Copyright: ©Author(s) 2026.
World J Diabetes. Mar 15, 2026; 17(3): 115433
Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.115433
Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.115433
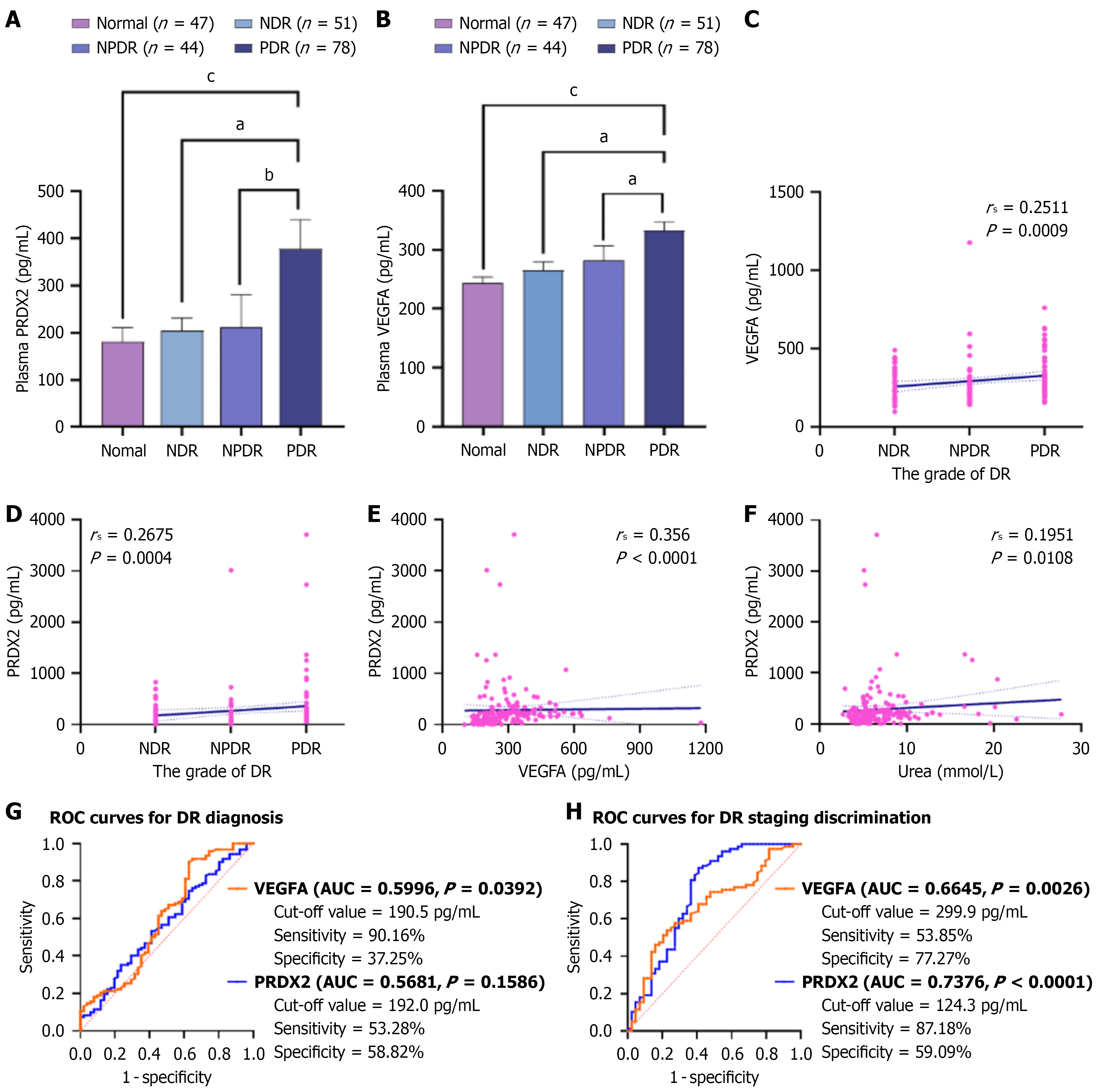
Figure 1 Biomarker profiling of plasma PPDX2 and vascular endothelial growth factor A in diabetic retinopathy progression.
A and B: Plasma PRDX2 and vascular endothelial growth factor A (VEGFA) concentrations across study cohorts: Healthy controls (normal, n = 47), non-diabetic retinopathy (n = 51), nonproliferative diabetic retinopathy (DR; n = 44), proliferative DR (n = 78). Kruskal-Wallis test for trend; C and D: Spearman correlations between DR severity and plasma VEGFA (C) or PRDX2 (D) levels; E and F: Spearman correlations between plasma PRDX2 and plasma VEGFA (E) or serum urea (F); G: Receiver operating characteristic (ROC) curves for DR diagnosis using plasma VEGFA [area under the curve (AUC) = 0.5996] and PRDX2 (AUC = 0.5681) in the diabetic cohort (n = 173); H: ROC curves for DR staging discrimination using plasma VEGFA (AUC = 0.6645) and PRDX2 (AUC = 0.7376) in confirmed DR cases (n = 122). aP < 0.05; bP < 0.0001; cP < 0.001. DR: Diabetic retinopathy; NDR: Non-diabetic retinopathy; PDR: Proliferative diabetic retinopathy; NPDR: Non-proliferative diabetic retinopathy; VEGFA: Vascular endothelial growth factor; ROC: Receiver operating characteristic; DR: Diabetic retinopathy; AUC: Area under the curve.
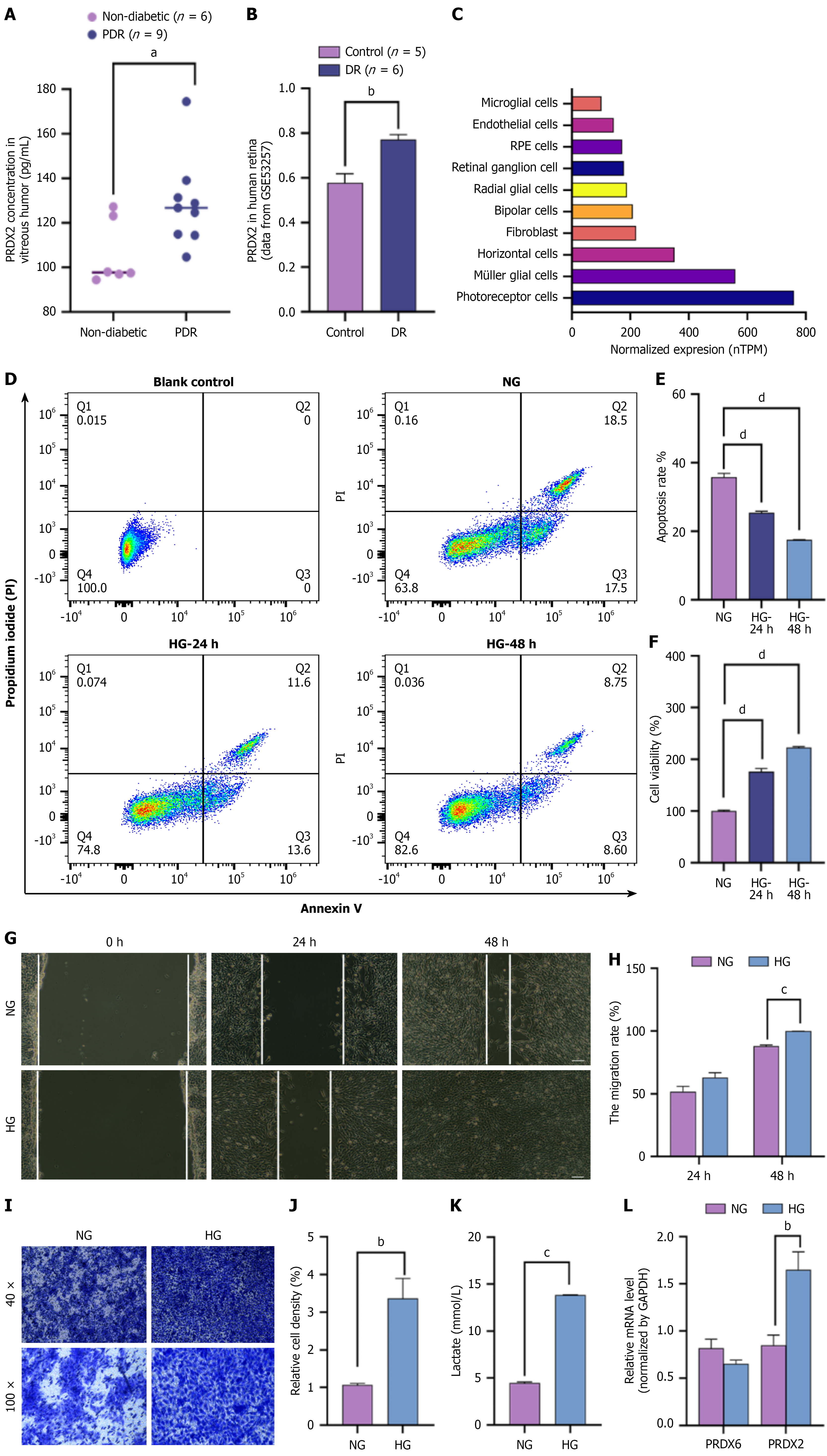
Figure 2 PRDX2 is upregulated in ocular tissues of patients with diabetic retinopathy and high-glucose-induced Müller cells.
A: Vitreous PRDX2 protein in proliferative diabetic retinopathy (DR; n = 9) vs non-diabetic controls (n = 6; ELISA); B: Retinal PRDX2 expression in six DR patients vs five healthy controls (GSE53257); C: PRDX2 gene expression in the single cell type clusters in the human retina from the Human Protein Atlas; D-L: RMC-1 cells were treated with high glucose (25 mmol/L) or normal glucose (5.5 mmol/L) for 24 hours or 48 hours to mimic diabetes-related stress; Apoptosis by Annexin V/PI flow cytometry. The bar histogram shows the quantitative results of flow cytometry. Living cell populations are clustered in the Q4 quadrant; apoptotic or necrotic cells are in the Q2 and Q3 quadrants (D and E). Cell viability of rMC-1 cells by CCK8 assay (F). Migration of rMC-1 cells by wound-healing assay (G and H; magnification 100 ×; scale bars: 100 μm). The bar histogram shows the percentage of the cell migration area. Photomicrographs (40 × and 100 ×) show the invasive ability of rMC-1 cells as measured by the Transwell chamber assay (I and J). Lactate content of rMC-1 cells (K). mRNA expression of PRDX2 and PRDX6 measured by quantitative PCR (L). All values are presented as means ± SEM. Each experiment was independently repeated 3-5 times. P value for trends assessed using the t-test or ordinary one-way analysis of variance. aP < 0.05; bP < 0.01; cP < 0.001; dP < 0.0001. PDR: Proliferative diabetic retinopathy; DR: Diabetic retinopathy; NG: Normal glucose; HG: High glucose.
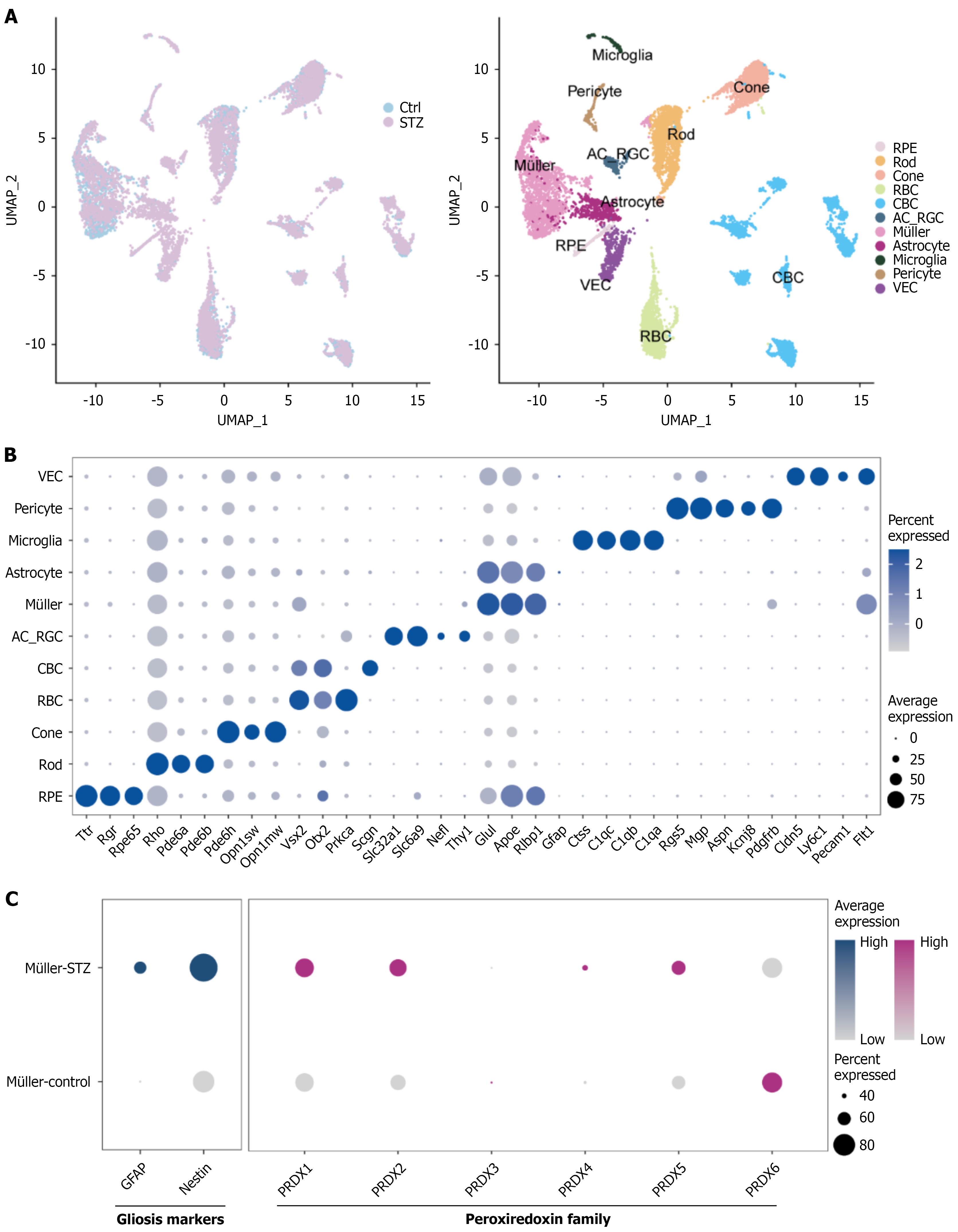
Figure 3 Single-cell transcriptomic profiling of diabetic murine retinas.
A: Uniform Manifold Approximation and Projection visualization of integrated retinal single-cell data from control vs streptozotocin (STZ)-induced diabetic mice (GEO: GSE178121). Left: Batch-corrected integration output. Right: Annotated clusters of 11 transcriptionally distinct retinal cell types; B: Dot plot representation of canonical retinal cell-type marker expression across clusters. Circle size indicates percentage of cells expressing the marker within each cluster; color intensity denotes average expression level (log-normalized counts); C: Differential expression heatmap of Müller gliosis markers (GFAP and nestin) and peroxiredoxin family genes (PRDX1-PRDX6) in Müller cells from STZ-diabetic vs control mice. The color scale represents the average expression level. STZ: Streptozotocin.
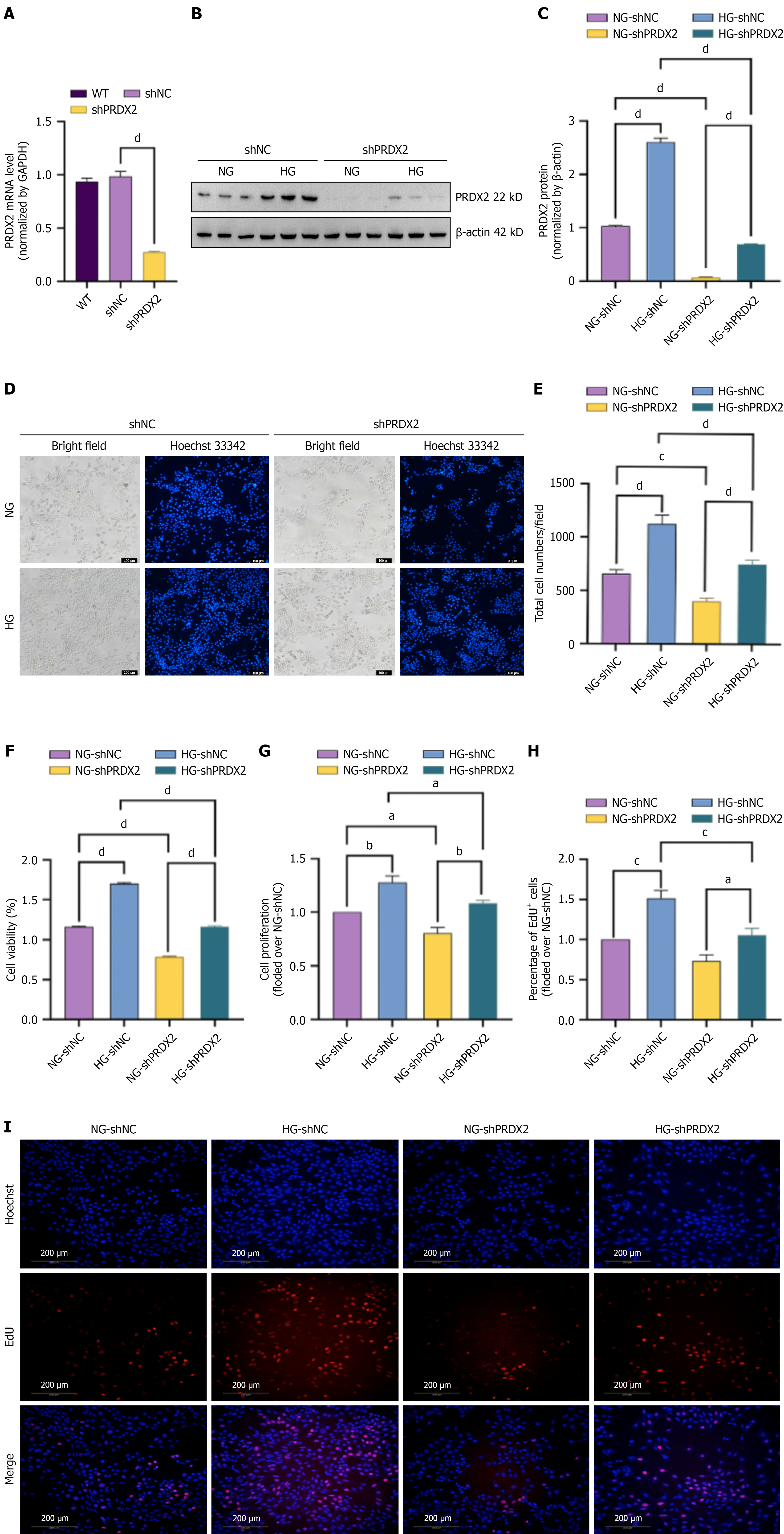
Figure 4 PPDX2 silencing attenuates high glucose-induced Müller cell proliferation.
rMC-1 cells transfected with control (shNC) or PRDX2 shRNA (shPRDX2) before being subjected to high glucose or normal glucose for 48 hours. A: mRNA expression of PRDX2 measured by quantitative polymerase chain reaction in wild-type, shNC, and shPRDX2 rMC-1 cells; B and C: Western blotting analysis of PRDX2 in rMC-1 cells. The bar histogram is representative of measurements of the gray values of PRDX2/β-actin (C); D and E: Hoechst staining was used to measure the total cell number (magnification: 100 ×; scale bars: 100 μm; blue: Hoechst); F: Cell viability as measured by CCK8 assay; G: Cell proliferation was measured by the CyQUANT NF cell proliferation assay kit; H and I: Cell proliferation detected by EdU staining (magnification: 200 ×; scale bars: 200 μm; red: EdU; blue: Hoechst). Bar histogram represents the quantitative results of rMC-1 cell proliferation rate under EdU staining (H). All values are presented as means ± SEM. Each experiment was independently repeated 3-5 times. P value for trends assessed using ordinary one-way analysis of variance. aP < 0.05; bP < 0.01; cP < 0.001; dP < 0.0001. WT: Wild type; NG: Normal glucose; HG: High glucose.
Figure 5 Proteome profiling of PPDX2-silenced Müller cells.
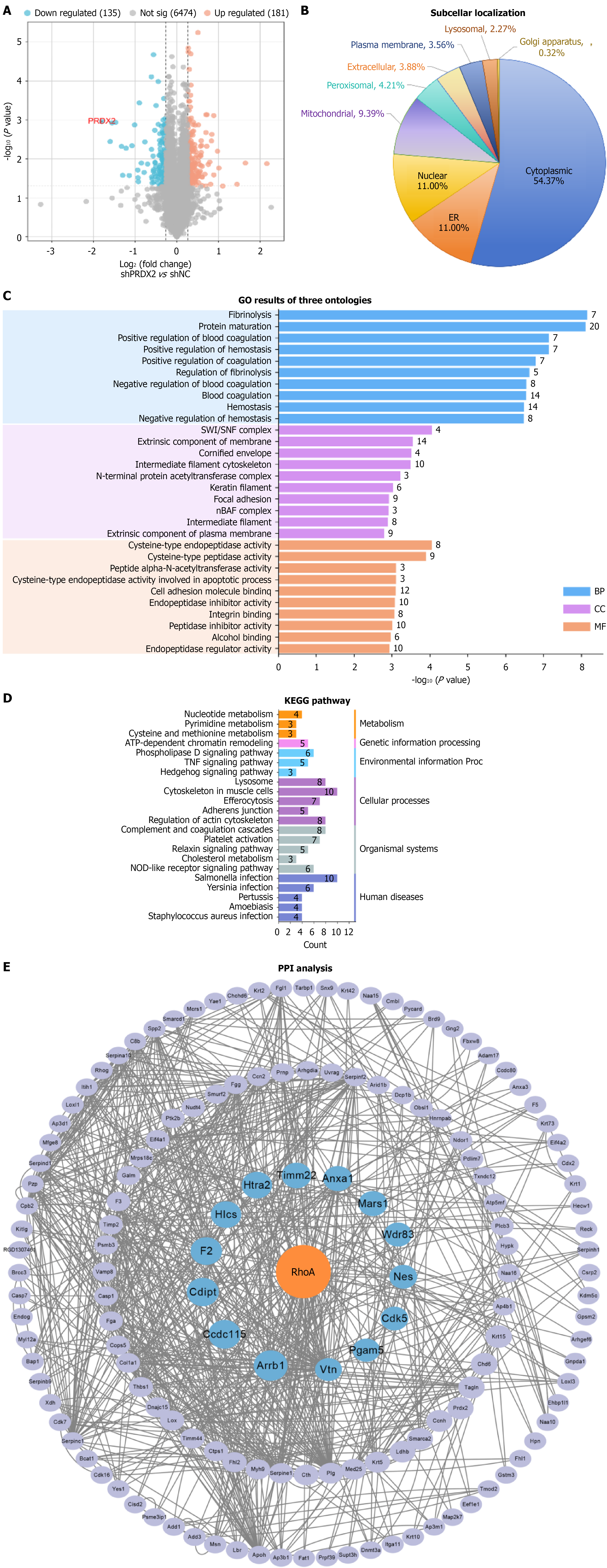
Tandem mass tag quantitative proteomics analysis of rMC-1 cells transfected with PRDX2 shRNA (n = 3) vs shNC (n = 3). A: Volcano plot of 316 significantly differentially expressed proteins [DEPs; fold change > 1.2 and P < 0.05; X axis: Log2 (fold change), Y axis: -log10 (P value); gray: Nonsignificant; orange: Upregulated; blue: Downregulated]; B: Subcellular localization of 316 DEPs; C: Gene Ontology (GO) annotation of the DEPs. The total DEPs were evaluated using biological processes, cellular components, and molecular function terms (X axis: P value; Y axis: GO categories); D: Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis of the DEPs (X axis: Number of proteins; Y axis: Pathway categories); E: Protein-protein interaction (PPI) network of DEPs. Hub proteins were selected using the Cytoscape tool. Proteins are labeled with gene names, and the circular size indicates their ranking in the PPI analysis. RhoA protein was the hub node for all nodes. KEGG: Kyoto Encyclopedia of Genes and Genomes; GO: Gene Ontology; PPI: Protein-protein interaction; BP: Biological process; CC: Cellular components; MF: Molecular function.
Figure 6 PRDX2 silencing inhibits RhoA/ROCK signaling in Müller cells.
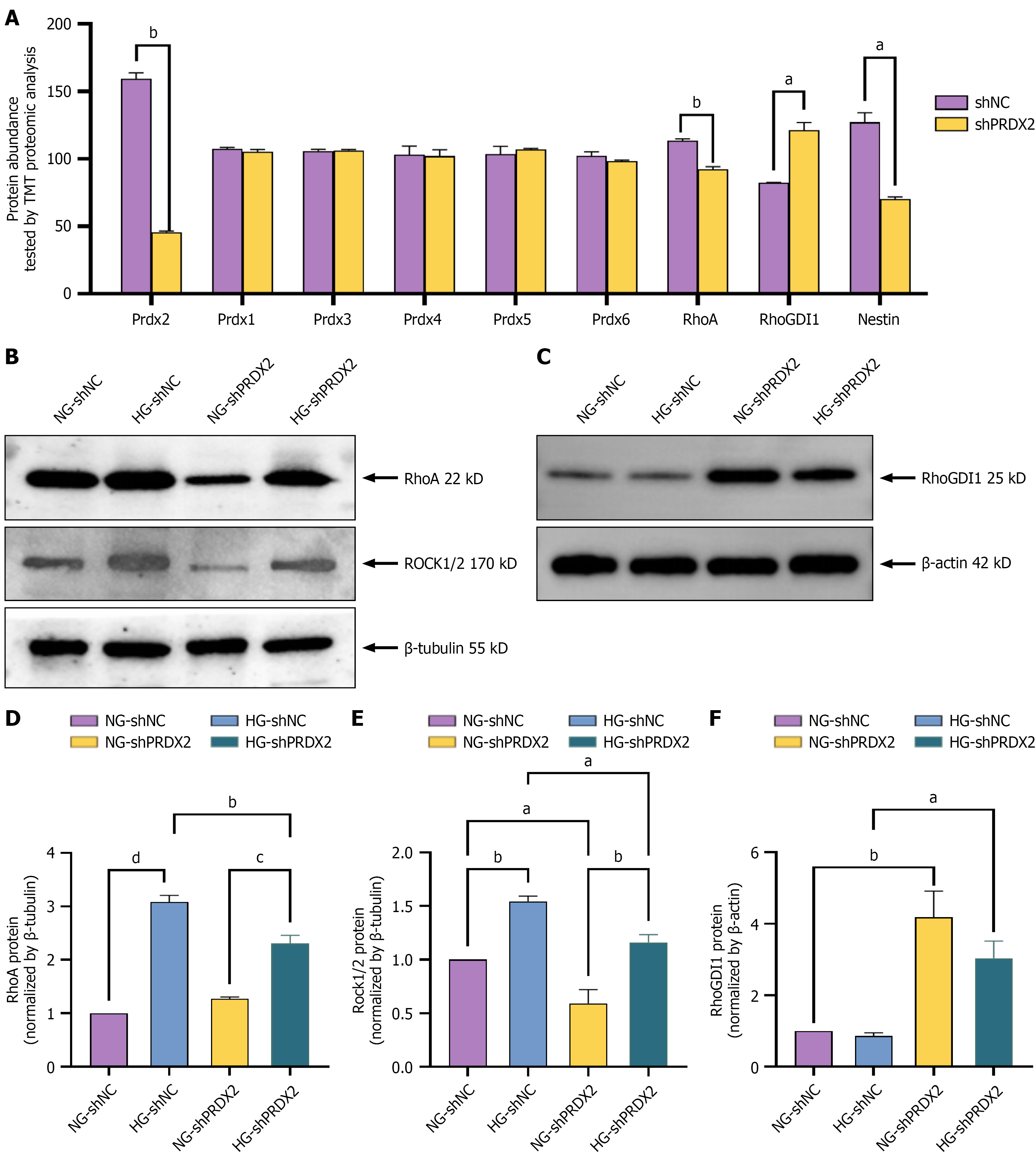
A: Tandem mass tagging proteomics of PRDX family proteins, RhoA, RhoGDI1, nestin in shPRDX2 and shNC groups; B and C: Protein expression of PRDX2, RhoA, ROCK1/2, and RhoGDI1 between shPRDX2 and shNC groups after high glucose- or normal glucose-treated rMC-1 cells for 48 hours by western blotting; D-F: The bar histograms represent measurements of the target protein's gray values. All values are presented as means ± SEM. Each experiment was independently repeated three times. P for trends assessed using the t test or ordinary one-way analysis of variance. aP < 0.05; bP < 0.01; cP < 0.001; dP < 0.0001. TMT: Tandem mass tag; NG: Normal glucose; HG: High glucose.
- Citation: Du XL, Ouyang S, Wang YS, Mao YS, Ren BC, Yu WH. PRDX2 silencing alleviates reactive hyperplasia of Müller glial cells in diabetic retinopathy by inhibiting the RhoA/ROCK signaling pathway. World J Diabetes 2026; 17(3): 115433
- URL: https://www.wjgnet.com/1948-9358/full/v17/i3/115433.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i3.115433