Copyright: ©Author(s) 2026.
World J Diabetes. May 15, 2026; 17(5): 118275
Published online May 15, 2026. doi: 10.4239/wjd.v17.i5.118275
Published online May 15, 2026. doi: 10.4239/wjd.v17.i5.118275
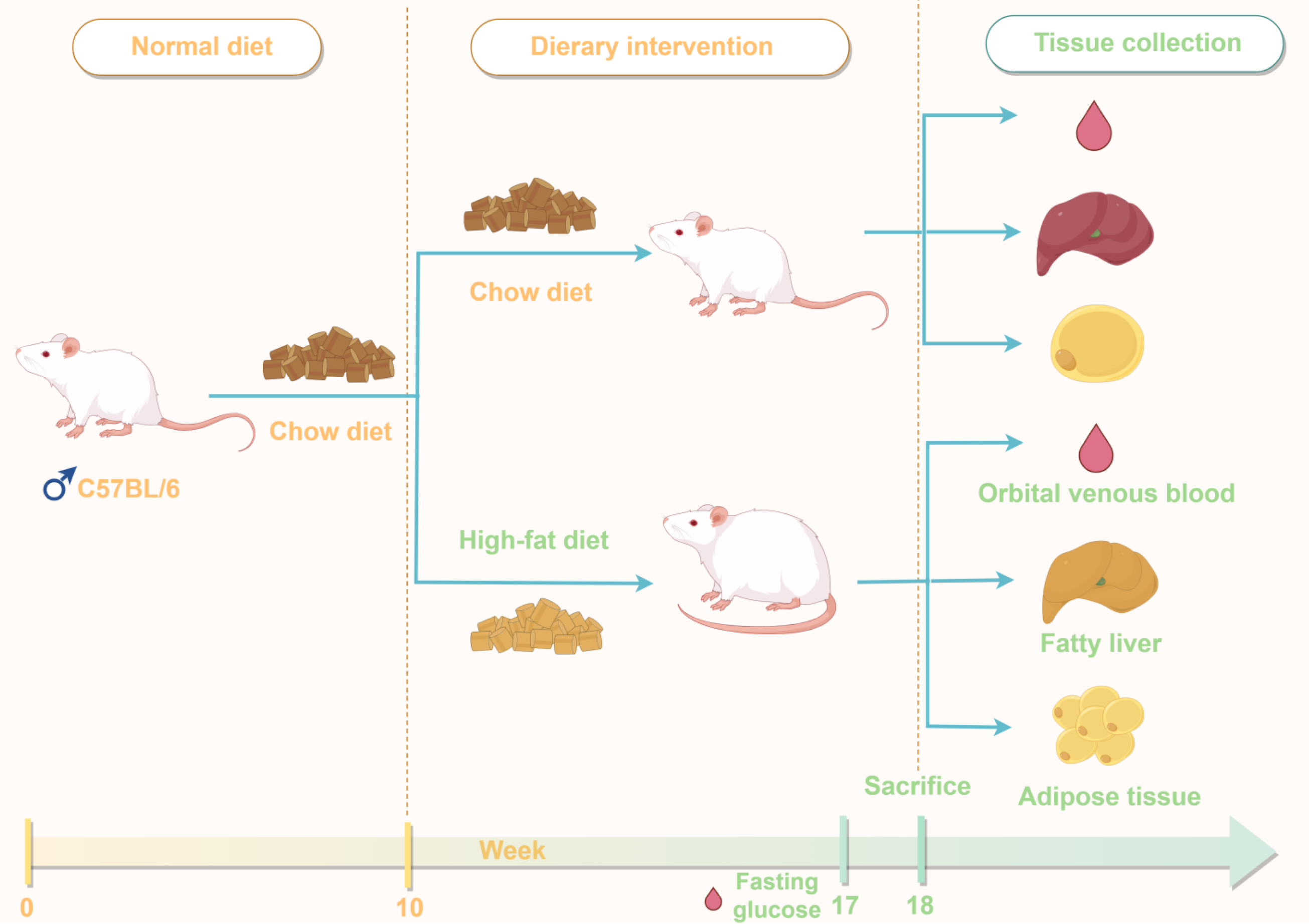
Figure 1 Schematic of the dietary intervention and sample collection timeline.
This diagram outlines the key timeline of the animal experiment. C57BL/6 mice were assigned to either a chow diet or high-fat diet regimen. Obesity was induced by high-fat diet feeding from week 0 to 10. Glucose assessment was performed at week 17. At week 18, mice were sacrificed after fasting, and orbital venous blood, fatty liver tissue, and adipose tissue were collected for subsequent analyses.
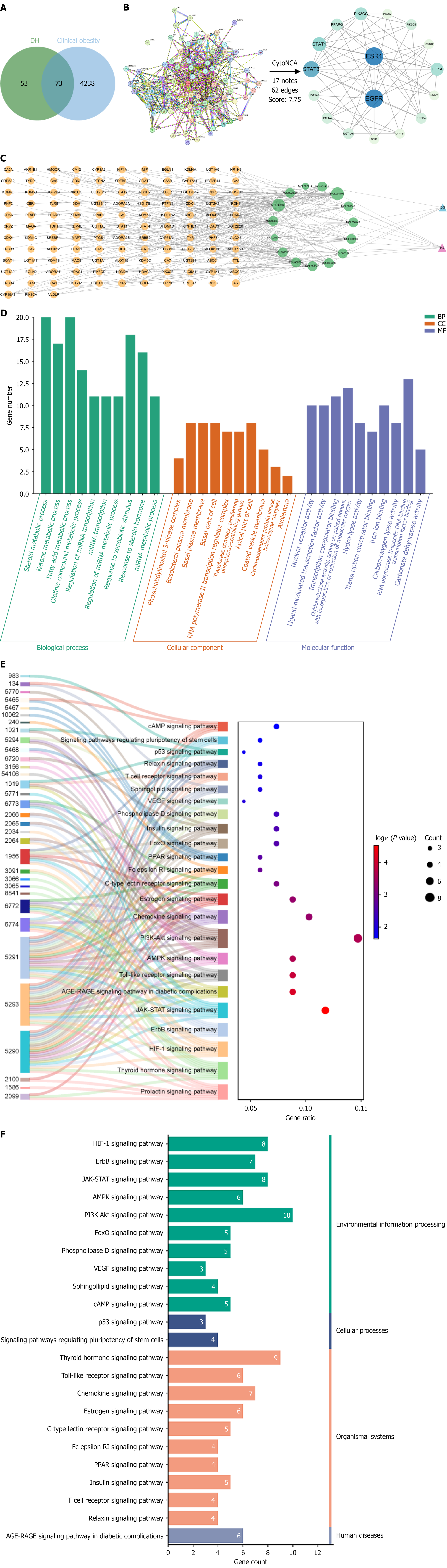
Figure 2 Results of the network pharmacology analysis of a combination of Danggui-Huangjing on clinical obesity.
A: Venn diagram of overlapping targets between Danggui-Huangjing (DH) and clinical obesity (CO). DH targets: Screened from Traditional Chinese Medicine System Pharmacology Database and Analysis Platform (oral bioavailability ≥ 30%, drug likeness ≥ 0.18), Herb.ac.cn, CNKI, verified by PubChem, standardized via UniProt (Homo sapiens); CO targets: Collected from GeneCards (relevance score > 2), Online Mendelian Inheritance in Man, CTDbase, Therapeutic Target Database; B: Protein-protein interaction network and core subnetwork. Left: Initial network (73 nodes, 448 edges) via STRING (confidence > 0.400, hidden isolated nodes); right: Core subnetwork (17 nodes, 62 edges) mined by Cytoscape 310.3/CytoNCA (hub genes: Degree/betweenness/closeness/eigenvector centrality > mean values), epidermal growth factor receptor/estrogen receptor 1 highlighted. Nodes: Larger size/darker color = higher degree; grey edges = target-target interactions; C: “DH-active ingredient-disease target” network. Orange = 73 CO targets; green = 15 DH bioactive components (larger size = higher degree); blue = Danggui (Angelica sinensis); purple = Huangjing (Polygonatum sibiricum); grey edges = compound-target interactions; D: Gene Ontology enrichment (DAVID, P < 0.05). Green = biological process, orange = cellular component, blue = molecular function; E: Kyoto Encyclopedia of Genes and Genomes enrichment (DAVID, P < 0.05). Bubble plot (top 20 pathways): X-axis = enrichment ratio, Y-axis = pathway name, bubble size = target number, purple-yellow gradient = significance (-log10P value); F: Kyoto Encyclopedia of Genes and Genomes pathway hierarchical tree. BP: Biological process; CC: Cellular component; DH: Danggui-Huangjing; MF: Molecular function.
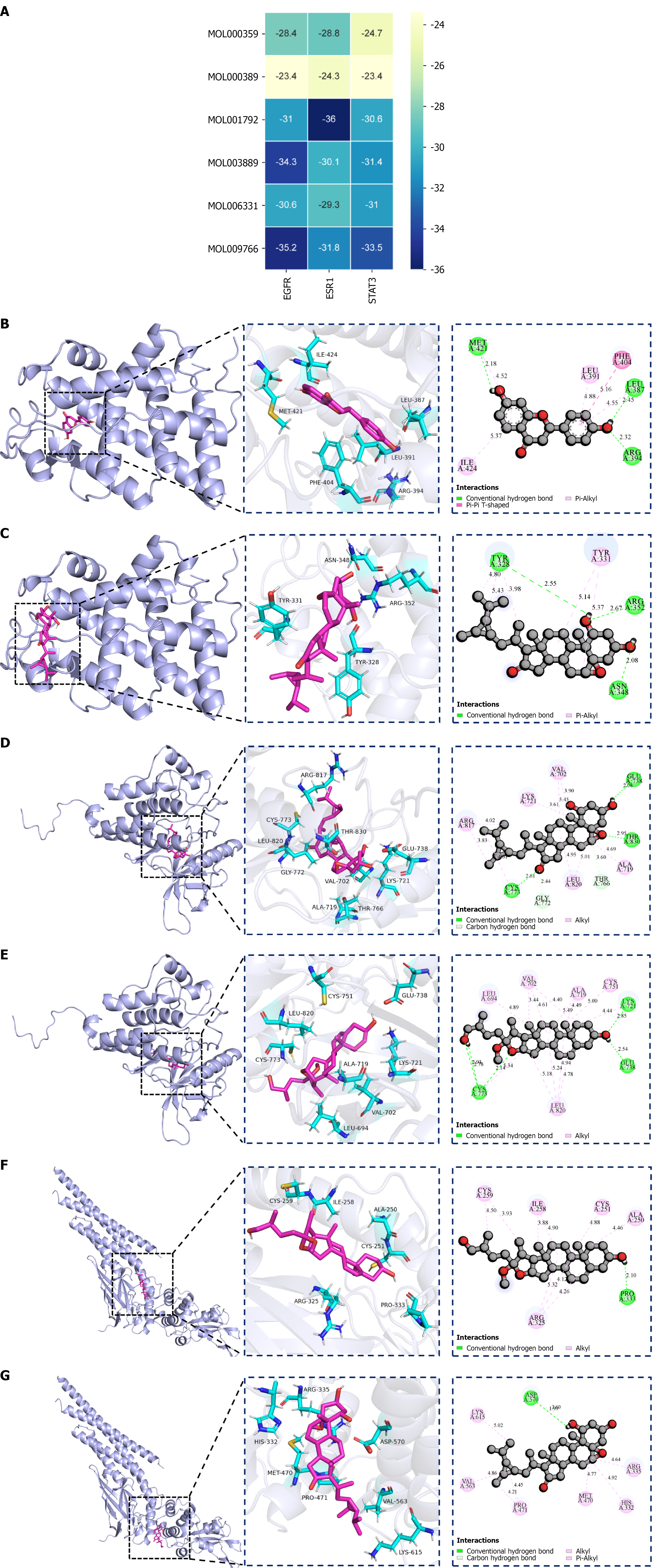
Figure 3 Molecular docking binding energies and the key active ingredient–core target pairs.
A: Heatmap of binding energies (kJ/mol) between 6 Danggui-Huangjing components (rows, MOL IDs) and 3 targets [columns: Epidermal growth factor receptor (EGFR)/estrogen receptor 1 (ESR1)/signal transducer and activator of transcription 3 (STAT3)]. Energies converted from kcal/mol (1 kcal/mol = 4.184 kJ/mol); more negative values = stronger binding; color gradient = affinity; B-G: Docking conformations (PyMOL 2.6.0) and two-dimensional chemical structure of the ligand [B: EGFR-zhonghualiaoine 1 (MOL009766); C: EGFR-methylprotodioscin_qt (MOL003889); D: STAT3-methylprotodioscin_qt (MOL003889); E: STAT3-zhonghualiaoine (MOL009766); F: ESR1-DFV (MOL001792); G: ESR1-zhonghualiaoine 1 (MOL009766)]. Docking via AutoDock 4.2.6 (semi-rigid); receptors (Protein Data Bank structures) processed by PyMOL 2.5.7 (ligands/water removed); binding pockets identified by Getbox Plugin.
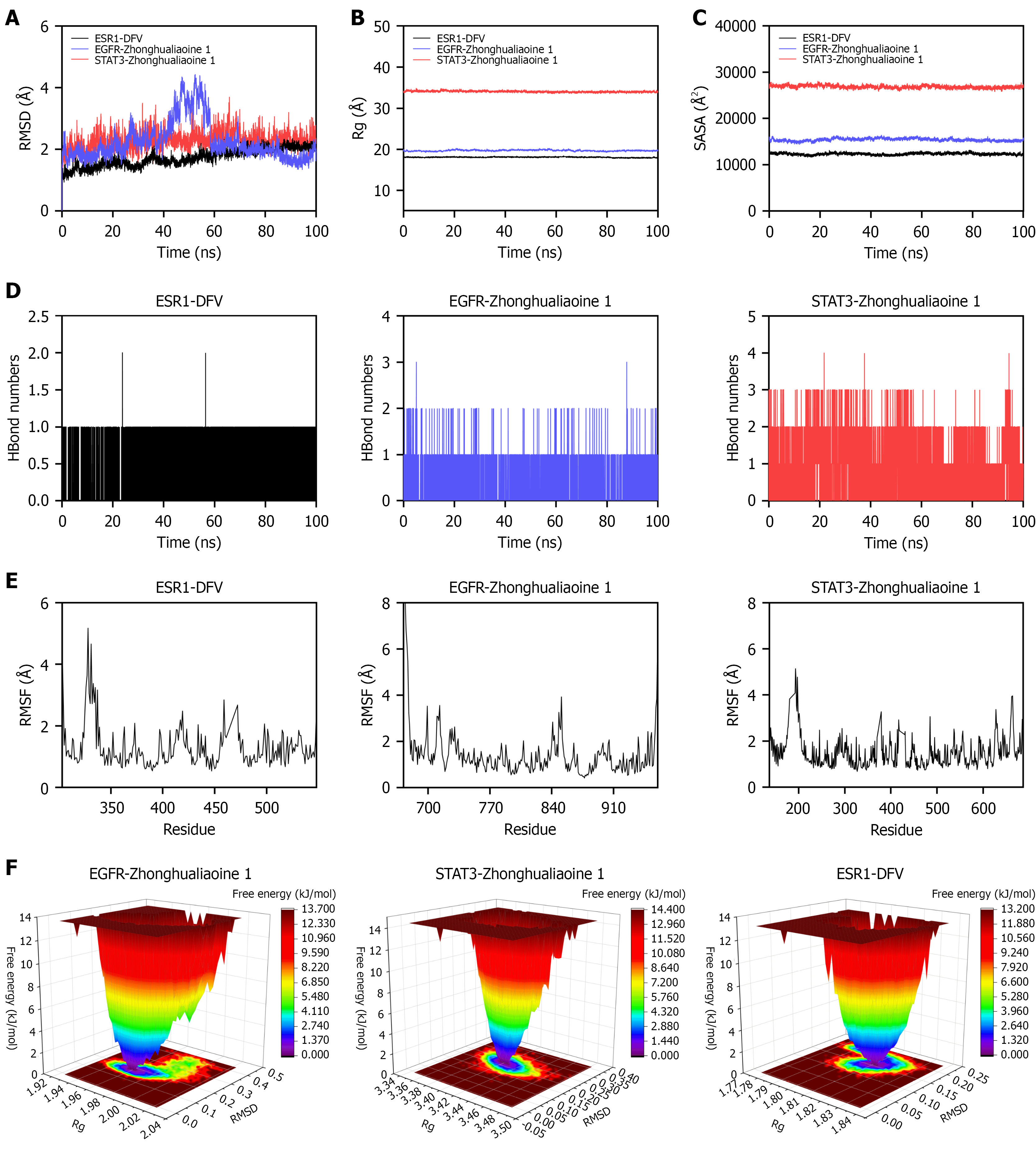
Figure 4 Molecular dynamics simulation of the protein-ligand complexes.
A: Root-mean-square deviation; B: Radius of gyration; C: Solvent-accessible surface area; D: Hydrogen bond count; E: Root-mean-square fluctuation; F: Free energy landscape. Receptors: AMBER14SB force field; ligands: GAFF2 (RESP charges); solvated in TIP3P water box (1 nm edge), 0.15 M NaCl; long-range electrostatics: Particle mesh Ewald method. Molecular dynamics results (GROMACS 2022, 100 nanoseconds, normal pressure and temperature: 310 K, 1 bar) of estrogen receptor 1/diosgenin-3-O-β-D-fructofuranosyl-α-L-rhamnopyranoside, epidermal growth factor receptor/zhonghualiaoine 1, signal transducer and activator of transcription 3/zhonghualiaoine 1 complexes. DFV: Diosgenin-3-O-β-D-fructofuranosyl-α-L-rhamnopyranoside; EGFR: Epidermal growth factor receptor; ESR1: Estrogen receptor 1; Rg: Radius of gyration; RMSD: Root-mean-square deviation; RMSF: Root-mean-square fluctuation; SASA: Solvent-accessible surface area; STAT3: Signal transducer and activator of transcription 3.
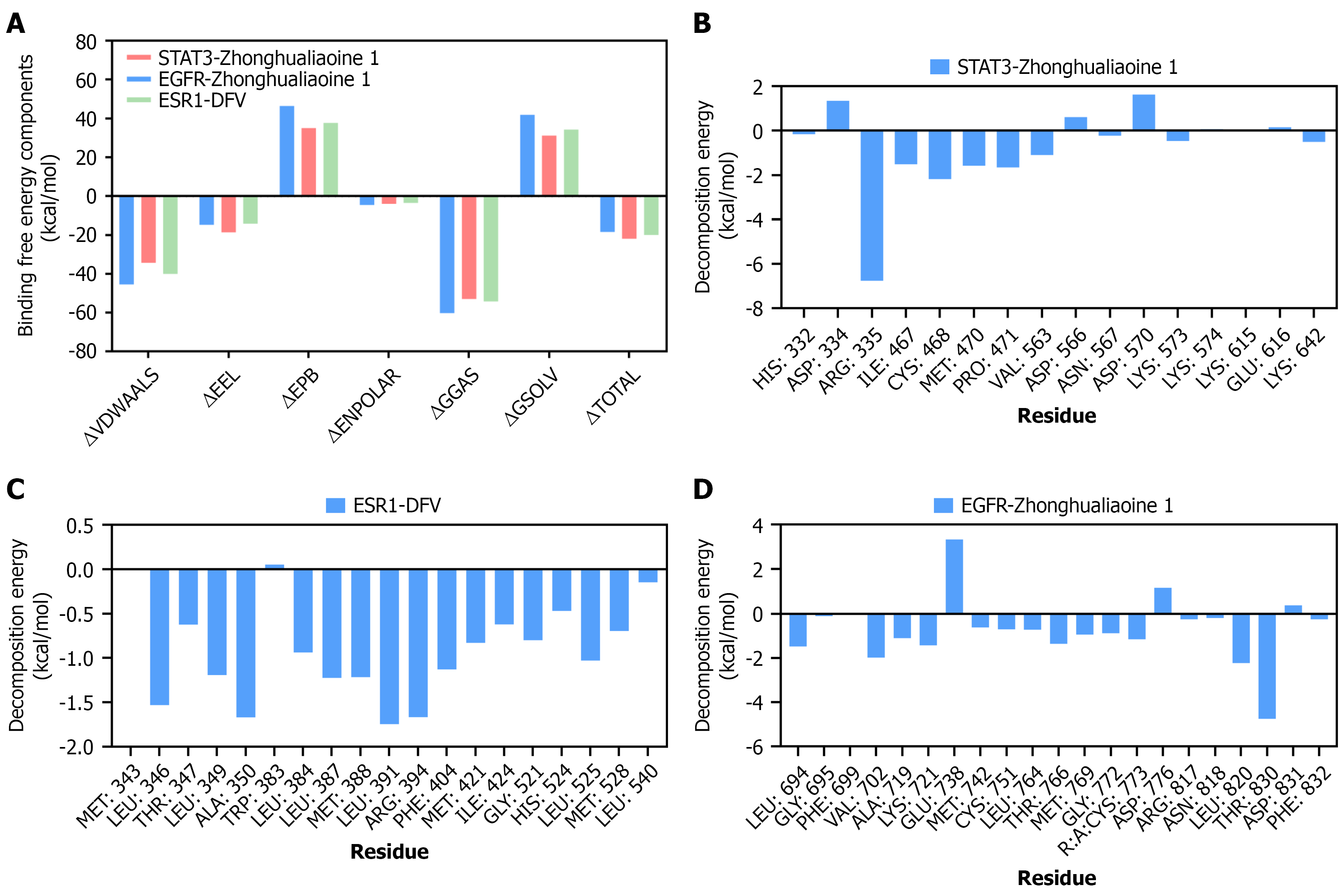
Figure 5 Binding free energy calculation via molecular mechanics-Poisson Boltzmann surface area.
A: Molecular mechanics-Poisson Boltzmann surface area binding free energy decomposition of three complexes; total energies: (1) -91.63 kcal/mol; (2) -77.07 kcal/mol; and (3) -83.93 kcal/mol; B: Residue decomposition energy of signal transducer and activator of transcription 3-zhonghualiaoine 1 complex; C: Residue decomposition energy of estrogen receptor 1-diosgenin-3-O-β-D-fructofuranosyl-α-L-rhamnopyranoside complex; D: Residue decomposition energy of epidermal growth factor receptor-zhonghualiaoine 1 complex. Binding free energy of protein-ligand complexes calculated by g_mmpbsa package (solvent model = Poisson Boltzmann surface area, ionic strength = 0.15 M). Data: mean ± SD (n = 3 independent simulations). DFV: Diosgenin-3-O-β-D-fructofuranosyl-α-L-rhamnopyranoside; EGFR: Epidermal growth factor receptor; ESR1: Estrogen receptor 1; STAT3: Signal transducer and activator of transcription 3.
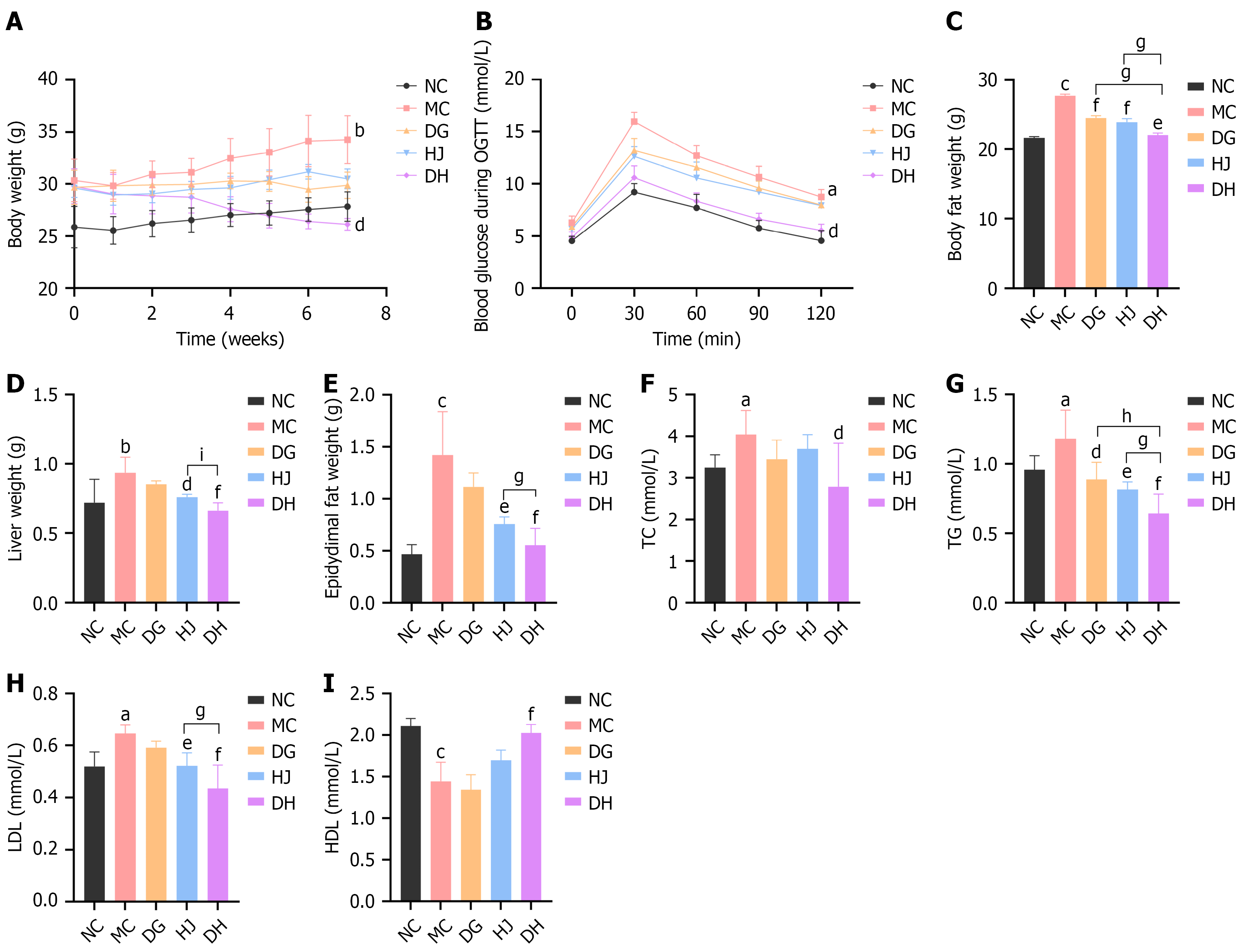
Figure 6 Danggui-Huangjing improves the phenotype and histology of obese C57BL/6 mice.
A: Body mass curves (n = 6/group). Mice fed standard chow (normal control) or high-fat diet (45% fat, model control/Danggui/Huangjing/Danggui-Huangjing) for 18 weeks (induction: Weeks 1-10; treatment: Weeks 11-18); Danggui/Huangjing/Danggui-Huangjing: Intragastric administration (1.82 g/kg/day); B: Oral glucose tolerance testing results (n = 6/group). Fasted 12 hours, glucose (2 g/kg, oral gavage), blood glucose measured at 0/30/60/90/120 minutes, area under the curve (0-120) via trapezoidal rule; C-E: Body fat percentage (echo magnetic resonance imaging 100H), liver, and epididymal white adipose tissue masses (n = 6/group); F-I: Serum total cholesterol, triglyceride, high-density lipoprotein-cholesterol, low-density lipoprotein-cholesterol (n = 6/group), measured by BS-420 analyzer with commercial kits. Data are presented as mean ± SD and were analyzed by one-way analysis of variance followed by the Bonferroni post hoc test. aP < 0.05, bP < 0.01, cP < 0.001 vs normal control group; dP < 0.05, eP < 0.01, fP < 0.001 vs model control group; gP < 0.05, hP < 0.01, iP < 0.001 vs Danggui-Huangjing group. DG: Danggui; DH: Danggui-Huangjing; HDL: High-density lipoprotein; HJ: Huangjing; LDL: Low-density lipoprotein; MC: Model control; NC: Normal control; OGTT: Oral glucose tolerance testing; TC: Total cholesterol; TG: Triglyceride.
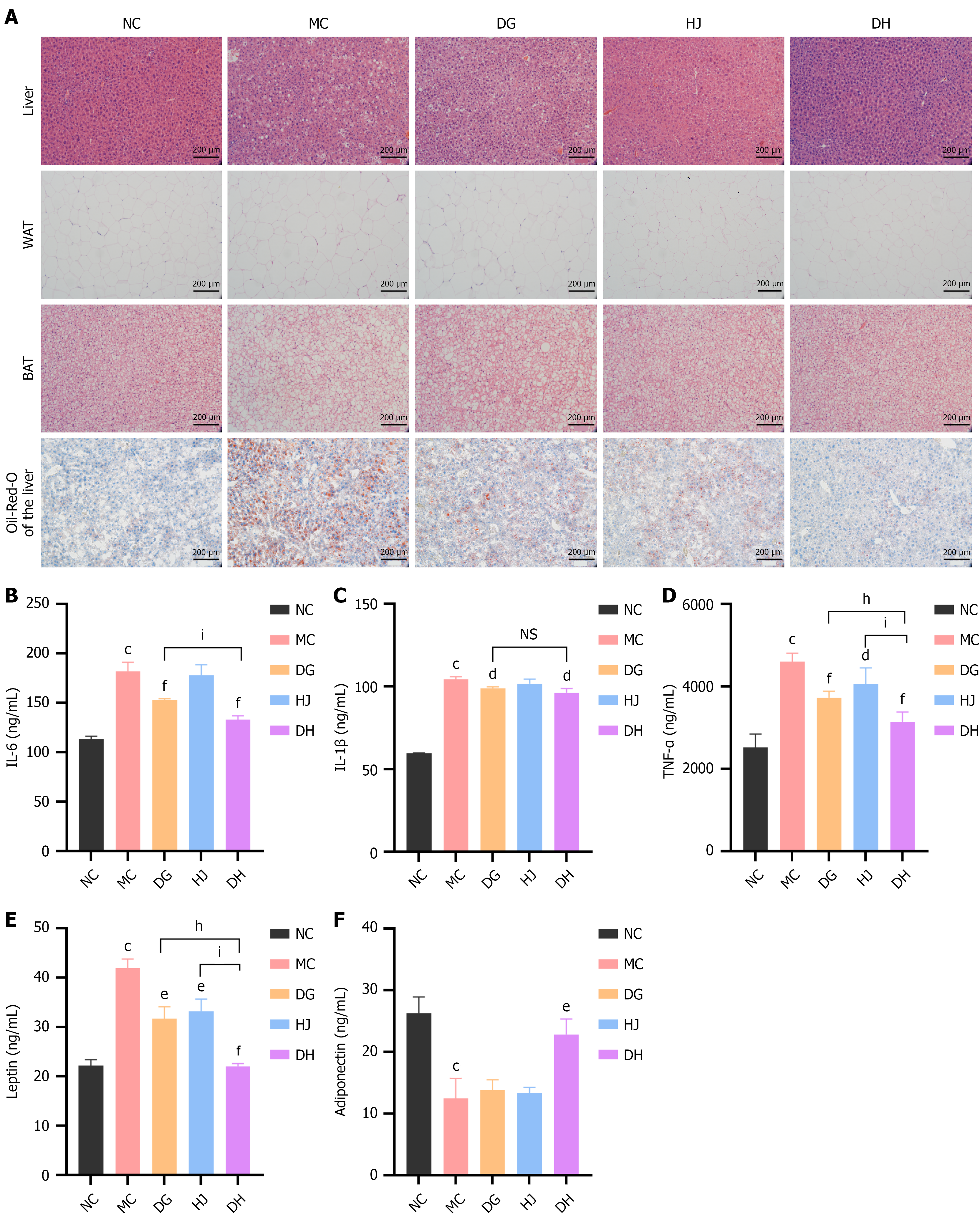
Figure 7 Danggui-Huangjing alleviates adipose inflammation and hepatic steatosis in obese C57BL/6 mice.
A: Representative micrographs (200 ×): Hematoxylin and eosin/Oil Red O-stained sections (processing as Figure 6A); B-F: Serum interleukin-6, interleukin-1β, tumor necrosis factor-α, leptin, adiponectin (n = 6/group), measured by enzyme-linked immunosorbent assay kits (Ruixin Biotechnology) with E9032 microplate reader (duplicate assays). Data are presented as mean ± SD and were analyzed by one-way analysis of variance followed by the Bonferroni post hoc test. aP < 0.05, bP < 0.01, cP < 0.001 vs normal control group; dP < 0.05, eP < 0.01, fP < 0.001 vs model control group; gP < 0.05, hP < 0.01, iP < 0.001 vs Danggui-Huangjing group. NS: Not significant (for Danggui-Huangjing vs Danggui/Huangjing groups, no statistical difference). BAT: Brown adipose tissue; DG: Danggui; DH: Danggui-Huangjing; HJ: Huangjing; IL: Interleukin; MC: Model control; NC: Normal control; TNF-α: Tumor necrosis factor-α; WAT: White adipose tissue.
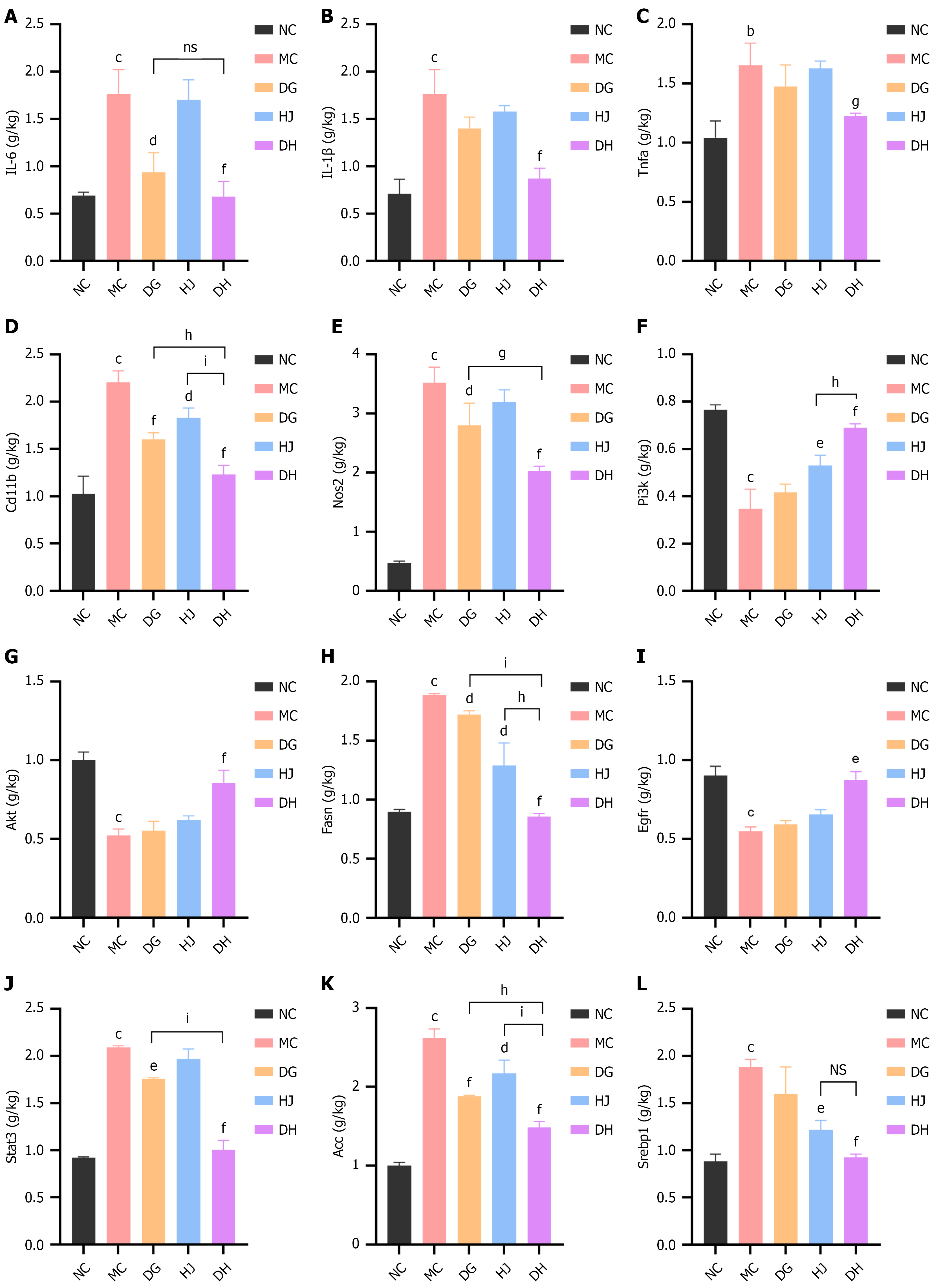
Figure 8 Danggui-Huangjing robustly reduces the expression of inflammation and lipogenesis-related genes in obese C57BL/6 mice.
A: Relative hepatic mRNA expression of IL6; B: Relative hepatic mRNA expression of IL1b; C: Relative hepatic mRNA expression of Tnfa; D: Relative hepatic mRNA expression of Cd11b; E: Relative hepatic mRNA expression of Nos2; F: Relative hepatic mRNA expression of Pi3k; G: Relative hepatic mRNA expression of Akt; H: Relative hepatic mRNA expression of Fasn; I: Relative hepatic mRNA expression of Egfr; J: Relative hepatic mRNA expression of Stat3; K: Relative hepatic mRNA expression of Acc; L: Relative hepatic mRNA expression of Srebp1. RNA extraction (AG21017 kit), complementary DNA synthesis (AG11728 kit), qPCR (StepOne Plus system, SYBR Green kit AG11739); β-Actin as internal reference, relative expression via 2-ΔΔCt method. Data are presented as mean ± SD and were analyzed by one-way analysis of variance followed by the Bonferroni post hoc test (n = 6/group). aP < 0.05, bP < 0.01, cP < 0.001 vs normal control group; dP < 0.05, eP < 0.01, fP < 0.001 vs model control group; gP < 0.05, hP < 0.01, iP < 0.001 vs Danggui-Huangjing group. NS: Not significant (for Danggui-Huangjing vs Danggui/Huangjing groups, no statistical difference). DG: Danggui; DH: Danggui-Huangjing; HJ: Huangjing; MC: Model control; NC: Normal control.
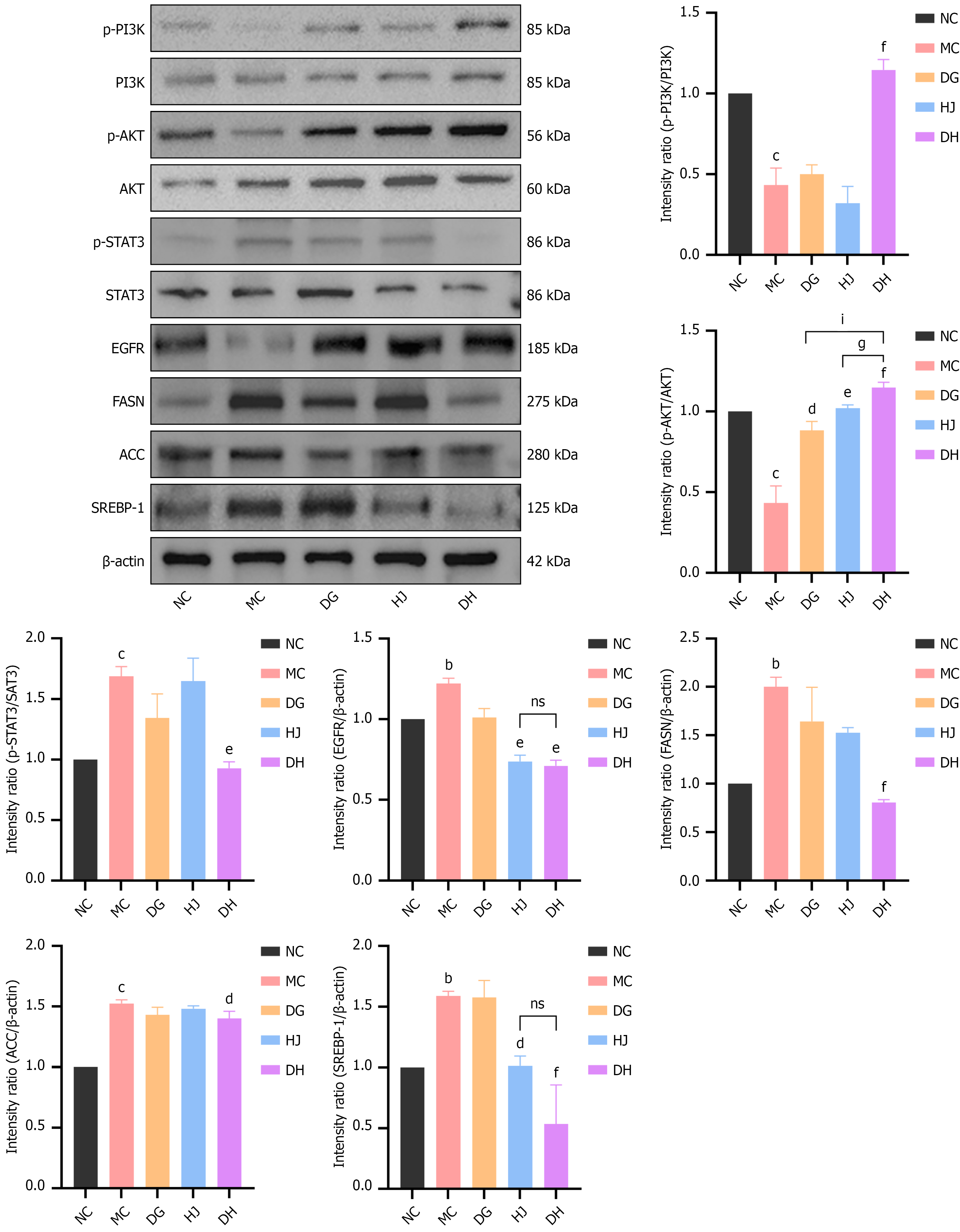
Figure 9 Expression of relevant proteins in the livers of C57BL/6 mice from the various groups.
Representative western blot images and quantitative analysis of phosphorylated phosphatidylinositol 3-kinase (PI3K), PI3K, phosphorylated protein kinase B, protein kinase B, phosphorylated signal transducer and activator of transcription 3, signal transducer and activator of transcription 3, epidermal growth factor receptor, fatty acid synthase, acetyl-CoA carboxylase, sterol regulatory element-binding protein 1 in liver tissues (n = 3/group). Liver lysates: RIPA buffer (protease inhibitors), 20 μg protein loaded per lane; primary antibodies from Cell Signaling Technology (phosphorylated PI3K No. 4228, etc.) and Abcam (anti-acetyl-CoA carboxylase, No. ab45174, etc.); β-actin as internal reference; bands quantified by ImageJ. Data are presented as mean ± SD and were analyzed by one-way analysis of variance followed by the Bonferroni post hoc test. aP < 0.05, bP < 0.01, cP < 0.001 vs normal control group; dP < 0.05, eP < 0.01, fP < 0.001 vs model control group; gP < 0.05, hP < 0.01, iP < 0.001 vs Danggui-Huangjing group. NS: Not significant (for Danggui-Huangjing vs Danggui/Huangjing groups, no statistical difference). ACC: Acetyl-CoA carboxylase; AKT: Protein kinase B; DG: Danggui; DH: Danggui-Huangjing; HJ: Huangjing; EGFR: Epidermal growth factor receptor; FASN: Fatty acid synthase; MC: Model control; NC: Normal control; PI3K: Phosphatidylinositol 3-kinase; p-AKT: Phosphorylated protein kinase B; p-PI3K: Phosphorylated phosphatidylinositol 3-kinase; p-STAT3: Phosphorylated signal transducer and activator of transcription 3; SREBP-1: Sterol regulatory element-binding protein 1; STAT3: Signal transducer and activator of transcription 3.
- Citation: Li WJ, Xu JJ, Tiang HS, Liu TH, Wu LL. Network pharmacology: Anti-obesity and insulin-sensitizing effects of Danggui-Huangjing herb-pair via epidermal growth factor receptor/phosphatidylinositol 3-kinase/protein kinase B pathway. World J Diabetes 2026; 17(5): 118275
- URL: https://www.wjgnet.com/1948-9358/full/v17/i5/118275.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i5.118275