Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.113947
Revised: October 25, 2025
Accepted: December 25, 2025
Published online: March 15, 2026
Processing time: 185 Days and 23.7 Hours
Solute carrier family 30 member 8 (SLC30A8) is a protein encoded for a zinc efflux transporter in insulin secretory granules of pancreatic islets. Genome wide asso
To evaluate whether DNA methylation changes in SLC30A8 is associated with T2D and diabetic kidney disease (DKD).
A total of 820 Chinese subjects, including non-diabetic control subjects, newly diagnosed T2D, T2D with and without DKD and additional 230 healthy subjects before and after physical exercise were enrolled in this study. DNA methylation levels of six CpG sites in the SLC30A8 gene promoter were analyzed with bisulfite pyrose
DNA methylation levels of SLC30A8 were found to be remarkably high (81.01% ± 3.9%) in this Chinese cohort. Compared to non-diabetic controls, SLC30A8 promoter methylation levels in T2D and DKD patients were significantly higher (81.7% ± 3.9% and 81.6% ± 3.8% vs 79.4% ± 2.5%, P < 0.001 and P < 0.001). Mendelian randomization analysis implicated a causal relationship between decreased SLC30A8 expression and higher risk of T2D. We further investigated SLC30A8 promoter methylation changes in 230 healthy subjects before and after physical exercise and found that the specificity of SLC30A8 promoter hypermethylation was not influenced by environmental factors such as physical exercise.
The current study provides evidence that SLC30A8 gene promoter region exhibits hypermethylation. The SLC30A8 promoter hypermethylation is associated with both T2D and DKD in a Chinese population but not interfered by physical intervention. The findings suggest that SLC30A8 may have its potential value as a novel target of epi
Core Tip: Solute carrier family 30 member 8 (SLC30A8) is a β-cell specific Zn transporter in insulin secretory granules. DNA in the SLC30A8 gene promoter region is hypermethylated. SLC30A8 promoter hypermethylation is associated with both type 2 diabetes and diabetic kidney disease. This study aims to evaluate whether DNA methylation changes in SLC30A8 is associated with type 2 diabetes and diabetic kidney disease.
- Citation: Li N, Li YP, Xu Q, Xu J, Yu YH, Su J, Shen C, Yu JY, Gu HF. SLC30A8 promoter hypermethylation is associated with type 2 diabetes and diabetic kidney disease in a Chinese population. World J Diabetes 2026; 17(3): 113947
- URL: https://www.wjgnet.com/1948-9358/full/v17/i3/113947.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i3.113947
Epigenetic factors mainly including DNA methylation changes have been considered to be involved in the pathogenesis of type 2 diabetes (T2D) and diabetic kidney disease (DKD). DNA methylation analysis may reveal the epigenetic mechanisms and provide new strategies for diagnosis, classification, personalized treatment and prevention of the diseases[1-3]. Solute carrier family 30 member 8 (SLC30A8, also known as Znt8) is a protein encoded for a zinc efflux transporter. This protein is highly expressed in pancreas, particularly in the islets of Langerhans, and functions as a β-cell specific Zn transporter in insulin secretory granules[4]. Clinically, SLC30A8 has been applied as a new antigenic target in type 1 diabetes patients[5,6]. Genome wide association study (GWAS) have confirmed that SLC30A8 is a susceptibility gene for T2D[7-9]. In the promoter region of human SLC30A8 gene, there are six CpG sites. Ten years ago, our former research group has reported that DNA methylation in the promoter region of SLC30A8 is hypermethylated. The SLC30A8 promoter hypermethylation is associated with T2D but not with DKD[10]. To investigate the role of SLC30A8 in DKD, our current research group has recently carried out the experiments by using animal models for diabetes, including streptozotocin-induced diabetic, db/db, ZnT8-KO, ZnT8-KO-streptozotocin and ZnT8-KO-db/db mice. We have demonstrated that SLC30A8 is highly expressed in kidneys and plays a protective role against mesenchymal transition tubulointerstitial fibrosis though the restrain of transforming growth factor-β1/Smads signaling activation in DKD[11].
In the present study, we first analyzed DNA methylation levels in the promoter of this gene in a Chinese population of T2D and DKD. Concurrently, we utilized DNA samples from human pancreatic tissue to verify the specificity of SLC30A8 promoter hypermethylation in pancreas. Mendelian randomization (MR) is a convenient analysis that reduces the effects of confounding bias and yields more robust causal interferences[12,13]. To further validate the relationship between SLC30A8 expression and T2D, we performed MR analysis using SLC30A8 expression as the exposure factor, while T2D as the outcome variable. Subsequently, we employed animal models for study of T2D and DKD to analyze the changes of DNA methylation and RNA expression levels of SLC30A8 in kidneys. Finally, by using DNA samples from Chinese healthy subjects before and after physical exercise, we determined whether SLC30A8 promoter hypermethylation levels were influenced by environmental factors, such as exercise intervention. The purpose of this study was to further evaluate the association of SLC30A8 hypermethylation with T2D and DKD.
Two cohorts were enrolled in this study. In the 1st cohort, there were a total of 820 Chinese subjects, including non-diabetic control subjects (n = 132), newly diagnosed T2D (n = 135), T2D without DKD (n = 399) and DKD (n = 154). T2D patients had diabetic duration ≥ 10 years and normal albuminuria, while DKD patients had urine albumin-to-creatinine ratio (≥ 300 mg/g). All diagnoses were done based upon American Diabetes Association criteria. Clinical characteristics of all subjects in the first cohort are represented in Table 1. We have recently carried out a physical exercise intervention program in the youth (PiPy). All participants were interviewed with the predesigned electronic questionnaire regarding their demographic information, food habits and physical activity and also included in the physical examination for admission of PiPy for 3 months[14]. The 2nd cohort of 230 healthy male subjects from PiPy was included in this study. Their DNA samples at the baseline and 3-month follow-up were used for comparative analysis of SLC30A8 promoter methylation levels. Physiological characteristics of healthy subjects of the 2nd cohort at the baseline and after PiPy are summarized in Table 2. In addition, DNA samples extracted from pancreatic tissues of pancreatic serous cystadenoma (SCA) patients with (n = 3) and without T2D (n = 10) were included in the detection of SLC30A8 promoter methylation levels. Clinical information of SCA patients is attached in Supplementary Table 1. This study was conducted in ac
| NDC | Newly T2D | T2D | DKD | P value, a/b/c | |
| N (females) | 132 (43.2%) | 135 (43.0%) | 399 (41.1%) | 154 (35.1%) | |
| Age (years) | 55 ± 17 | 52 ± 14 | 55 ± 16 | 60 ± 14 | 0.548/0.192/0.005 |
| BMI (kg/m2) | 25.2 ± 5.2 | 25.1 ± 4.1 | 25.5 ± 3.7 | 25.6 ± 3.7 | 0.994/0.725/0.995 |
| SBP (mmHg) | 135 ± 18 | 130 ± 16 | 135 ± 18 | 141 ± 20 | 0.029/0.047/0.001 |
| DBP (mmHg) | 85 ± 15 | 83 ± 9 | 82 ± 11 | 81 ± 13 | 0.729/0.812/0.649 |
| FBG (mol/L) | 5.6 ± 1.9 | 9.2 ± 4.1 | 8.0 ± 3.3 | 8.1 ± 3.7 | < 0.001/0.002/0.992 |
| HbA1c (%)/(mmol/mol)1 | 6.0 ± 1.5/42.4 ± 16.1 | 10.1 ± 2.7/86.5 ± 26.3 | 8.8 ± 2.2/72.7 ± 23.9 | 9.2 ± 2.1/77.1 ± 23.0 | < 0.001/< 0.001/0.204 |
| Fc-peptide (ng/mL) | 1.94 ± 1.34 | 1.59 ± 0.97 | 1.83 ± 1.08 | 2.04 ± 1.65 | 0.124/0.245/0.266 |
| UACR (mg/g) | 8.6 ± 11.5 | 11.7 ± 12.9 | 142.5 ± 51.6 | 914.3 ± 346.7 | 0.996/0.214/< 0.001 |
| SCr (μmol/L) | 70.2 ± 25.3 | 63.0 ± 15.2 | 72.5 ± 25.4 | 120.1 ± 106.2 | 0.650/0.235/< 0.001 |
| BUN (mmol/L) | 7.9 ± 3.1 | 5.4 ± 1.6 | 6.8 ± 6.8 | 9.6 ± 14.3 | 0.146/0.492/0.016 |
| LDL (mmol/L) | 2.6 ± 0.6 | 3.1 ± 0.9 | 2.8 ± 0.9 | 2.8 ± 1.0 | < 0.001/0.027/0.995 |
| HDL (mmol/L) | 1.3 ± 0.3 | 3.5 ± 0.8 | 1.3 ± 0.3 | 1.3 ± 0.3 | 0.063/0.014/0.996 |
| TG (mmol/L) | 1.7 ± 1.2 | 2.0 ± 1.7 | 2.2 ± 2.7 | 2.2 ± 2.3 | 0.708/0.936/0.992 |
| TC (mmol/L) | 4.4 ± 0.9 | 4.9 ± 1.3 | 4.7 ± 1.3 | 4.8 ± 1.3 | 0.004/0.298/0.959 |
| AST (U/L) | 21.8 ± 14.8 | 19.5 ± 16.5 | 20.6 ± 18.5 | 19.3 ± 13.7 | 0.649/0.897/0.825 |
| ALT (U/L) | 31.3 ± 32.2 | 26.6 ± 22.9 | 24.9 ± 31.7 | 20.8 ± 14.7 | 0.528/0.935/0.431 |
| Characteristics | Baseline | PiPy | Changes | P value1 |
| BW (kg) | 67.74 ± 12.96 | 68.80 ± 12.36 | 1.11 ± 2.28 | 0.373 |
| BMI (kg/m2) | 22.18 ± 3.64 | 22.59 ± 3.47 | 0.43 ± 0.85 | 0.218 |
| Hipline (m) | 93.91 ± 7.44 | 93.64 ± 10.06 | -0.27 ± 9.81 | 0.747 |
| Sit-and-reach (cm) | 7.03 ± 8.28 | 14.84 ± 7.58 | 7.87 ± 6.84 | < 0.0001 |
| Handgrip (N) | 36.93 ± 7.47 | 43.03 ± 10.84 | 5.95 ± 10.13 | < 0.0001 |
| 1000 m-run (minutes) | 4.08 ± 0.53 | 3.73 ± 0.42 | -0.35 ± 0.34 | < 0.0001 |
| PA (°) | 5.40 ± 0.57 | 5.54 ± 0.46 | 0.14 ± 0.37 | 0.005 |
| SBP (mmHg) | 118.67 ± 10.24 | 116.17 ± 11.27 | -2.50 ± 9.32 | 0.013 |
| DBP (mmHg) | 68.28 ± 7.07 | 65.19 ± 9.69 | -3.10 ± 8.48 | < 0.0001 |
| HR (bmp) | 80.70 ± 12.09 | 70.79 ± 10.21 | -10.09 ± 10.96 | <0.0001 |
| Relafat value (%) | 15.85 ± 8.02 | 17.34 ± 7.52 | 1.49 ± 2.40 | 0.041 |
| Abdfat value (%) | 11.54 ± 7.71 | 12.67 ± 7.61 | 1.12 ± 1.89 | 0.116 |
| HDL-C (mmol/L) | 1.48 ± 0.22 | 1.50 ± 0.40 | 0.02 ± 0.31 | 0.592 |
| LDL-C (mmol/L) | 2.54 ± 0.78 | 1.65 ± 0.66 | -0.89 ± 0.54 | < 0.0001 |
| TG (mmol/L) | 1.00 ± 0.57 | 0.79 ± 0.56 | -0.21 ± 0.51 | < 0.0001 |
| TC (mmol/L) | 4.48 ± 0.83 | 3.51 ± 0.83 | -0.97 ± 0.75 | < 0.0001 |
| FBG (mmol/L) | 3.62 ± 0.41 | 4.51 ± 0.67 | 0.89 ± 0.74 | < 0.0001 |
| INS (uIU/mL) | 6.82 ± 4.80 | 5.28 ± 2.28 | -1.55 ± 4.64 | 0.013 |
| HOMA-IR | 1.11 ± 0.78 | 1.05 ± 0.49 | -0.06 ± 0.77 | 0.598 |
Genomic DNA was extracted from peripheral blood or pancreas tissue samples using a DNeasy blood and tissue extraction kit according to the manufacturer’s protocol (Qiagen, Germany) and quantified using a spectrophotometry (NanoDrop 2000, Thermo Scientific, MA, United States).
The bisulfite pyrosequencing is a sensitive and accurate technique for DNA methylation analysis[15]. All DNA samples were treated with sodium bisulfite using EpiTect Bisulfite kit (Qiagen, Germany) and cleanup of bisulfite-converted DNA was done. Polymerase chain reaction (PCR) amplification was carried out using PyroMark CpG assay (Qiagen, Germany), PyroMark Gold Q96 reagent kits (Qiagen, Germany) and sequenced in PyroMark Q48 Autoprep (Qiagen, Germany). The PCR amplicon (185 bp) covers the sequence in human chromosome 8: 117962434 to 117962479. The unmethylated and unconverted DNA samples (Qiagen, Germany) were used for control of conversion efficiency in bisulfite treatment and accuracy in DNA methylation analysis.
We performed a MR analysis and the analytical procedure is shown in Supplementary Figure 1. To acquire the exposure factor dataset, we retrieved 15 papers by searching “QTL (quantitative trait loci)” and “East Asian” in the PubMed database (Date to June, 2025) and selected the dataset from the top-ranked article (“best match”) as the exposure variable dataset[16]. Concerning the outcome variable dataset, we identified four East Asian GWAS datasets for T2D in the IEU OpenGWAS database. The reported datasets were open-access with the closest publication date and thereby chosen as the outcome variable dataset[17]. To select the instrumental variable (IV), we primarily extracted single nucleotide polymorphisms (SNPs) in SLC30A8 with genome-wide significance (P < 5.0E-8) from the QTL data as IV Set 1. To strictly satisfy MR’s relevance assumption, we queried the ensemble database for all SNPs within chr8: 113,950,273-120,176,714 annotated to SLC30A8 (field: External_gene_name) as IV Set 2. After verified palindromic SNPs to ensure accurate exposure effect estimation, the screened SNPs (rs11558471, rs13266634 and rs3802177) in GWAS catalog were confirmed to be associated with SLC30A8 and T2D-related traits. Additional information concerning the packet tools and references is summarized under this table. Pruned SNPs via linkage disequilibrium clumping (r2 < 0.1 and kb > 1000) were used to mitigate horizontal pleiotropy. Finally, MR analysis was performed using R (v4.4.2) with the TwoSampleMR package (v0.6.16). The harmonized IV set was aligned to ensure the comparisons of consistent allele effects. Information of SNPs (rs2466293, rs2466300 and rs57177943) used for MR analysis and related references are summarized in Supplementary Table 2.
Db/db (BKS.Cg-Dock7m +/+ Lepr db/J) mice are commonly used model for study of T2D and DKD[18]. The male db/db mice used in the present study were purchased from Institute of Model Animal Research at Nanjing University, Nanjing, China. The db/db mice with DKD (at the age of 14 weeks) was identified based on the presence of blood glucose > 16.7 mmol/L and urine albumin-to-creatinine ratio > 200 mg/g on two consecutive tests. The number of db/db mice with and without DKD was 4 each group. All mice resided in a specific pathogen-free barrier environment (three mice in each cage), kept at a temperature of 24 ± 2 °C and humidity of 50% ± 10%, experiencing a 12-hour cycle of light and darkness at the Animal Experimental Center of China Pharmaceutical University[19-21]. The animal experiments were conducted according to the relevant experimental regulations of China Pharmaceutical University.
Extraction of genomic DNA from kidney tissues was done with the animal genomic DNA kit (Omega Bio-Tek, GA, United States). DNA bisulfite transformation was performed using a Zymo Research transformation kit (Shanghai RainBio Technology Co., Ltd., Shanghai, China). The sequences of bisulfite transformed DNA was analysed with Illumina HiSeq 2500 (Illumina, CA, United States). The proportion of methylated cytosines in the SLC30A8 gene was analyzed using Bismark software (v2.90) and R-package Methylkit to estimate the methylation levels[22,23]. Total RNAs were extracted from crushed kidney tissues with Trizol protocol (Invitrogen, PO, United States). The RNA samples that met the criteria were reverse transcribed to complementary DNAs and subsequently sequenced with PE150-Illumina Hiseq platform (Illumina, CA, United States). Expression matrices were analyzed for differential expression using DESeq2 software (v3.17)[24]. SLC30A8 gene expression at mRNA levels was confirmed with real time real time-PCR as we have previously described[11].
Tests for comparison of continuous variables between groups were conducted using either an unpaired t-test or one-way analysis of variance, followed by Tukey’s post hoc test for pairwise comparisons. For non-normally distributed traits, data were transformed using the natural logarithm to achieve normality prior to statistical analysis. A P value of less than 0.05 was considered statistically significant. All statistical analyses were performed using IBM SPSS Statistics 25 (SPSS Inc., Chicago, IL, United States).
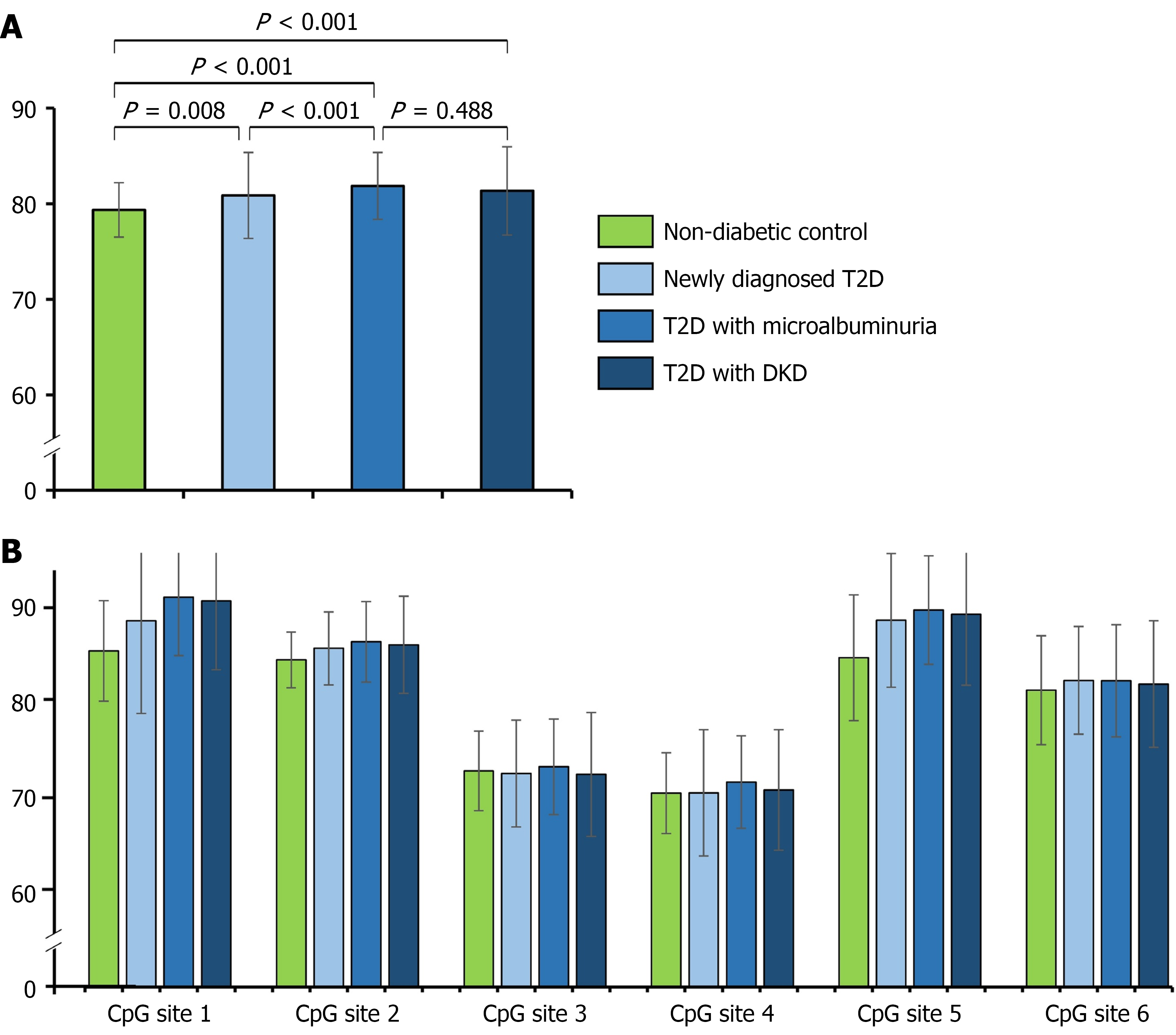
We implemented the age- and body mass index-matched groups, introduced a newly diagnosed T2D to minimize the confounding effects and analyzed the DNA methylation levels in the promoter region of SLC30A8 in a Chinese population. There are 6 CpG sites in the promoter region of the SLC30A8 gene in human as previously described[25]. The average DNA methylation levels of SLC30A8 promoter in this population were remarkably high (81.2% ± 3.9%). Figure 1A illustrated that compared to non-diabetic control, SLC30A8 promoter methylation levels were significantly higher in T2D either newly diagnosed or with diabetic duration more than 10 years, as well as in DKD compared to non-diabetic controls (81.2% ± 4.4%, 81.7% ± 3.9% and 81.6% ± 3.8% vs 79.4% ± 2.5%, P = 0.001, P < 0.001 and P < 0.001, respectively). Figure 1B showed DNA methylation levels at six CpG sites of SLC30A8 promoter region in the groups of non-diabetic control, newly diagnosed T2D, T2D and DKD.
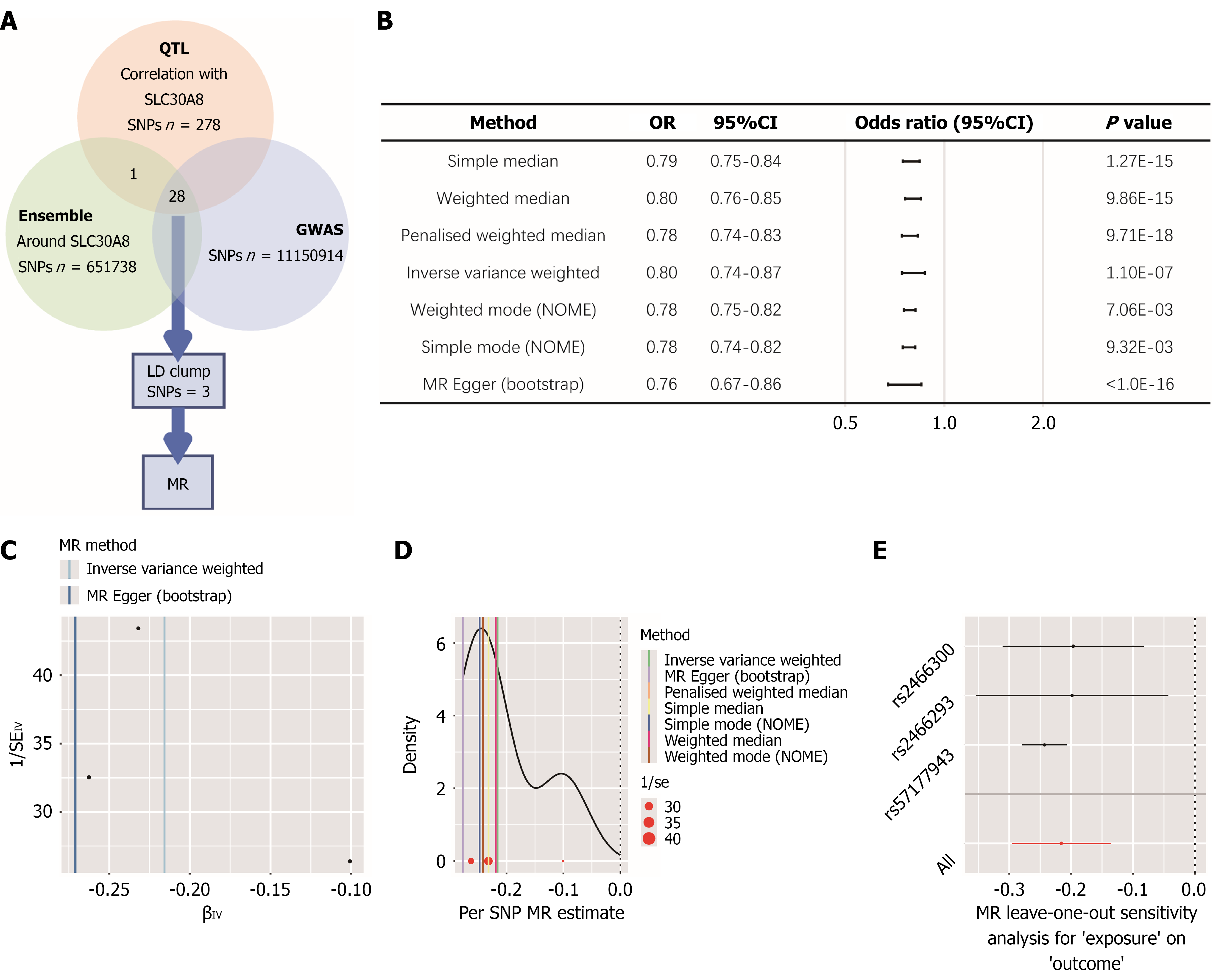
We utilized QTL data from the East Asian populations (n = 2099) and GWAS for T2D as the outcome (n = 433540) to conduct MR analysis, with manual verification confirming no cohort duplication (Figure 2A). Based on the selected IVs, we implemented multiple MR methods using SLC30A8 expression as the exposure and T2D as the outcome (Figure 2B). Specifically, three methods suited for the limited SNP counts, i.e. simple median [odds ratio (OR) = 0.79, 95% confidence interval (CI): 0.75-0.84, P = 1.27E-15], weighted median (OR = 0.80, 95%CI: 0.76-0.85, P = 9.86E-15), and penalized weighted median (OR = 0.78, 95%CI: 0.74-0.83, P = 9.71E-18), which consistently indicated that SLC30A8 expression negatively regulated T2D with high confidence. The data were supported by the analyses with other methods, including inverse variance weighted (OR = 0.80, 95%CI: 0.74-0.87, P = 1.10E-07), weighted mode (NOME) (OR = 0.78, 95%CI: 0.75-0.82, P = 7.06E-03), simple mode (NOME) (OR = 0.78, 95%CI: 0.74-0.82, P = 9.32E-03), and MR Egger (bootstrap) (OR = 0.76, 95%CI: 0.67-0.86, P < 1.0E-16). Additionally, we assessed each IV’s contribution to the inverse variance weighted method (Figure 2C-E) to evaluate heterogeneity. The IV assumptions for rs2466293, rs2466300 and rs57177943 were validated. MR analysis thereby established a significant causal relationship between decreased SLC30A8 expression and higher risk of T2D. We initially screened SNPs that were significantly associated with both the exposure and outcome (P < 1E-6) and located within a 1E+6 base-pair physical distance from genes, yielding a total of 28 SNPs as candidate IVs. To minimize horizontal pleiotropy, we performed linkage disequilibrium-clumping, ultimately retaining 3 SNPs for the final MR analysis.
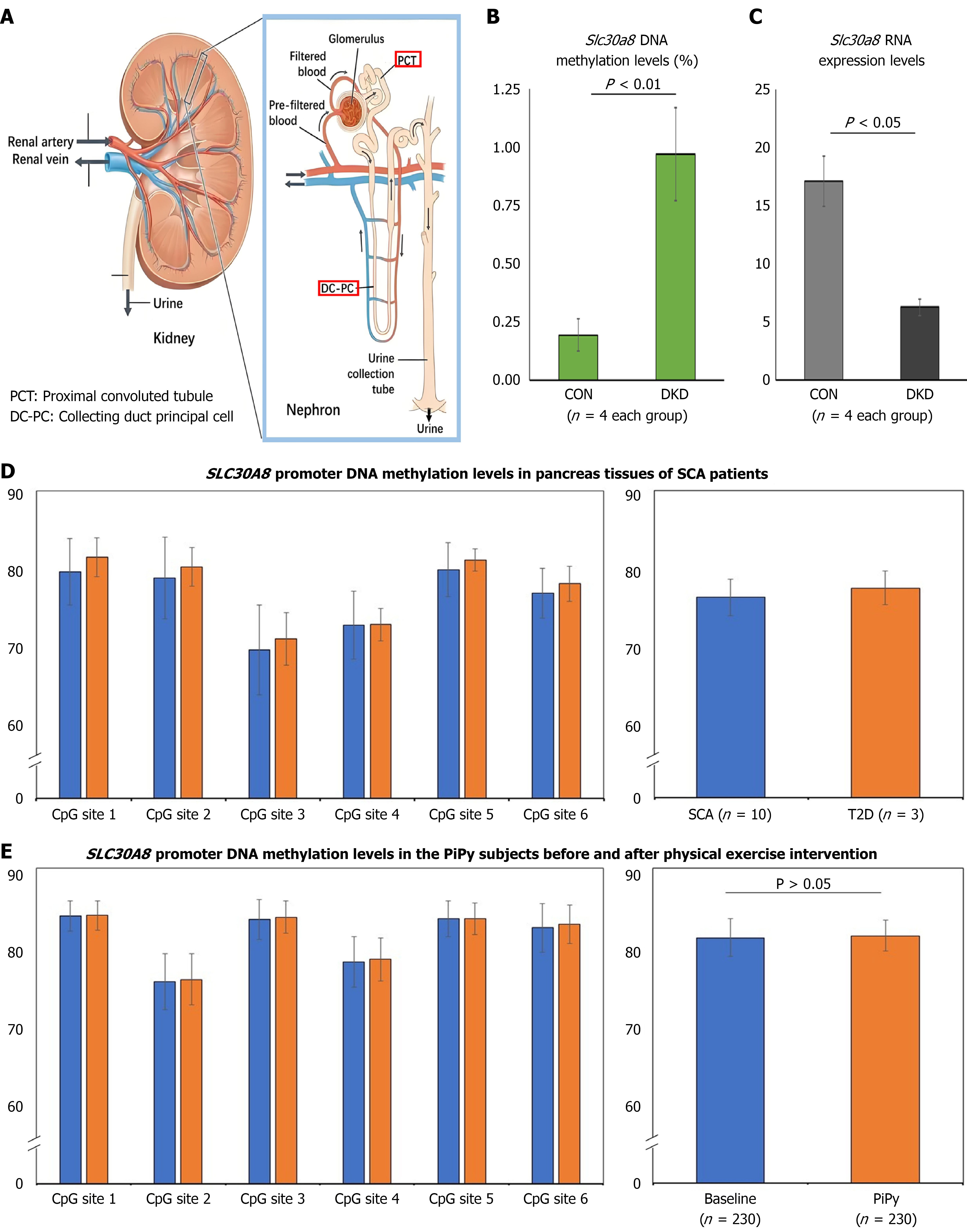
Our research group has recently employed the technology of single-cell transcriptomic analysis to analyze the kidneys in db/db mice and found that SLC30A8 is primarily expressed in the proximal convoluted tubule and collecting duct principal cells (Figure 3A)[19]. In the present study, we further verified the variation of DNA methylation and mRNA expression of SLC30A8 in kidneys of db/db mice with and without DKD. Since the promoter region of this gene in mice lacks CpG sites, we analyzed the overall DNA methylation and mRNA expression levels of this gene in kidney tissues. Data demonstrated that higher methylation levels of SLC30A8 in kidneys of db/db mice with DKD compared to those without DKD, while mRNA expression of this gene showed an inverse trend (Figure 3B and C).
SLC30A8 is primarily highly expressed in pancreatic tissue[4]. Due to the difficulty in obtaining pancreatic tissues from patients with T2D and DKD, we extracted DNA from pancreatic tissues of SCA patients (with and without T2D) and detected similarly high promoter methylation levels of this gene as observed in peripheral blood cells (Figure 3D). However, the limited sample size precluded formal statistical analysis, which is a limitation of this study. To ascertain whether environmental factors such as physical activity could influence DNA methylation of this gene, we investigated the changes of SLC30A8 promoter methylation whether in PiPy subjects from baseline to 3-month follow-up. The average methylation levels of six CpG sites in the SLC30A8 gene promoter were 77.9% ± 5.9% before exercise intervention and 78.5% ± 4.9% after intervention, but no significant difference between the two groups (P = 0.270) was seen (Figure 3E).
Over the past decade, despite laboratory relocations, our research group has been dedicated to investigate the changes of SLC30A8 promoter methylation levels and their associations with T2D and DKD. Ten years ago, we unexpectedly observed that SLC30A8 promoter methylation levels were very high in Malaysia population and found that SLC30A8 promoter hypermethylation is associated with T2D in Malaysian population but failed to detect the association with DKD[10]. In the present study, we have investigated the changes of SLC30A8 promoter DNA methylation levels in non-diabetic controls, the subjects of newly diagnosed T2D, T2D with diabetic duration and DKD. Data from this study have consistently demonstrated that the promoter region of SLC30A8 is hypermethylated in Chinese population. Furthermore, we found that SLC30A8 promoter hypermethylation is associated with not only T2D but also DKD. Generally, the DNA methylation levels of genes associated with diabetes and diabetic nephropathy are around 20% (2, 25-27). However, the methylation levels of this gene promoter are as high as 80%, which is rare in epigenetic methylation studies.
MR analysis is commonly applied to estimate the causal relationship between a modifiable environmental exposure and a medical relevant trait or disease such as T2D and its complications by utilizing genetic variants as IVs[12,13,16,17]. In the present study, our MR analysis has employed rigorously selected IVs to minimize confounding factors, demonstrating robust causal inference even when some MR assumptions are violated, while maintaining strong statistical power for detecting the weak effects. Notably, MR-Egger yielded discordant results (OR = 1.32, 95%CI: 0.99-1.76, P = 0.313), potentially influenced by limited SNP counts and intercept values. Therefore, we have considered MR-Egger (bootstrap) more reliable for assessing pleiotropy’s impact on inverse variance weighted estimates, with both methods converging on minimal pleiotropic effects. The data of MR analysis may support the causal relationship between decreased SLC30A8 expression and higher risk of T2D.
SLC30A8 is primarily highly expressed in pancreatic tissue[4]. Due to the difficulty in obtaining pancreatic tissues from patients with T2D and DKD, we extracted DNA from pancreatic tissues of SCA patients (with and without T2D) and detected similarly high promoter methylation levels of this gene as observed in peripheral blood cells. However, there are a couple of limitations in the present study. First, the sample sizes of DNA extracted from pancreatic tissues of SCA patients with and without T2D are limited, which may preclude the formal statistical analysis. Second, MR fundamentally reanalyzes the existing datasets constrained by summary statistics. Although we have mitigated horizontal pleiotropy through methodological controls, its complete elimination still remains challenging. While prioritizing East Asian populations for consistency with observational studies, potential heterogeneity across subpopulations may affect robustness due to the dataset limitations. In addition, the database searches (National Center for Biotechnology Information, ensemble) have revealed only one annotated CpG site (cg20041597 at Pos.4 of CpG island in SLC30A8 promoter), with no mQTL-derived IVs available even at lenient thresholds (P < 1.0E-06).
Accumulating evidence has indicated that increased methylation levels of genes can suppress their function, playing a significant role in the pathology of T2D and DKD[26-30]. The above findings have indicated that the promoter hypermethylation of SLC30A8 is associated with T2D and DKD, while this association is likely attributed to reduced gene function, warranting further investigation through biological experiments. In the recent years, our research group has been utilizing animal models of T2D and DKD such as db/db mice to investigate the molecular mechanisms underlying the pathophysiology of these diseases. We have identified the expression of SLC30A8 in proximal convoluted tubule and collecting duct principal cell of kidneys and also discovered its anti-inflammatory and anti-fibrotic roles during the pathological progression of DKD[11]. In this study, we have verified the variation of DNA methylation and mRNA expression of SLC30A8 in kidneys of db/db mice with and without DKD. Since the promoter region of this gene in mice lacks methylation sites, we analyzed the overall DNA methylation and mRNA expression levels of this gene and found that higher methylation levels of SLC30A8 in kidneys of db/db mice with DKD compared to those without DKD, while mRNA expression of this gene showed an inverse trend. The data obtained from animal experiments further demonstrate that the elevated DNA methylation and decreased mRNA expression levels of SLC30A8 are involved in the pathogenesis and progression of T2D and DKD.
During our data analysis, we observed that patients with DKD exhibited a downward trend in SLC30A8 promoter hypermethylation levels compared to T2D, albeit remaining higher than in non-diabetic individuals. This phenomenon may result from pharmacological effects, which could explain the lack of significant association with DKD observed in our earlier Malaysian cohort study[10]. Given the diversity in medication regimens (including both Western and traditional Chinese medicines) and substantial variability in patient compliance, we were unable to identify specific drug contributions despite conducting multiple analyses. The methylation levels of a gene may be influenced by environmental factors such as age, body weight, physical activity, and pharmacological interventions[31,32]. To ascertain whether environmental factors such as physical activity could influence DNA methylation of this gene, we further investigated the changes of SLC30A8 promoter methylation whether in PiPy subjects from baseline to 3-month follow-up and found that the methylation status of remains unaffected. In this study, the utilization of diverse cohorts - encompassing human, animal, and exercise models - enhances the generalizability and robustness of the findings. Thereby, the study may have significant potential implications for early diagnosis and epigenetic therapy. Notably, SLC30A8 emerges as a promising target for pharmacological intervention; for instance, demethylating agents could be explored given that the gene’s methylation status remains largely unaffected by exercise, underscoring its stability as a therapeutic target.
A high methylation level of up to 80% in gene promoters typically carries significant biological implications, primarily involving gene silencing. Such hypermethylation (e.g., 80%) in promoter regions can impede transcription factor binding, leading to substantial suppression of gene expression. For instance, aberrant hypermethylation of tumor suppressor genes (e.g., BRCA1) may inactivate their function, thereby promoting tumorigenesis[33,34]. Additionally, it is associated with disease progression and epigenetic regulatory dysregulation. In this study, we observed that SLC30A8 promoter hypermethylation correlates with T2D and DKD. However, it is likely unrelated to environmental factors such as physical exercise. Further experimental validation is required to determine whether SLC30A8 promoter hypermethylation indeed inhibits its gene function. Based on our project experience, a major challenge will be the collection of pancreatic tissue samples from T2D and DKD patients for this purpose.
The current study provides evidence that DNA methylation levels in the promoter region of SLC30A8 are extremely high in a Chinese population. The SLC30A8 promoter hypermethylation is associated with T2D and DKD, but not interfered by environmental factors such as physical exercise. The findings suggest that SLC30A8 may be a new epigenetic pharmacological target in medication of diabetes and its complications.
The authors wish to thank Pancreas Biobank of the First Affiliated Hospital with Nanjing Medical University for providing with the pancreatic tissue samples in this study.
| 1. | Ling C, Bacos K, Rönn T. Epigenetics of type 2 diabetes mellitus and weight change - a tool for precision medicine? Nat Rev Endocrinol. 2022;18:433-448. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 95] [Article Influence: 23.8] [Reference Citation Analysis (0)] |
| 2. | Kato M, Natarajan R. Epigenetics and epigenomics in diabetic kidney disease and metabolic memory. Nat Rev Nephrol. 2019;15:327-345. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 513] [Cited by in RCA: 447] [Article Influence: 63.9] [Reference Citation Analysis (1)] |
| 3. | Gu HF. Epigenetics of Diabetic Nephropathy. In: Handbook of Nutrition, Diet, and Epigenetics. Switzerland: Springer, Cham, 2019: 1-20. [DOI] [Full Text] |
| 4. | Chimienti F, Devergnas S, Favier A, Seve M. Identification and cloning of a beta-cell-specific zinc transporter, ZnT-8, localized into insulin secretory granules. Diabetes. 2004;53:2330-2337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 409] [Cited by in RCA: 367] [Article Influence: 16.7] [Reference Citation Analysis (0)] |
| 5. | Lernmark Å, Agardh D, Akolkar B, Gesualdo P, Hagopian WA, Haller MJ, Hyöty H, Johnson SB, Elding Larsson H, Liu E, Lynch KF, McKinney EF, McIndoe R, Melin J, Norris JM, Rewers M, Rich SS, Toppari J, Triplett E, Vehik K, Virtanen SM, Ziegler AG, Schatz DA, Krischer J. Looking back at the TEDDY study: lessons and future directions. Nat Rev Endocrinol. 2025;21:154-165. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 22] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 6. | Regnell SE, Lernmark Å. Early prediction of autoimmune (type 1) diabetes. Diabetologia. 2017;60:1370-1381. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 128] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 7. | Scott LJ, Mohlke KL, Bonnycastle LL, Willer CJ, Li Y, Duren WL, Erdos MR, Stringham HM, Chines PS, Jackson AU, Prokunina-Olsson L, Ding CJ, Swift AJ, Narisu N, Hu T, Pruim R, Xiao R, Li XY, Conneely KN, Riebow NL, Sprau AG, Tong M, White PP, Hetrick KN, Barnhart MW, Bark CW, Goldstein JL, Watkins L, Xiang F, Saramies J, Buchanan TA, Watanabe RM, Valle TT, Kinnunen L, Abecasis GR, Pugh EW, Doheny KF, Bergman RN, Tuomilehto J, Collins FS, Boehnke M. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science. 2007;316:1341-1345. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2216] [Cited by in RCA: 2042] [Article Influence: 107.5] [Reference Citation Analysis (0)] |
| 8. | Kirchhoff K, Machicao F, Haupt A, Schäfer SA, Tschritter O, Staiger H, Stefan N, Häring HU, Fritsche A. Polymorphisms in the TCF7L2, CDKAL1 and SLC30A8 genes are associated with impaired proinsulin conversion. Diabetologia. 2008;51:597-601. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 204] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 9. | Strawbridge RJ, Dupuis J, Prokopenko I, Barker A, Ahlqvist E, Rybin D, Petrie JR, Travers ME, Bouatia-Naji N, Dimas AS, Nica A, Wheeler E, Chen H, Voight BF, Taneera J, Kanoni S, Peden JF, Turrini F, Gustafsson S, Zabena C, Almgren P, Barker DJ, Barnes D, Dennison EM, Eriksson JG, Eriksson P, Eury E, Folkersen L, Fox CS, Frayling TM, Goel A, Gu HF, Horikoshi M, Isomaa B, Jackson AU, Jameson KA, Kajantie E, Kerr-Conte J, Kuulasmaa T, Kuusisto J, Loos RJ, Luan J, Makrilakis K, Manning AK, Martínez-Larrad MT, Narisu N, Nastase Mannila M, Ohrvik J, Osmond C, Pascoe L, Payne F, Sayer AA, Sennblad B, Silveira A, Stancáková A, Stirrups K, Swift AJ, Syvänen AC, Tuomi T, van 't Hooft FM, Walker M, Weedon MN, Xie W, Zethelius B; DIAGRAM Consortium; GIANT Consortium; MuTHER Consortium; CARDIoGRAM Consortium; C4D Consortium, Ongen H, Mälarstig A, Hopewell JC, Saleheen D, Chambers J, Parish S, Danesh J, Kooner J, Ostenson CG, Lind L, Cooper CC, Serrano-Ríos M, Ferrannini E, Forsen TJ, Clarke R, Franzosi MG, Seedorf U, Watkins H, Froguel P, Johnson P, Deloukas P, Collins FS, Laakso M, Dermitzakis ET, Boehnke M, McCarthy MI, Wareham NJ, Groop L, Pattou F, Gloyn AL, Dedoussis GV, Lyssenko V, Meigs JB, Barroso I, Watanabe RM, Ingelsson E, Langenberg C, Hamsten A, Florez JC. Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes. Diabetes. 2011;60:2624-2634. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 327] [Cited by in RCA: 277] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 10. | Seman NA, Mohamud WN, Östenson CG, Brismar K, Gu HF. Increased DNA methylation of the SLC30A8 gene promoter is associated with type 2 diabetes in a Malay population. Clin Epigenetics. 2015;7:30. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 34] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 11. | Zhang X, Guan T, Yang B, Gu HF, Chi Z. Effects of ZnT8 on epithelial-to-mesenchymal transition and tubulointerstitial fibrosis in diabetic kidney disease. Cell Death Dis. 2020;11:544. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 12. | Emdin CA, Khera AV, Kathiresan S. Mendelian Randomization. JAMA. 2017;318:1925-1926. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3188] [Cited by in RCA: 2654] [Article Influence: 294.9] [Reference Citation Analysis (0)] |
| 13. | Skrivankova VW, Richmond RC, Woolf BAR, Yarmolinsky J, Davies NM, Swanson SA, VanderWeele TJ, Higgins JPT, Timpson NJ, Dimou N, Langenberg C, Golub RM, Loder EW, Gallo V, Tybjaerg-Hansen A, Davey Smith G, Egger M, Richards JB. Strengthening the Reporting of Observational Studies in Epidemiology Using Mendelian Randomization: The STROBE-MR Statement. JAMA. 2021;326:1614-1621. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3468] [Cited by in RCA: 2814] [Article Influence: 562.8] [Reference Citation Analysis (1)] |
| 14. | Su J, Wang H, Tian Y, Hu H, Gu W, Zhang T, Li M, Shen C, Gu HF. Impact of physical exercise intervention and PPARγ genetic polymorphisms on cardio-metabolic parameters among a Chinese youth population. BMJ Open Sport Exerc Med. 2020;6:e000681. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 15. | Higashimoto K, Hara S, Soejima H. DNA Methylation Analysis Using Bisulfite Pyrosequencing. Methods Mol Biol. 2023;2577:3-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 16. | Hatton AA, Cheng FF, Lin T, Shen RJ, Chen J, Zheng Z, Qu J, Lyu F, Harris SE, Cox SR, Jin ZB, Martin NG, Fan D, Montgomery GW, Yang J, Wray NR, Marioni RE, Visscher PM, McRae AF. Genetic control of DNA methylation is largely shared across European and East Asian populations. Nat Commun. 2024;15:2713. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 19] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 17. | Spracklen CN, Horikoshi M, Kim YJ, Lin K, Bragg F, Moon S, Suzuki K, Tam CHT, Tabara Y, Kwak SH, Takeuchi F, Long J, Lim VJY, Chai JF, Chen CH, Nakatochi M, Yao J, Choi HS, Iyengar AK, Perrin HJ, Brotman SM, van de Bunt M, Gloyn AL, Below JE, Boehnke M, Bowden DW, Chambers JC, Mahajan A, McCarthy MI, Ng MCY, Petty LE, Zhang W, Morris AP, Adair LS, Akiyama M, Bian Z, Chan JCN, Chang LC, Chee ML, Chen YI, Chen YT, Chen Z, Chuang LM, Du S, Gordon-Larsen P, Gross M, Guo X, Guo Y, Han S, Howard AG, Huang W, Hung YJ, Hwang MY, Hwu CM, Ichihara S, Isono M, Jang HM, Jiang G, Jonas JB, Kamatani Y, Katsuya T, Kawaguchi T, Khor CC, Kohara K, Lee MS, Lee NR, Li L, Liu J, Luk AO, Lv J, Okada Y, Pereira MA, Sabanayagam C, Shi J, Shin DM, So WY, Takahashi A, Tomlinson B, Tsai FJ, van Dam RM, Xiang YB, Yamamoto K, Yamauchi T, Yoon K, Yu C, Yuan JM, Zhang L, Zheng W, Igase M, Cho YS, Rotter JI, Wang YX, Sheu WHH, Yokota M, Wu JY, Cheng CY, Wong TY, Shu XO, Kato N, Park KS, Tai ES, Matsuda F, Koh WP, Ma RCW, Maeda S, Millwood IY, Lee J, Kadowaki T, Walters RG, Kim BJ, Mohlke KL, Sim X. Identification of type 2 diabetes loci in 433,540 East Asian individuals. Nature. 2020;582:240-245. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 404] [Cited by in RCA: 347] [Article Influence: 57.8] [Reference Citation Analysis (0)] |
| 18. | Sharma K, McCue P, Dunn SR. Diabetic kidney disease in the db/db mouse. Am J Physiol Renal Physiol. 2003;284:F1138-F1144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 401] [Cited by in RCA: 385] [Article Influence: 16.7] [Reference Citation Analysis (0)] |
| 19. | Wu C, Tao Y, Li N, Fei J, Wang Y, Wu J, Gu HF. Prediction of cellular targets in diabetic kidney diseases with single-cell transcriptomic analysis of db/db mouse kidneys. J Cell Commun Signal. 2023;17:169-188. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 20. | Wu C, Tang H, Cui X, Li N, Fei J, Ge H, Wu L, Wu J, Gu HF. A single-cell profile reveals the transcriptional regulation responded for Abelmoschus manihot (L.) treatment in diabetic kidney disease. Phytomedicine. 2024;130:155642. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 12] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 21. | Yu H, Tang H, Wang M, Xu Q, Yu J, Ge H, Qiang L, Tang W, Gu HF. Effects of total flavones of Abelmoschus manihot (L.) on the treatment of diabetic nephropathy via the activation of solute carriers in renal tubular epithelial cells. Biomed Pharmacother. 2023;169:115899. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 16] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 22. | Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571-1572. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4957] [Cited by in RCA: 3842] [Article Influence: 256.1] [Reference Citation Analysis (0)] |
| 23. | Akalin A, Kormaksson M, Li S, Garrett-Bakelman FE, Figueroa ME, Melnick A, Mason CE. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 2012;13:R87. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1973] [Cited by in RCA: 1551] [Article Influence: 110.8] [Reference Citation Analysis (0)] |
| 24. | Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010;26:136-138. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3874] [Cited by in RCA: 3165] [Article Influence: 197.8] [Reference Citation Analysis (0)] |
| 25. | Gu HF. Genetic, Epigenetic and Biological Effects of Zinc Transporter (SLC30A8) in Type 1 and Type 2 Diabetes. Curr Diabetes Rev. 2017;13:132-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 17] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 26. | Gu HF. Genetic and Epigenetic Studies in Diabetic Kidney Disease. Front Genet. 2019;10:507. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 60] [Article Influence: 8.6] [Reference Citation Analysis (1)] |
| 27. | Sandholm N, Dahlström EH, Groop PH. Genetic and epigenetic background of diabetic kidney disease. Front Endocrinol (Lausanne). 2023;14:1163001. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 30] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 28. | Smyth LJ, Dahlström EH, Syreeni A, Kerr K, Kilner J, Doyle R, Brennan E, Nair V, Fermin D, Nelson RG, Looker HC, Wooster C, Andrews D, Anderson K, McKay GJ, Cole JB, Salem RM, Conlon PJ, Kretzler M, Hirschhorn JN, Sadlier D, Godson C, Florez JC; GENIE consortium, Forsblom C, Maxwell AP, Groop PH, Sandholm N, McKnight AJ. Epigenome-wide meta-analysis identifies DNA methylation biomarkers associated with diabetic kidney disease. Nat Commun. 2022;13:7891. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 39] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 29. | Schlosser P, Tin A, Matias-Garcia PR, Thio CHL, Joehanes R, Liu H, Weihs A, Yu Z, Hoppmann A, Grundner-Culemann F, Min JL, Adeyemo AA, Agyemang C, Ärnlöv J, Aziz NA, Baccarelli A, Bochud M, Brenner H, Breteler MMB, Carmeli C, Chaker L, Chambers JC, Cole SA, Coresh J, Corre T, Correa A, Cox SR, de Klein N, Delgado GE, Domingo-Relloso A, Eckardt KU, Ekici AB, Endlich K, Evans KL, Floyd JS, Fornage M, Franke L, Fraszczyk E, Gao X, Gào X, Ghanbari M, Ghasemi S, Gieger C, Greenland P, Grove ML, Harris SE, Hemani G, Henneman P, Herder C, Horvath S, Hou L, Hurme MA, Hwang SJ, Jarvelin MR, Kardia SLR, Kasela S, Kleber ME, Koenig W, Kooner JS, Kramer H, Kronenberg F, Kühnel B, Lehtimäki T, Lind L, Liu D, Liu Y, Lloyd-Jones DM, Lohman K, Lorkowski S, Lu AT, Marioni RE, März W, McCartney DL, Meeks KAC, Milani L, Mishra PP, Nauck M, Navas-Acien A, Nowak C, Peters A, Prokisch H, Psaty BM, Raitakari OT, Ratliff SM, Reiner AP, Rosas SE, Schöttker B, Schwartz J, Sedaghat S, Smith JA, Sotoodehnia N, Stocker HR, Stringhini S, Sundström J, Swenson BR, Tellez-Plaza M, van Meurs JBJ, van Vliet-Ostaptchouk JV, Venema A, Verweij N, Walker RM, Wielscher M, Winkelmann J, Wolffenbuttel BHR, Zhao W, Zheng Y; Estonian Biobank Research Team; Genetics of DNA Methylation Consortium, Loh M, Snieder H, Levy D, Waldenberger M, Susztak K, Köttgen A, Teumer A. Meta-analyses identify DNA methylation associated with kidney function and damage. Nat Commun. 2021;12:7174. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 56] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 30. | Nadiger N, Veed JK, Chinya Nataraj P, Mukhopadhyay A. DNA methylation and type 2 diabetes: a systematic review. Clin Epigenetics. 2024;16:67. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 34] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 31. | Światowy WJ, Drzewiecka H, Kliber M, Sąsiadek M, Karpiński P, Pławski A, Jagodziński PP. Physical Activity and DNA Methylation in Humans. Int J Mol Sci. 2021;22:12989. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 62] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 32. | Aronica L, Levine AJ, Brennan K, Mi J, Gardner C, Haile RW, Hitchins MP. A systematic review of studies of DNA methylation in the context of a weight loss intervention. Epigenomics. 2017;9:769-787. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 33. | Baylin SB. DNA methylation and gene silencing in cancer. Nat Clin Pract Oncol. 2005;2 Suppl 1:S4-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 907] [Cited by in RCA: 808] [Article Influence: 38.5] [Reference Citation Analysis (0)] |
| 34. | Fiegl H, Schnaiter S, Reimer DU, Leitner K, Nardelli P, Tsibulak I, Wieser V, Wimmer K, Schamschula E, Marth C, Zeimet AG. BRCA loss of function including BRCA1 DNA-methylation, but not BRCA-unrelated homologous recombination deficiency, is associated with platinum hypersensitivity in high-grade ovarian cancer. Clin Epigenetics. 2024;16:171. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (0)] |