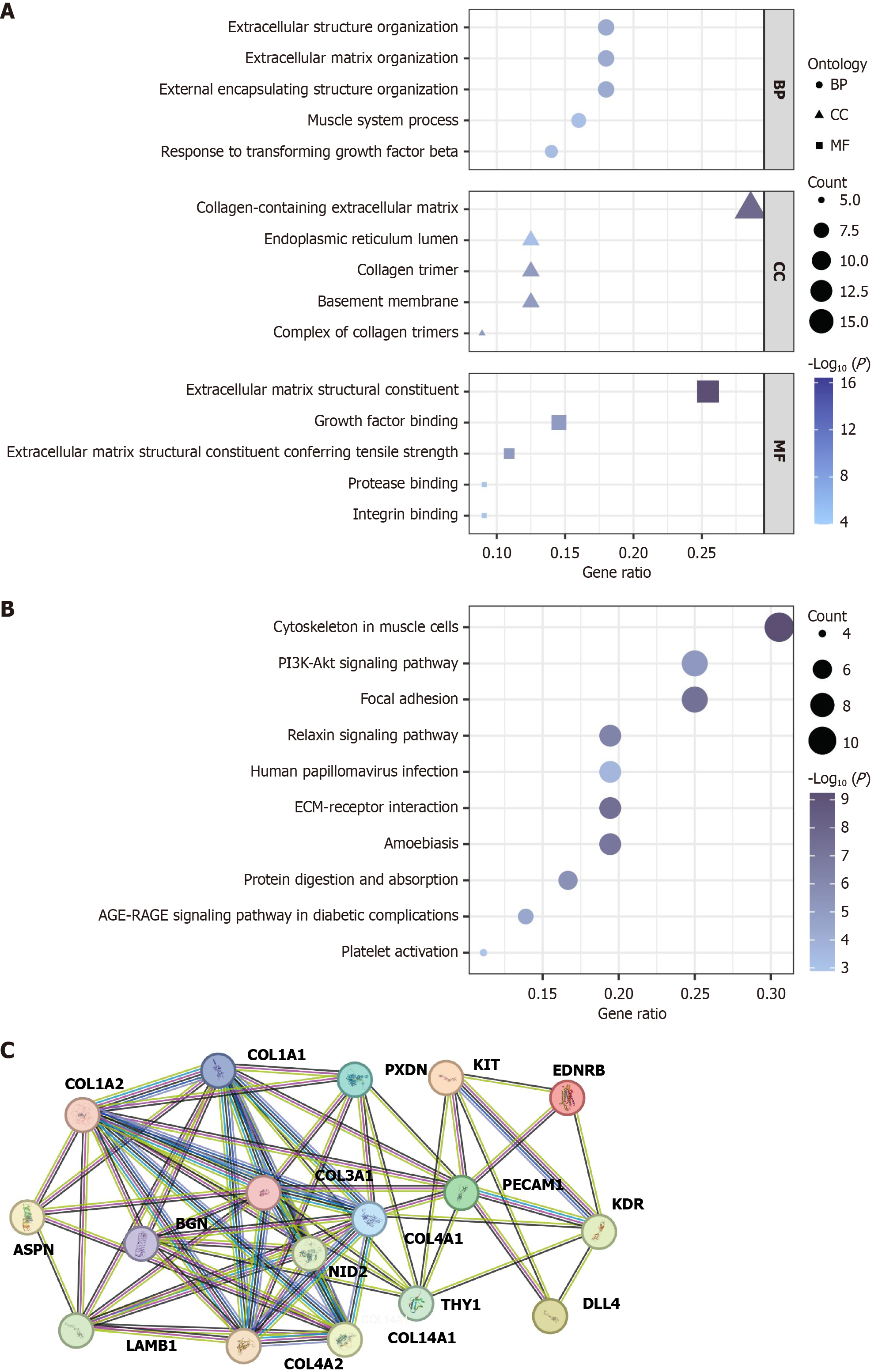
Figure 3 Functional enrichment analysis of differentially expressed genes.
A: Bubble charts showing the enrichment of Gene Ontology terms among differentially expressed genes (DEGs); B: Bubble charts depicting the enrichment of Kyoto Encyclopedia of Genes and Genomes pathways among DEGs; C: Protein-protein interaction network diagram for 17 DEGs. BP: Biological processes; CC: Cellular components; MF: Molecular functions; ECM: Extracellular matrix; PI3K: Phosphatidylinositol 3-kinase; Akt: Protein kinase B; AGE-RAGE: Advanced glycation end products-receptor for advanced glycation end products.
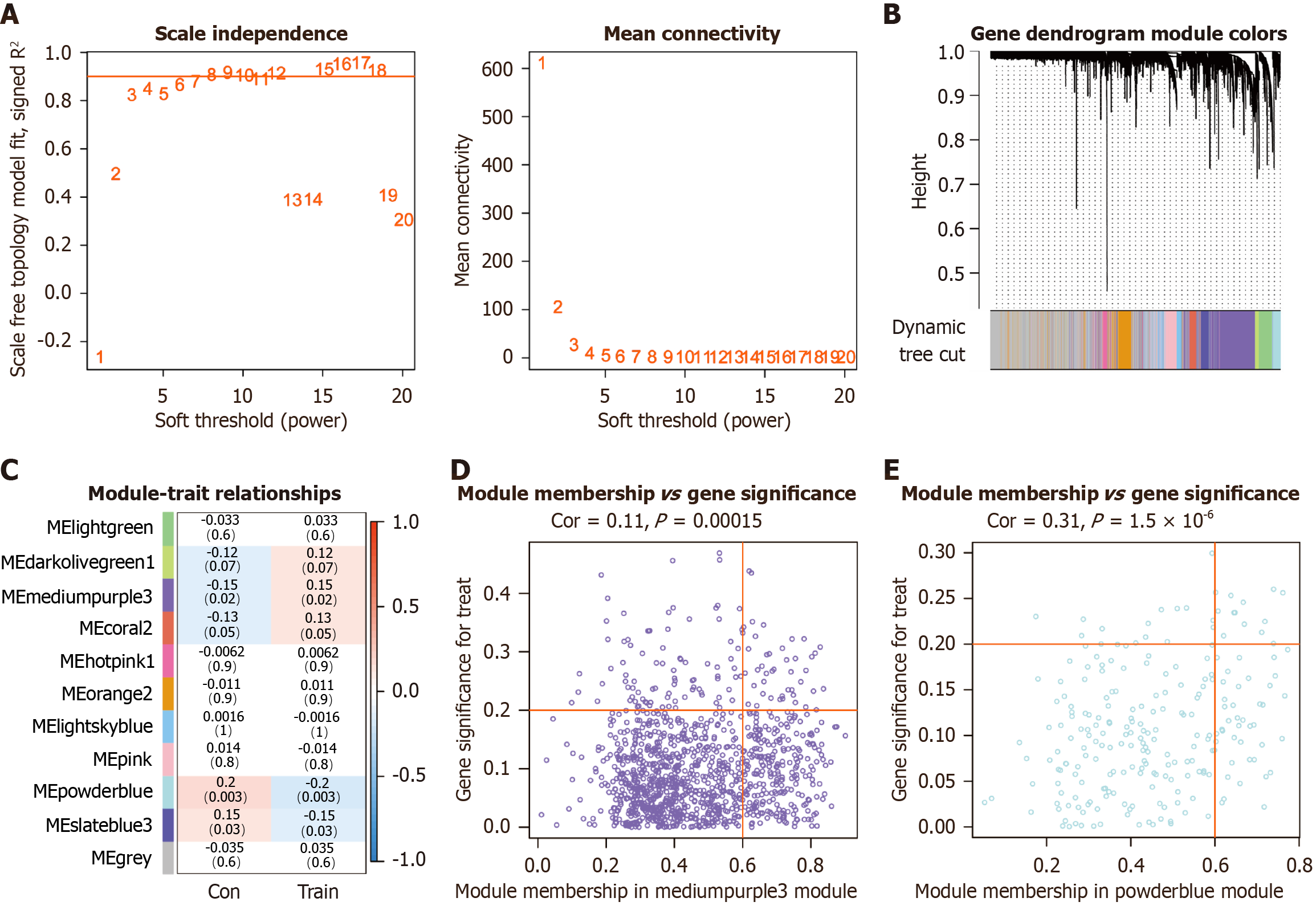
Figure 4 Weighted gene co-expression network analysis of differentially expressed genes.
A: Scale-free topology fitting index (left) and mean connectivity (right) across soft-thresholding powers (β), with the red line indicating the threshold correlation coefficient of 0.9; B: Gene clustering dendrogram constructed using the topological overlap matrix, with color bars representing an co-expression modules; C: Heatmap illustrating the correlation between module eigengenes and clinical traits in metabolic dysfunction-associated steatotic liver disease subjects before and after exercise, with positive correlations in red and negative in blue; D: Scatter plot of gene significance vs module membership in the mediumpurple3 module; E: Scatter plot of gene significance vs module membership in the powder blue module. Con: Pre-exercise; Train: Post-exercise.
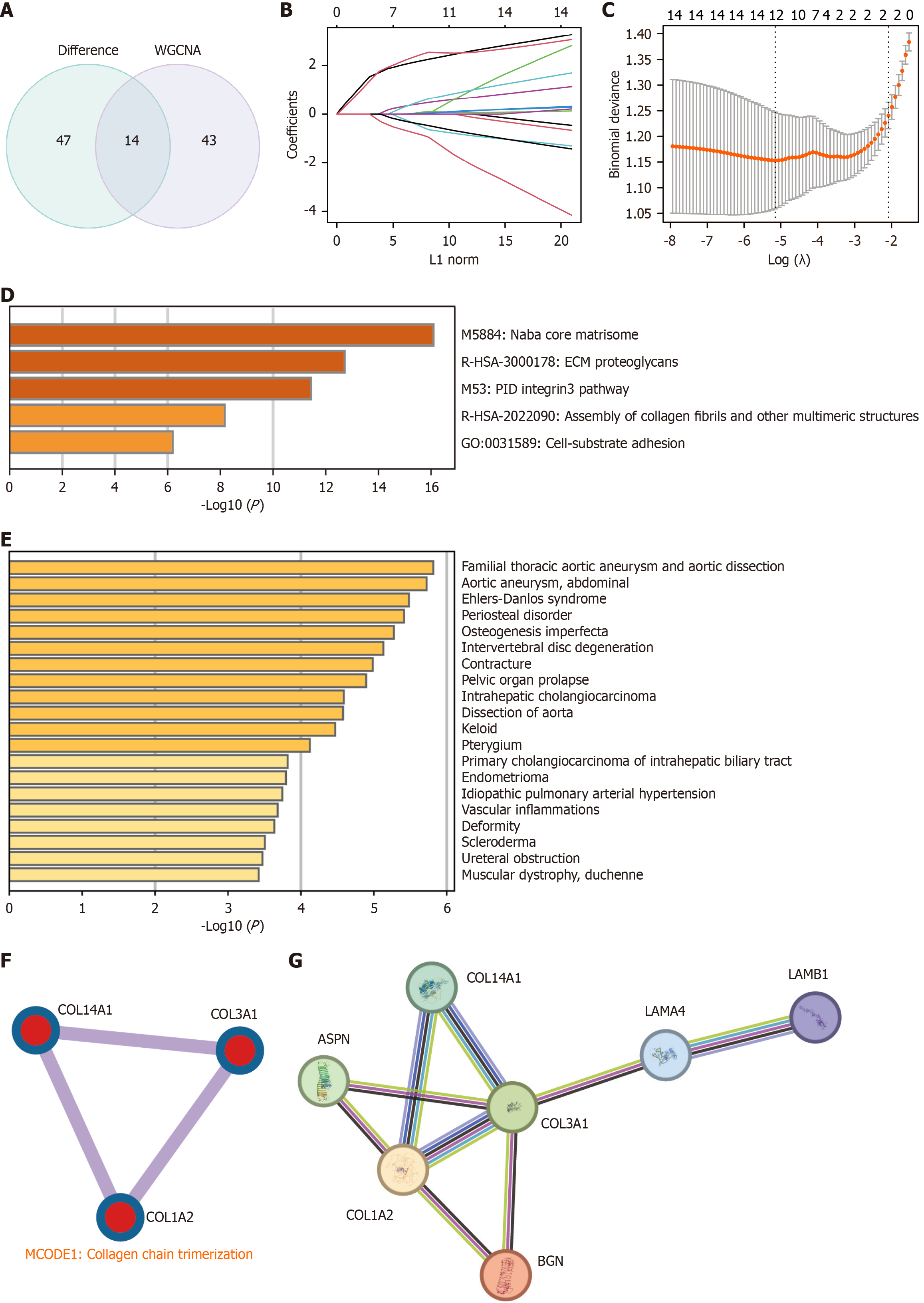
Figure 5 Identification and functional enrichment analysis of key hub genes.
A: Venn diagram showing the overlap of hub genes from differentially expressed genes and weighted gene co-expression network analysis; B and C: Identification of twelve genes through least absolute shrinkage and selection operator regression; curves represent individual genes, with the dashed line marking the optimal lambda; D: Gene Ontology (GO) enrichment analysis of hub genes, with -log10 P values on the X-axis; E: DisGeNET enrichment of hub genes, X-axis showing -log10 P values; F: Top molecular complex detection clusters of hub genes from the protein-protein interaction (PPI) network, with GO analysis highlighting key biological functions; G: PPI network of the 7 key hub genes. WGCNA: Weighted gene co-expression network analysis; GO: Gene ontology; MCODE: Molecular complex detection.
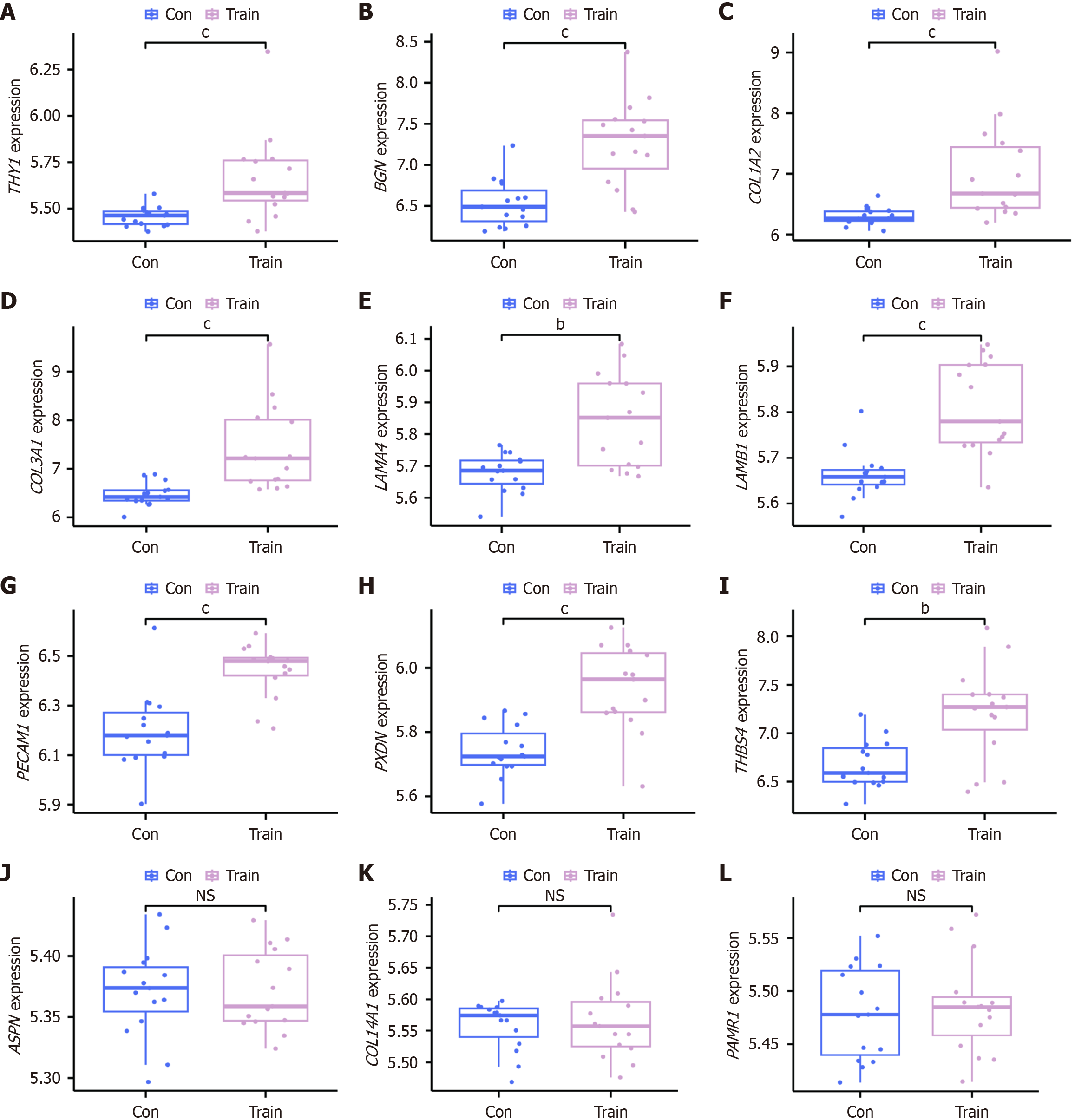
Figure 6 Expression of key hub genes in the validation cohort.
A-L: Box plots illustrating the expression levels of genetic biomarkers in the validation cohort (GSE58249): THY1 (A); BGN (B); COL1A2 (C); COL3A1 (D); LAMA4 (E); LAMB1 (F); PECAM1 (G); PXDN (H); THBS4 (I); ASPN (J); COL14A1 (K); PAMR1 (L). bP < 0.01. cP < 0.001. NS: Not significant; Con: Pre-exercise; Train: Post-exercise.
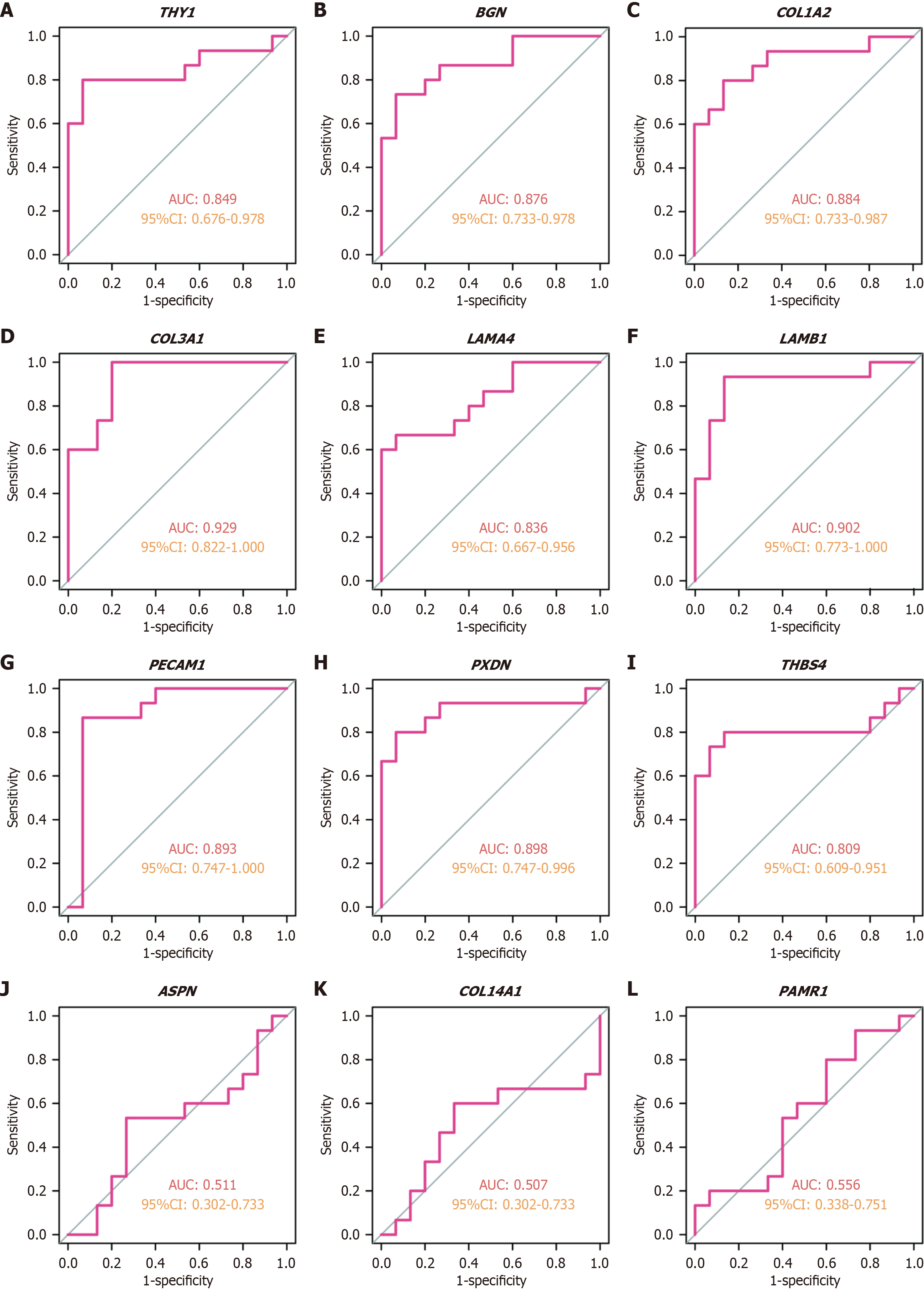
Figure 7 Diagnostic performance of key hub genes in the validation cohort.
A-L: Receiver operating characteristic curve analysis for the validation cohort (GSE58249): THY1 (A); BGN (B); COL1A2 (C); COL3A1 (D); LAMA4 (E); LAMB1 (F); PECAM1 (G); PXDN (H); THBS4 (I); ASPN (J); COL14A1 (K); PAMR1 (L). AUC: Area under the curve; CI: Confidence interval.
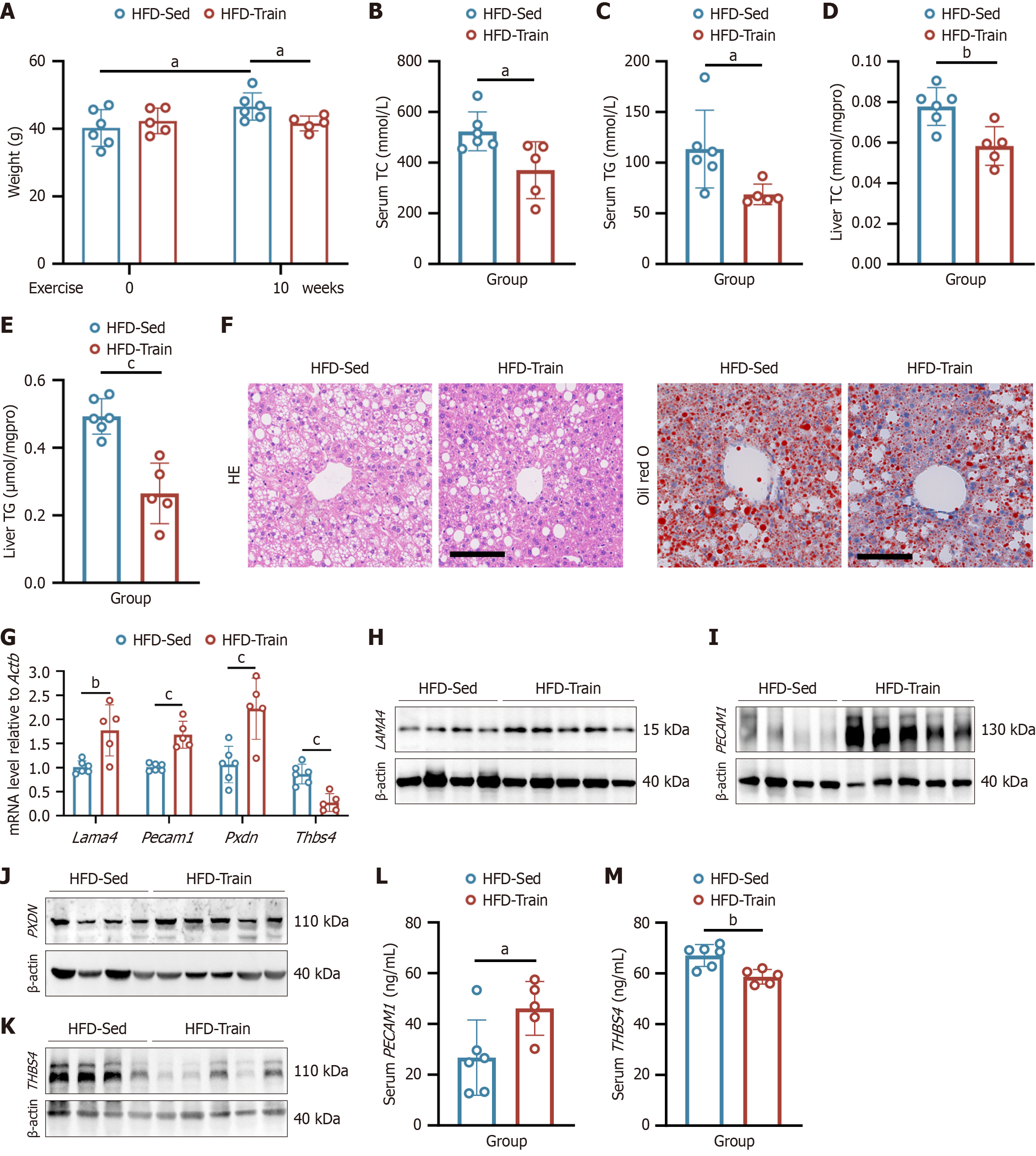
Figure 8 Expression of genetic biomarkers in high-fat diet-induced metabolic dysfunction-associated steatotic liver disease combined with endurance exercise.
A: Body weight changes in mice subjected to 14 weeks of high-fat diet (HFD) followed by 10 weeks of endurance exercise. Measurements were taken before and after the exercise intervention, n ≥ 5; B and C: Serum total cholesterol (TC) (B) and triglyceride (TG) (C) levels in HFD mice with no exercise (HFD-Sed) and HFD with exercise (HFD-Train), n ≥ 5; D and E: Liver TC (D) and TG (E) levels in mice with HFD-Sed and HFD-Train group, n ≥ 5; F: Representative images of hematoxylin and eosin staining and oil red O staining of liver sections. Scale bars, 100 μm; G: The messenger RNA levels of Lama4, Pecam1, Pxdn and Thbs4 in gastrocnemius muscle of HFD-Sed and HFD-Train mice were measured by quantitative polymerase chain reaction, Actb was used as an internal control, n ≥ 5; H-K: Western blot analysis to assess the protein expression levels of laminin subunit α 4 (H), platelet endothelial cell adhesion molecule 1 (PECAM1) (I), peroxidasin (J), thrombospondin 4 (THBS4) (K), n ≥ 4; L and M: The serum level of PECAM1 (L) and THBS4 (M) were measured using an enzyme-linked immunosorbent assay, n ≥ 5. aP < 0.05. bP < 0.01. cP < 0.001. HFD: High-fat diet; HFD-Sed: High-fat diet with no exercise; HFD-Train: High-fat diet with exercise; TC: Total cholesterol; TG: Triglyceride; HE: Hematoxylin and eosin; mRNA: Messenger RNA; LAMA4: Laminin subunit α 4; PECAM1: Platelet endothelial cell adhesion molecule 1; PXDN: Peroxidasin; THBS4: Thrombospondin 4.
- Citation: Zhang JH, Chen K, Zhu XM, Zhou H, Jiang JM, Zou YQ, Liu KR, Zhang L, Li Y. Exercise-responsive skeletal muscle genes mechanistically linked to metabolic dysfunction-associated steatotic liver disease. World J Gastroenterol 2026; 32(13): 113985
- URL: https://www.wjgnet.com/1007-9327/full/v32/i13/113985.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i13.113985