Published online May 15, 2026. doi: 10.4251/wjgo.v18.i5.117323
Revised: December 29, 2025
Accepted: February 11, 2026
Published online: May 15, 2026
Processing time: 161 Days and 2.8 Hours
Metastasis-associated colon cancer 1 (MACC1) is known to promote tumor grow
To systematically evaluate the multiomics characteristics and clinical significance of MACC1 across cancers and to investigate its biological functions and molecular mechanisms in PC.
MACC1 expression, DNA methylation, copy number changes, and genetic alterations were analyzed across The Cancer Genome Atlas, Genotype-Tissue Expression, Gene Expression Omnibus, cBioPortal, TIMER2.0, CancerSEA, and single-cell datasets. The diagnostic and prognostic value of MACC1 was assessed using Cox models, Kaplan-Meier analysis, and receiver operating characteristic curves. Functional studies were performed by silencing or overexpressing MACC1 in PANC-1 cell and ASPC-1 cell. Cell proliferation, colony formation, migration, invasion, and apoptosis were evaluated. Enriched pathways were screened using gene set enrichment analyses and confirmed by western blot.
MACC1 was significantly upregulated in PC at both mRNA and protein levels in public datasets and our local cohort. High MACC1 expression was linked to advanced stage, poor differentiation, and worse survival. Multiomics analysis showed that MACC1 alterations were common across cancers and were associated with copy number gain and promoter hypomethylation. Functional assays demonstrated that MACC1 enhanced pancreatic tumor proliferation, colony formation, migration, and invasion while reducing apoptosis. Gene set enrichment analyses, dual luciferase gene assay and western blot confirmed that MACC1 activated p53-related and Notch-related signaling in PC cells.
MACC1 is a clinically relevant oncogene in PC. Its overexpression promotes aggressive tumor behavior through activation of p53 and Notch pathways. These findings support MACC1 as a potential diagnostic marker and therapeutic target in PC.
Core Tip: Metastasis-associated colon cancer 1 (MACC1) is strongly upregulated in pancreatic cancer (PC) and shows consistent alterations across multiomics datasets. In this study, we combined pan-cancer bioinformatic profiling with functional experiments to clarify its role in PC. Silencing MACC1 suppressed proliferation, migration, invasion, and increased apoptosis, while overexpression produced the opposite effects. We also confirmed that MACC1 activates both p53 and Notch signaling. These findings show that MACC1 is not only a useful diagnostic and prognostic marker but also a potential therapeutic target in PC.
- Citation: Cong L, Tong S, Xu YJ, Zhang DY, Zhang XM, Yu H. Metastasis-associated colon cancer 1 drives pancreatic cancer via p53/Notch signaling. World J Gastrointest Oncol 2026; 18(5): 117323
- URL: https://www.wjgnet.com/1948-5204/full/v18/i5/117323.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i5.117323
Pancreatic cancer (PC) is recognized as the seventh most common cause of cancer death globally, and results in more than 300000 deaths per year. It is estimated that PC will become the second leading cause of tumor-related mortality in the next 10 years[1,2]. The most common malignant PC type is known as pancreatic ductal adenocarcinoma, which is con
Metastasis-associated colon cancer 1 (MACC1) was first identified in colorectal cancer as its name, is an important regulator for tumor proliferation and metastasis[4]. It has been found that MACC1 encodes hepatocyte growth factor (HGF), regulates mTOR signaling pathway, HGF/Met signaling pathway and accelerates cancer cell to migrate, invade, and metastasize[4-6]. Moreover, six studies have discussed that the overexpression of MACC1 is significantly correlated with the poor clinical outcomes in colorectal carcinoma, hepatocellular carcinoma, lung adenocarcinoma, gastric carcinoma, esophageal cancer, and ovarian cancer[7-12]. However, the deep mechanism of the carcinogenesis of MACC1 in multiple cancers, especially the precise mRNA expression, survival analysis and mechanism of MACC1 in PC patients and PC carcinogenesis remain unclear.
High-throughput technologies (sequencing), such as gene chip-based sequencing and RNA-sequencing, gene-related analyses have been able to identify the significance of target genes from overall gene expression data. Bioinformatics based methods have not been applied for identification of function of MACC1 in multiple cancers, including the PC. Therefore, in the current study, we intended to investigate the clinical value and the pivotal role of MACC1 with targeted signal pathway in multiple cancers, especially in pancreatic carcinogenesis using pan-cancer bioinformatical analysis and experiment validation. The integrative analysis will employ state-of-the-art bioinformatics tools to sift through vast datasets from repositories like The Cancer Genome Atlas (TCGA) described as the Cancer Genome Atlas, as well as Genotype-Tissue Expression (GTEx) described as and the Genotype-Tissue Expression project, identifying patterns of MACC1 expression and its genetic alterations across different cancers. This comprehensive approach is designed to validate the oncogenic role of MACC1 and association of this gene with clinical outcomes, including survival rates and response to therapeutic application. Furthermore, by pinpointing the signaling pathways modulated by MACC1, such as the HGF/Met pathway in pancreatic carcinogenesis, our study aims to illuminate the molecular drivers of MACC1-mediated tumorigenesis. These findings may have great significance in basic research and clinical translational practice and offer a new perspective for the treatment of multiple-cancer patients, especially PC patients.
We collected the human PC tumor tissues and adjacent normal control tissues from 25 patients (age range, 45-72 years) between January 2021 and January 2023 at Changhai Hospital. A pair of PC and paracancerous tissue samples from 15 males and 10 females were collected during pancreatectomy with pathologically confirmed diagnosis. After resection, the collected tissues were snap-frozen by using liquid nitrogen and stored at -80 °C until polymerase chain reaction (PCR) assay. The patient’s age, sex, pathology grade, American Joint Committee on Cancer (AJCC) stage, lymphatic metastasis, and distant metastasis, were recorded and analyzed. This study was approved after careful review and consideration by the Ethics Committee of Changhai Hospital (No. CHEC2020-173). In addition, informed and written consent was received from the enrolled patients before the start of the study.
MACC1 mRNA expression was determined using real-time reverse transcription-PCR. For this process, first we extracted total RNA from PC samples and paired paracancerous samples through Trizol reagent (Invitrogen, United States) following the guidelines provided by the manufacturer, which was then reverse transcribed into cDNA by using a chain synthesis kit for cDNA (Vazyme, China). For preforming real-time reverse transcription-PCR experiment Light Roche 480 System was used. Amplification, cDNA was done and GAPDH was used as reference primer for all the experiments. The quantification was estimated using the ΔΔCT method. The method of bioinfomatical analysis has been provided in Supplementary material.
PC cell lines from human, including PANC-1 and ASPC-1 were acquired from Zhengzhou University Hospital, First Affiliated Hospital. The PANC-1 cell line authentication has been provided in Supplementary material, and the ASPC-1 cell line authentication has been provided in Supplementary material.
These cells were cultured in a medium known as DMEM using 10% fetal bovine serum (Gibico, United States), 100 U/mL penicillin, and 100 μg/mL streptomycin (Invitrogen, United States) under 5% CO2 and 1% O2 at 37 °C. The expe
MACC1 small interfering RNA (siRNA) and its negative control (MACC1-NC) were purchased from RiboBio (Guangzhou, China). The siRNA sequences for MACC1 were as follows: (1) Forward, 5′-UUAGGAUGCCCUCAGCUAUCACUGC-3′; and (2) Reverse, 5′-AUUGUUGUAAAGCAGAAGCACUUCC-3′. The siRNA sequences for control were as follows: (1) Forward, 5′-CCAGTTAAGAACGTCCCCAAGCG-3′; and (2) Reverse, 5′-AAGCTTGAGGTCTAGGTAATTTC-3′.
To transfect PC cells, we used Lipofectamine 3000 Reagent (Invitrogen, United States). MACC1 and control vector plasmids were also obtained from RiboBio (Guangzhou, China). We then transfected siRNA (1 μg) into ASPC-1, through Lipofectamine 3000 reagent (Invitrogen, United States), following manufacturer’s guidelines.
We used the cell counting Kit8 (DojinDo Japan) for the assessment of cell proliferation. The collection of transfected cells done for the detection of cell viability from plated in 96-well culture plates (2 × 103 per well). For colony formation assay, the cells were seeded in 35 mm culture plates at 37 °C in 5% CO2 after transfection (2 × 103 per well). After 2 weeks, 4% paraformaldehyde and 0.1% crystal violet were used to fix the colony and stained the colony. A countable colony was defined as a cluster containing ≥ 50 cells to exclude small cell aggregate. The rate of colony formation was detected through the number of stained colonies.
We performed a trans well assay to evaluate cell migration. 5 × 105 cells were planted on trans well upper chambers with 100 μL Dulbecco’s modified Eagle’s medium (DMEM) [without fetal bovine serum (FBS)] and the lower chambers was infused with 600 μL of DMEM (with 10% FBS, Gibco, United States). After the 24 hours of incubation, methanol and crystal violet were used to fix and stain the cells. The cells on the outside surfaces of the upper chambers were then recorded by microscope (Olympus, Tokyo, Japan) at 100 × magnification.
Another trans well assay with Matrigel was performed to find out the cell invasion. The trans well upper chambers coated with Matrigel (Costar, United States) were used to plate PC cells (5 × 105 per chamber). The lower chambers of the trans well were infused with 600 μL of DMEM (with 10% FBS, Gibco, United States). After the incubation for 24 hours, we used methanol, hematoxylin, and eosin to fix and stain cells, and methanol and crystal violet were used to fix and stain the cells. The cells on the outside surfaces of the upper chambers were then recorded by microscope (Olympus, Tokyo, Japan) at 100 × magnification. The apoptosis of dead cells was determined using Apoptosis Kit (KeyGEN, China). We followed the manufacturer’s guidelines on a BD Accuri C6 flow cytometer (BD Bioscience, United States).
To extract all of the protein from transfected cells, combination of a solution known as RIPA (Beyotime, China) and protease inhibitor cocktail (Roche, United States) was done. In order to measure the concentration, we utilized BCA Protein Assay Kit (Thermo, United States). Following a uniform concentration adjustment, the protein samples were separated using SDS-PAGE (10%) and then transferred to membranes of polyvinylidene fluoride. After blocking, the obtained membrane was co-incubated with a number of antibodies. The antibodies contain: (1) Anti-p53 (Abcam, ab33889, 1:1000); (2) Anti-MACC1 (Abcam, ab226803, 1:1000); (3) Anti-HES1 (Abcam, ab71559, 1:1000); (4) Anti-HEY1 (Abcam, ab22614, 1:1000); (5) Anti-Notch1 (Abcam, ab83232,1:1000); (6) Anti-BAX (Abcam, ab32503, 1:1000); (7) Anti-Bcl-2 (Abcam, ab182858, 1:1000); (8) Anti-GAPDH (Abcam, EPR16891, 1:5000); and (9) Goat anti-Rabbit HRP (Dingguo Changsheng, IH-0011). Finally, fluorescent signals were detected using an ECL chromogenic apparatus (Amersham Imager 600, United States). After the experiment, the relative gray density was calculated and normalized through Image J software (version 1.53).
To perform statistical analysis, we used webtools GraphPad 5.0, and R software (version 4.3.0). For bioinformatics, Pearson was used for normal distribution data. The Spearman correlation analysis was used for other data analysis. We used Kruskal-Wallis Rank Sum Test Wilcoxon Rank Sum and Signed Rank Tests between variables to detect differences. Cox and Kaplan-Meier survival analysis were performed with the help of survival package. The Kaplan-Meier method uses log-rank test for significance test, and the “survminer” package to visualize Kaplan-Meier analysis. Relative risks were described using hazard ratios and 95%CI. Receiver operating characteristic (ROC) curve analysis was performed using the “pROC“ package to evaluate gene diagnostic performance. All tests/statistical analyses were bilateral. For experimental validation, results were presented as mean ± SD and analyzed using unpaired Student’s t-test and one-way analysis of variance. The significant values were evaluated based on the P < 0.05.
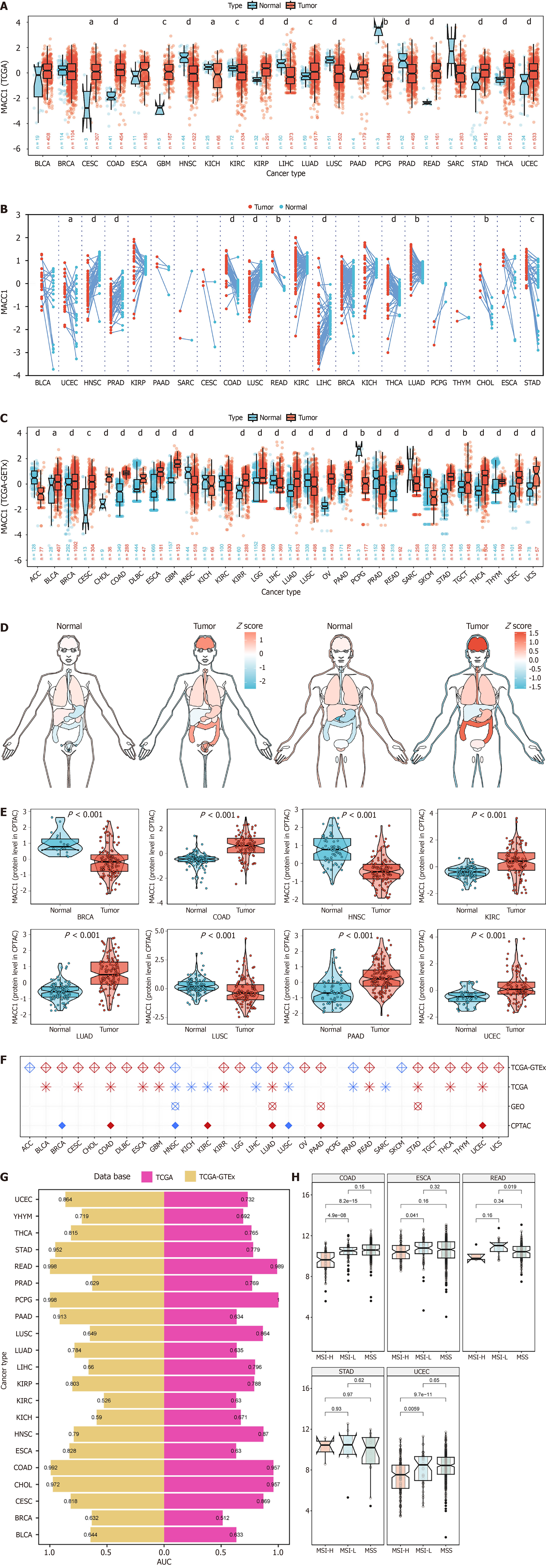
We performed difference analysis of samples received from TCGA (Figure 1A), and paired analysis of difference (Figure 1B) to determine dysregulation pattern of MACC1. Subsequently, the expression level was obtained for MACC1 from the perspective of PC to further expand the non-cancer sample size by combining TCGA and GTEx and mining these resources (Figure 1C). The distribution pattern of MACC1 expression was visualized by organ map (Figure 1D). We found that MACC1 was found expressed differentially in most types of cancer and depicted a regular and consistent pattern of expression across cancer species analyses, i.e., generally significantly upregulated. The verification of protein levels was based on the CPTAC database (Figure 1E). External verification of mRNA levels was conducted by Gene Expression Omnibus database (Supplementary Figure 1), and the above results were fully verified by regression analysis based on TCGA, TCGA-GTEx, Gene Expression Omnibus and CPTAC logistics (Figure 1F). We can observe that different omics and different databases have good consistency in the trend of data set expression. In fact, the estimated ROC curve showed (Figure 1G) that MACC1 mRNA expression levels had satisfactory sensitivity and specificity for the diagnosis of patients with multiple tumors [area under the curve (AUC) > 0.7]. MACC1 expression was correlated with 5 tumor stages (Supplementary Figure 2), suggesting that it was associated with the progression of these tumors. Interestingly, MACC1 differed across a large number of molecular subtypes (Figure 1H and Supplementary Figure 3).
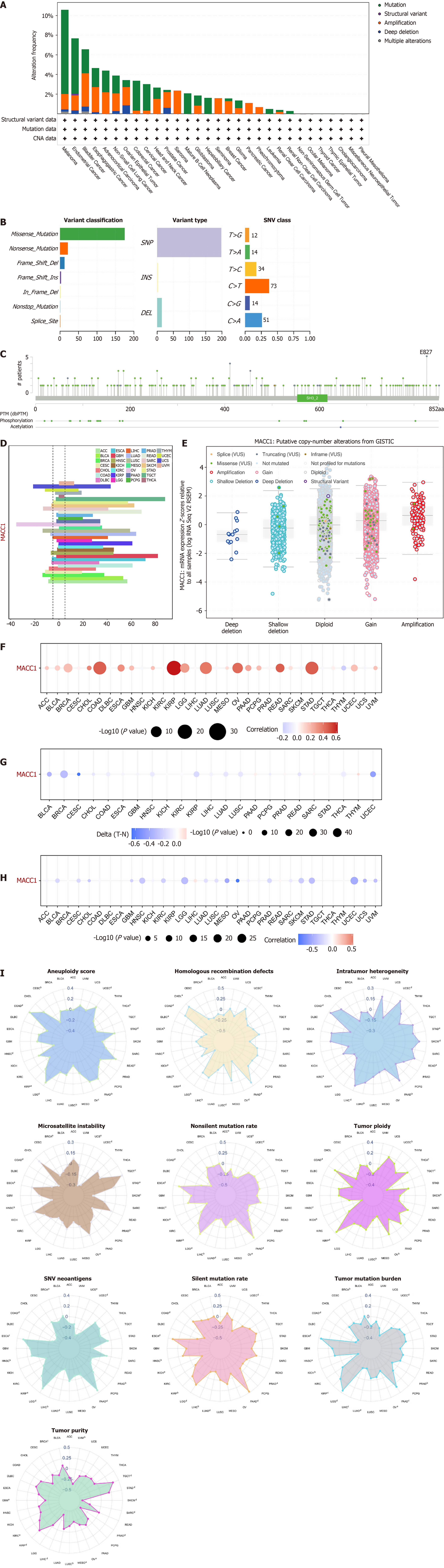
We examined genomics data of normal tumor tissues in the cohort of PC from TCGA, including genetic variation, DNA methylation data, somatic copy number alteration (SCNA), and mRNA expression data. The cBioPortal database suggests that MACC1 has a certain frequency of genetic changes in most cancers, with amplification and mutation being the most common types of genetic changes (Figure 2A). Missense mutation is the main type of mutation (Figure 2B). We systematically examined the two-dimensional mutation site structure of MACC1 (Figure 2C). To further investigate the genetic aberration of MACC1 in cancer, we examined the percentage of SCNA, which in general occurs more frequently in most types of different cancer and less frequently in only a very small tumors by numbers (Figure 2D). Clearly, SCNA plays a crucial role in the regulating gene expression in tumors (Figure 2E). We investigated the effect of SCNA on MACC1 mRNA expression by calculating the Spearman correlation. These obtained results have revealed that the MACC1 mRNA expression was significantly correlated with SCNA in different tumors types (Figure 2F). This result suggests that MACC1 copy number abnormalities are common among most of the tumors, and can affect gene expression. We found that MACC1 showed a relatively consistent methylation pattern in the pan-cancer cohort; most tumor tissues showed hypomethylation compared with normal tissues (Figure 2G). In general, MACC1 mRNA expression negatively correlates with DNA methylation as indicated by Figure 2H. Additionally, we calculated the spearman correlation of MACC1 with 10 genomics feature scores, including Aneuploidy Score, Homologous Recombination Defects, Intratumor Heterogeneity, Microsatellite Instability, Nonsilent Mutation Rate, Tumor Ploidy, SNV Neoantigens, Tumor Mutation Burden, Tumor Purity and Silent Mutation Rate (Figure 2I).
The correlation between MACC1 and major histocompatibility complex molecules, immune-activator genes, immunosuppressive genes, and chemokines showed surprising consistency, and an overall positive correlation trend was observed (Figure 3A). It is important to understand that we analyzed the anti-cancer immune status and response to each of the seven steps of the cancer immune cycle as described above. High and low MACC1 expression groups differences were calculated. Clearly, there was better consistency in pan carcinoma, i.e., a higher Tracking Tumor Immunophenotype score (Figure 3B). To elucidate the specific cell types regulated by MACC1 in the tumor microenvironment, we used the TIMER2.0 database to explore the association between MACC1 mRNA expression and immune infiltration levels and stromal cell abundance. These results suggest the involvement of MACC1 in the formation of immune rejection or infiltration of immune cells which plays a key role in immune escape (Figure 3C). The trend of this association is slightly different in different tumors, due to different proportions of immune invasion in different cancers and unique tumor microenvironment. However, the seven types of software (evaluation methods) corroborated each other and confirmed the accuracy of our analysis. In addition, multiple single-cell data sets in the TISCH database consistently described the expression landscape of MACC1 in most tumors, and the results showed that although MACC1 was not strongly expressed in most tumors, it mainly came from malignant cells, confirming the above results of immune infiltration (Figure 3D).
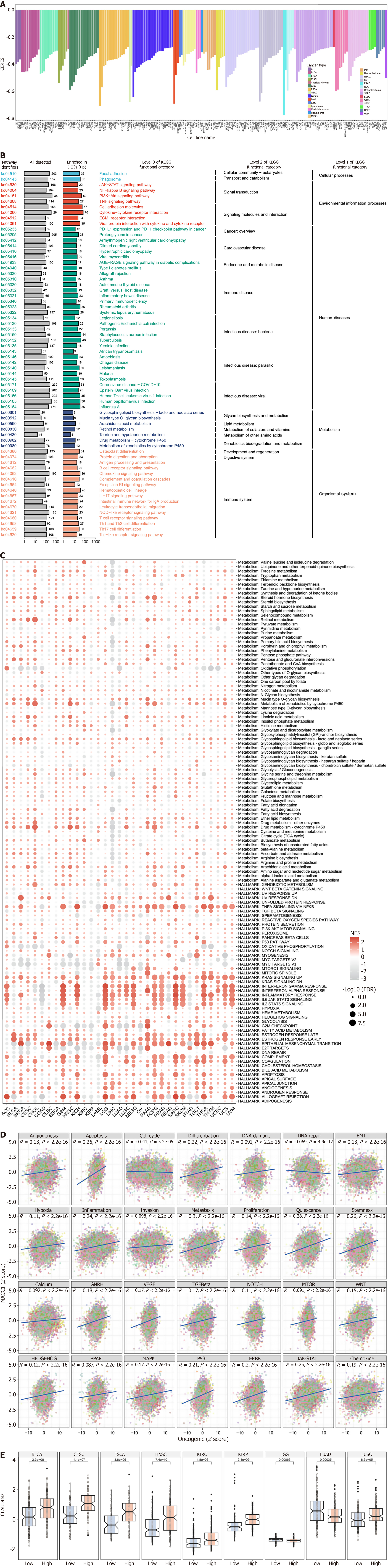
MACC1 was negative in a large number of cells, indicating that cell growth was inhibited after MACC1 knockout (Figure 4A), and cell adhesion related pathways were significantly activated in the group with high expression of MACC1 (Figure 4B). In addition, metabolic pathways analysis was carried systematically, which indicated consistency between cancer species. This implies the conservative nature of MACC1 function. We observed that immune and epithelial mesenchymal transformation pathways are often enriched in tumors with high MACC1. Moreover, MACC1 is thought to be associated with tumors and metabolism related disorders. Interestingly, MACC1 inhibits drug metabolism-related processes, which may be related to the sensitivity of chemotherapy (Figure 4C). We analyzed the relationship between 14 cancer markers and 14 tumor-related pathway scores and MACC1, and the main correlation was positive (Figure 4D). The Cancer Proteome Atlas data suggested that CLAUDIN7 had a higher expression intensity in the group with high expression of MACC1 mRNA (Figure 4E) and this finding confirmed constituency with transcriptome results.
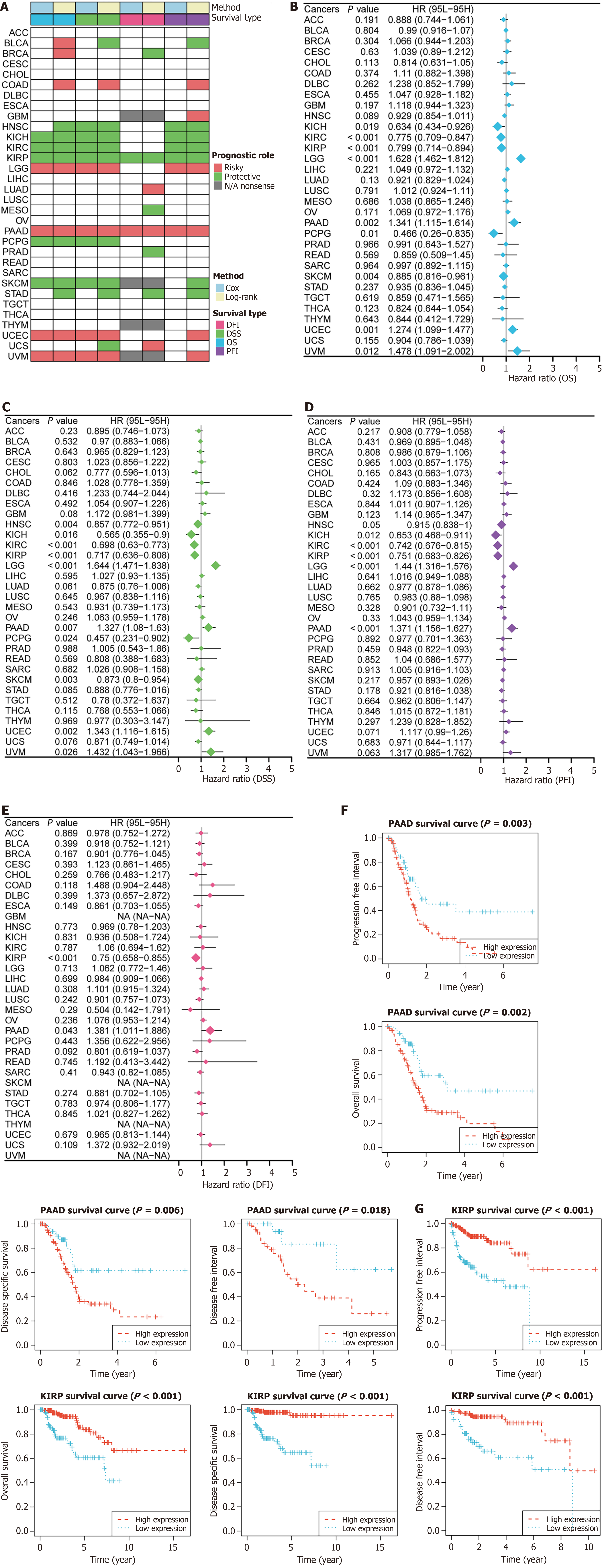
The Pan-cancer Atlas of survival analysis showed that all MACC1 was correlated with multiple survival times for multiple cancer types (Figure 5A), and the associations were relatively homogeneous, as MACC1 can be used as a protective factor for different cancers in most cases, especially kidney renal papillary cell carcinoma (KIRP), kidney chromophobe carcinoma, and kidney renal clear cell carcinoma (kidney tumors). A small number of tumors also present as risk factors, suggesting that MACC1 may play different roles in different types of cancers. Notably, MACC1 was associated with four lifetimes in KIRP and pancreatic adenocarcinoma (PAAD). Further information from the atlas of pan-cancer and results based on the analysis (cox survival) of four survival periods are shown in forest maps (Figure 5B-E), and log rank test results of KIRP and PAAD are shown in Kaplan-Meier curves (Figure 5F and G).
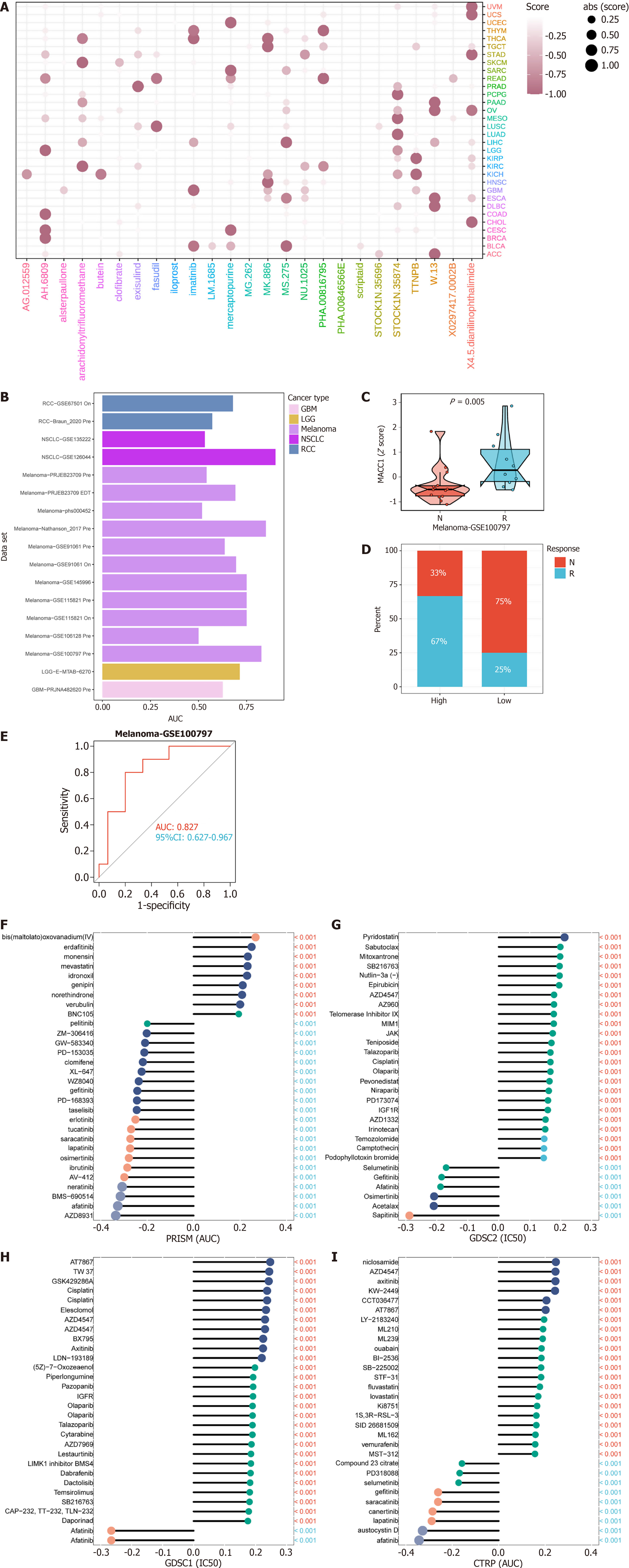
We performed CMAP analyses to identify potential treatment strategies, which can counteract MACC1-mediated tumor-promoting effects. We established an MACC1-associated gene signature, including the top-most upregulated and the most down-regulated genes, detected by the comparison of patients with low and high MACC1 expression in each cancer type. The similarity scores of 1288 compounds were obtained by comparing MACC1-related characteristics with CMAP gene characteristics using XSum, an optimal feature matching method. W.13 exhibit significantly lower scores in most cancer types, suggesting that they have the potential to inhibit MACC1-mediated cancer-promoting effects (Figure 6A). We analyzed MACC1 expression in a large number of immunotherapies, and it was clear that AUC values were not universally desirable (Figure 6B). However, we observed a skin melanoma cohort with higher MACC1 expression in the response group, a higher proportion of response patients in the MACC1 high expression group, and better sensitivity and specificity were also obtained by ROC curve analysis (Figure 6C-E). In the chemotherapy analysis, we evaluated the association between drug sensitivity and MACC1 expression using 3 different databases (Cancer Therapeutics Response Portal, genomics of drug sensitivity in cancer, and Preservice Research Institute for Science and Mathematics). Obviously, MACC1 is a potential drug resistance gene (Figure 6F-I).
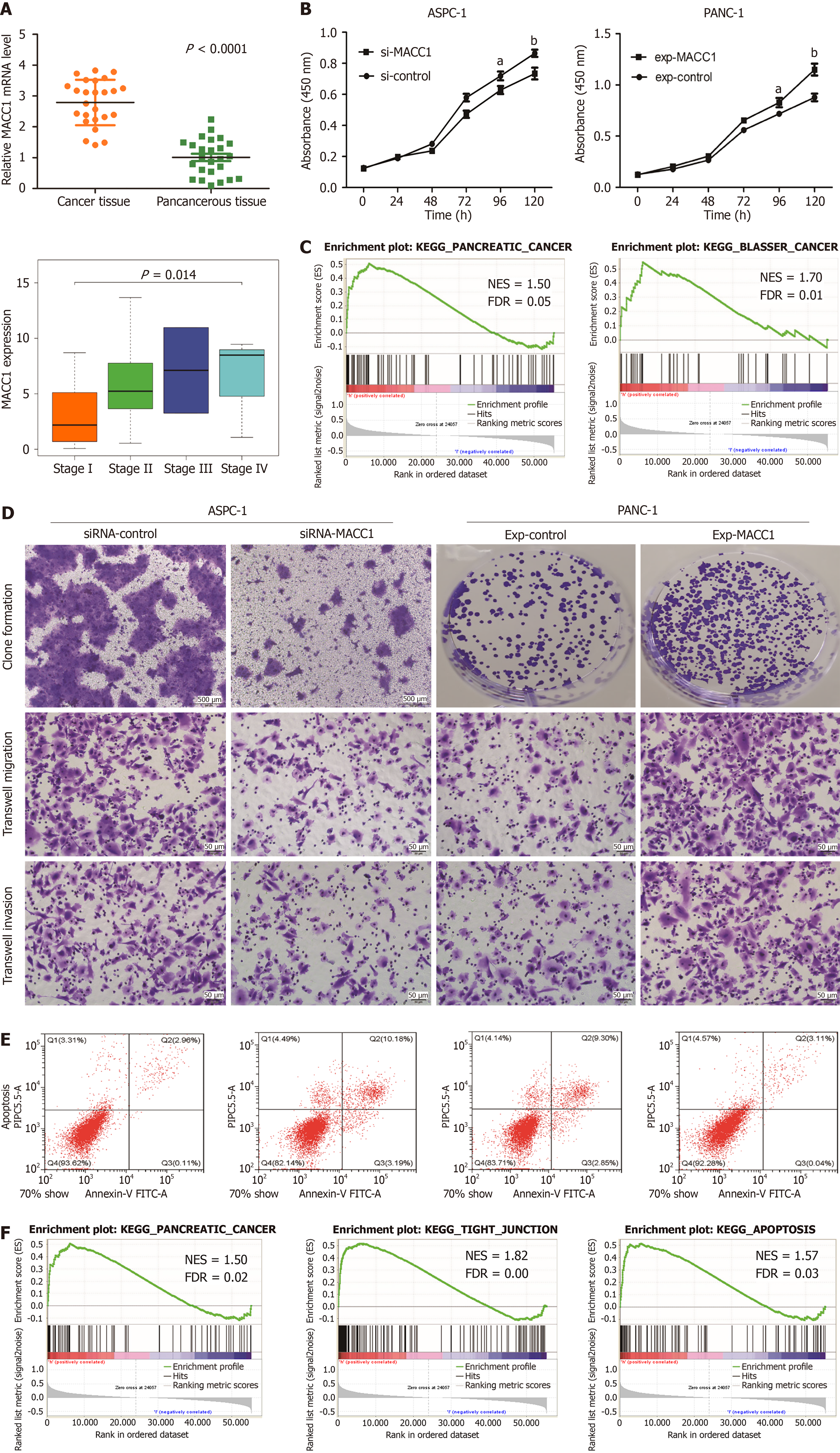
The upregulated MACC1 in PC is also verified according to our result in clinical samples between cancer tissues and normal tissues (Figure 7A left). The clinicopathologic parameters of these 25 subjects with MACC1 expression level shows upregulated MACC1 was related to poor pathology grade, higher AJCC stage and higher possibility of distant metastasis (P < 0.001; Figure 7A right, Table 1). We selected ASPC-1 for transfection with siRNA and PANC-1 to be transfected with MACC1 plasmid. The results of western blot showed that the transfection in these two cell lines was effective (Figure 7B). Overexpression of MACC1 can promote the proliferation and downregulated MACC1 can restrain the proliferation of PC cell. Moreover, according to the result of Hallmarks in gene set enrichment analyses (GSEA), the related pathway including cell cycle and PC are also enriched in TCGA-PAAD cohort (Figure 7C). The result of clone formation, transwell migration, invasion and flow cytometry indicated upregulated MACC1 can promote tumor migration, invasion and restrain apoptosis in PC cell. Down regulated MACC1 can restrain tumor migration, invasion and predispose to apoptosis in PC cell (Figure 7D and E). Interestingly, some related results of Hallmarks in GSEA term also demonstrated the potential function of MACC1 as a tumor promoter and apoptosis depressor (Figure 7F).
| Clinicopathologic parameters | Number | Metastasis-associated colon cancer 1 expression | P value |
| Sex | |||
| Male | 15 | 2.32 ± 0.52 | 0.432 |
| Female | 10 | 2.65 ± 0.35 | |
| Age | |||
| < 60 | 11 | 2.59 ± 0.47 | 0.483 |
| ≥ 60 | 14 | 2.67 ± 0.53 | |
| Pathology grade | |||
| G1 | 4 | 1.53 ± 0.43 | < 0.001 |
| G2 | 13 | 1.88 ± 0.32 | |
| G3 and G4 | 8 | 2.74 ± 0.56 | |
| American Joint Committee on Cancer stage | |||
| I and II | 11 | 1.67 ± 0.38 | < 0.001 |
| III and IV | 14 | 2.51 ± 0.42 | |
| Lymphatic metastasis | |||
| No | 8 | 2.16 ± 0.34 | 0.112 |
| Yes | 17 | 2.34 ± 0.57 | |
| Distant metastasis | |||
| No | 11 | 1.88 ± 0.48 | < 0.001 |
| Yes | 14 | 2.70 ± 0.54 |
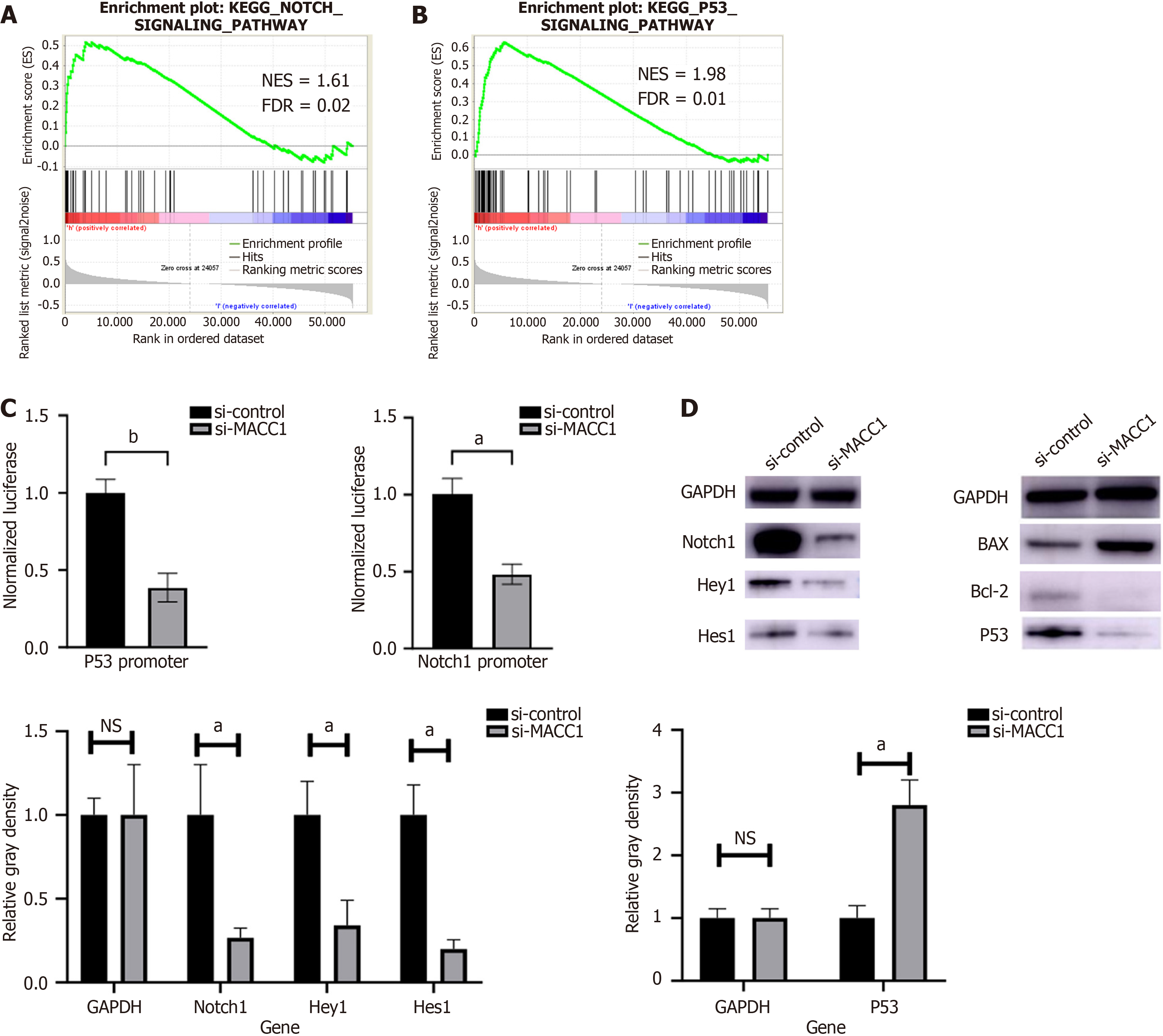
To evaluate the exact signaling pathway which MACC1 is participated in, we searched the potential signaling pathway in GSEA results above and some pathways were identified with high correlation (P < 0.05; Figure 8A and B), including p53 signaling pathway (false discovery rate = 0.01), Notch signaling pathway (false discovery rate = 0.02). Dual-luciferase reporter assays with Notch1 or p53 promoter constructs revealed attenuated transcriptional activity of both promoters upon MACC1 knockdown (Figure 8C). After verification through western blot assay, we confirmed that Notch signaling pathway related proteins, including Notch1, Hey1 and Hes1 were downregulated and p53 signaling pathway related protein, including p53 were significantly changed after knockdown of MACC1 expression level in PC cell (Figure 8D). These results indicated that MACC1 plays a crucial role in pancreatic tumorigenesis through p53 and Notch signaling pathway.
MACC1 is a novel oncogene which stimulates cancer cell to migrate and invade in vitro and vivo through HGF/Met pathway[13]. High-expression of MACC1 is associated with poor clinical outcomes in patients suffered from colorectal[14], hepatocellular[15], lung[16], gastric, esophageal[17], and ovarian[18] cancer. A recent meta-analysis has shown that overexpression of MACC1 is distinctly associated with unfavorable OS of patients with multiple solid cancers [hazard ratios = 2.11 (95%CI: 1.59-2.80), P < 0.001][19]. Another study have confirmed that over expression of MACC1 can activate PC cell chemoresistance[20]. Recently, a study reported that the MACC1 can regulate programmed death ligand-1 expression to influence tumor immunity in gastric cancer. Our data provide compelling evidence that MACC1 is overexpressed in PC and is closely associated with poor prognosis. Notably, while MACC1 exerts tumor-promoting effects in both colorectal cancer and PC, highlighting its potential as a pan-cancer oncogenic driver, their core regulatory me
PC has been reported as the seventh most common mortality causing cancer in men, and 8th in women[22]. Pancreatic ductal adenocarcinoma is the major subtype of PC which makes up the 85% of the overall PC cases. PC has the poorest prognosis among all the gastrointestinal cancer with a 5-year survival rate of 6%. The mortality rate of PC patients showed no significant improvement since 1997 until 2007[23]. The only treatment option which may provide prolonged survival is the complete surgical resection of the cancerous tissue. However, due to the lack of initial symptoms and high degree of malignancy, concrete diagnosis is made at the advanced stage of PC in most of the patients. At this stage of the diagnosis, most of patients have distance metastases or locoregional extension already[24,25]. In this grim situation, it is an urgent situation for the researchers to identify some sensitive biomarkers for PC to make timely diagnosis, and to find a potential therapeutic target. Many studies have discovered and discussed the relation between p53 and PC[26]. Notably, p53 is a well-characterized tumor suppressor gene that typically induces cell cycle arrest, apoptosis, or senescence to prevent malignant transformation. However, our findings demonstrate that MACC1 activates the p53 signaling pathway to promote PC progression, which seems contradictory to p53’s canonical tumor-suppressive function. This seeming contradiction is readily explained by the existence of gain-of-function (GOF) p53 – an oncogenic variant of p53. Accumulating evidence indicates that mutant p53, which is frequently detected in PC (with a mutation rate exceeding 70%), acquires oncogenic GOF properties that diverge from wild-type p53. Unlike wild-type p53, GOF p53 can promote cell proliferation, migration, invasion, and chemoresistance through interactions with other signaling molecules or transcriptional regulation of oncogenic target genes[27-29]. Notch signaling pathway play an important role in intercellular communication with high evolutionary conservation. Several functions can be regulated through Notch signaling pathway, and some of them are highly related to carcinogenesis including the regulation of tumor stem cell, the mediation of tumor proliferation, invasion and apoptosis[30]. To the best of our knowledge, the modulation between MACC1 and these two cancer-related signaling pathway have not been reported in any type of cancer. Our study for the first time presents these novel findings which is a significant addition to what is already available on this topic, and may provide a clinical perspective for the early diagnosis and related therapy of PC patients.
Another important spotlight of the current study is the prediction of molecular functions and pathways through bioinformatics analysis and genome-wide association study strategy with further verification through wet experiments. Since the first publication in 2005, the genome-wide association study strategy has contributed significantly to the understanding of the mechanisms of cancer[31]. GSEA is one of these important bioinformatics methods, it can facilitate researchers to predict the potential functions and activated pathways of a single gene according to the degree of variation of various genes in high-throughput sequencing profiles of certain cancers compared to normal samples. By using these methods, pathway-level changes could be identified and interpreted, and these results could lead to the orientation of further wet experiments and improve reliability of basic medical science experiment.
In our study, we firstly explore the potential role of MACC1 acted in PC. We predicted several function clusters and activated pathway and some of them have been proved in the results of functional experiment, including the proliferation, migration, invasion, apoptosis phenotype. Additionally, some critical protein in predicted pathways is also verified in western blot assay, including Notch signaling pathway, and p53 signaling pathway. The results of western blot showed that the transfection in these two cell lines were effective. Overexpression of MACC1 can promote the proliferation and downregulated MACC1 can restrain the proliferation of PC cell. Moreover, according to the result of GSEA, the related pathway including cell cycle and PC are also enriched in TCGA-PAAD cohort. The result of clone formation, transwell migration, invasion and flow cytometry indicated upregulated MACC1 can promote tumor migration, invasion and restrain apoptosis in PC cell. Down regulated MACC1 can restrain tumor migration, invasion and predispose to apoptosis in PC cell. Interestingly, some related results of GSEA also demonstrated the potential function of MACC1 as a tumor promoter and apoptosis depressor.
However, this study has some limitations as well as strength. One of the major limitations includes the lack of identification and verification of directly target of MACC1 in this pathway. The further study is needed to clarify the deep mechanism and identify the direct targets of MACC1 in p53 signaling pathway and Notch signaling pathway. The strength of our study includes bioinformatics analysis and hypothesis has been followed and confirmed by the wet experiment using our own cohort.
Our study indicated that MACC1 are significantly high-expressed in multiple cancer tissue with remarkable related changes in multi-omics, especially PC and distinctly poorly associated with AJCC stages, grades, OS and progression-free survival in PC. Overexpression of MACC1 could promote malignant phenotype of PC via activating p53 and Notch signaling pathway.
| 1. | Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359-E386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21479] [Cited by in RCA: 20392] [Article Influence: 1853.8] [Reference Citation Analysis (3)] |
| 2. | Kurdyn A, Pawłowska M, Paluszkiewicz E, Cichorek M, Augustin E. c-Myc inhibition and p21 modulation contribute to unsymmetrical bisacridines-induced apoptosis and senescence in pancreatic cancer cells. Pharmacol Rep. 2025;77:182-209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 3. | Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10534] [Cited by in RCA: 11973] [Article Influence: 1330.3] [Reference Citation Analysis (6)] |
| 4. | Stein U, Walther W, Arlt F, Schwabe H, Smith J, Fichtner I, Birchmeier W, Schlag PM. MACC1, a newly identified key regulator of HGF-MET signaling, predicts colon cancer metastasis. Nat Med. 2009;15:59-67. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 382] [Cited by in RCA: 389] [Article Influence: 22.9] [Reference Citation Analysis (1)] |
| 5. | Stein U, Dahlmann M, Walther W. MACC1 - more than metastasis? Facts and predictions about a novel gene. J Mol Med (Berl). 2010;88:11-18. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 61] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 6. | Yin Z, Ma T, Huang B, Lin L, Zhou Y, Yan J, Zou Y, Chen S. Macrophage-derived exosomal microRNA-501-3p promotes progression of pancreatic ductal adenocarcinoma through the TGFBR3-mediated TGF-β signaling pathway. J Exp Clin Cancer Res. 2019;38:310. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 130] [Cited by in RCA: 266] [Article Influence: 38.0] [Reference Citation Analysis (5)] |
| 7. | Shirahata A, Shinmura K, Kitamura Y, Sakuraba K, Yokomizo K, Goto T, Mizukami H, Saito M, Ishibashi K, Kigawa G, Nemoto H, Hibi K. MACC1 as a marker for advanced colorectal carcinoma. Anticancer Res. 2010;30:2689-2692. [PubMed] |
| 8. | Yao Y, Dou C, Lu Z, Zheng X, Liu Q. MACC1 suppresses cell apoptosis in hepatocellular carcinoma by targeting the HGF/c-MET/AKT pathway. Cell Physiol Biochem. 2015;35:983-996. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 46] [Article Influence: 4.2] [Reference Citation Analysis (3)] |
| 9. | Shimokawa H, Uramoto H, Onitsuka T, Chundong G, Hanagiri T, Oyama T, Yasumoto K. Overexpression of MACC1 mRNA in lung adenocarcinoma is associated with postoperative recurrence. J Thorac Cardiovasc Surg. 2011;141:895-898. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 83] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 10. | Wang L, Wu Y, Lin L, Liu P, Huang H, Liao W, Zheng D, Zuo Q, Sun L, Huang N, Shi M, Liao Y, Liao W. Metastasis-associated in colon cancer-1 upregulation predicts a poor prognosis of gastric cancer, and promotes tumor cell proliferation and invasion. Int J Cancer. 2013;133:1419-1430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 99] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 11. | Zhu M, Xu Y, Mao X, Gao Y, Shao L, Yan F. Overexpression of metastasis-associated in colon cancer-1 associated with poor prognosis in patients with esophageal cancer. Pathol Oncol Res. 2013;19:749-753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 29] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 12. | Jeong H, Jung J, Oh HE, Choi JW, Lee ES, Kim YS, Lee JH. Prognostic Characteristics of MACC1 Expression in Epithelial Ovarian Cancer. Biomed Res Int. 2018;2018:9207153. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 13. | Li H, Zhang H, Zhao S, Shi Y, Yao J, Zhang Y, Guo H, Liu X. Overexpression of MACC1 and the association with hepatocyte growth factor/c-Met in epithelial ovarian cancer. Oncol Lett. 2015;9:1989-1996. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 14. | Lin A, Zhang X, Zhang RL, He XF, Zhang JG, Yan WH. Prognostic and Risk Stratification Value of Lesion MACC1 Expression in Colorectal Cancer Patients. Front Oncol. 2019;9:28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 10] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 15. | Qian LQ, Li XQ, Ye PH, Su HY, Wang G, Liu Y, Shen GH, Gao QG. Downregulation of MACC1 inhibits the viability, invasion and migration and induces apoptosis in esophageal carcinoma cells through the phosphatase and tensin homolog/phosphoinositide 3-kinase/protein kinase B signaling pathway. Oncol Lett. 2017;14:4897-4905. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 16] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 16. | Xu G, Zhang B, Ye J, Cao S, Shi J, Zhao Y, Wang Y, Sang J, Yao Y, Guan W, Tao J, Feng M, Zhang W. Exosomal miRNA-139 in cancer-associated fibroblasts inhibits gastric cancer progression by repressing MMP11 expression. Int J Biol Sci. 2019;15:2320-2329. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 58] [Cited by in RCA: 134] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 17. | Wu J, Zhang D, Li J, Deng X, Liang G, Long Y, He X, Dai T, Ren D. MACC1 induces autophagy to regulate proliferation, apoptosis, migration and invasion of squamous cell carcinoma. Oncol Rep. 2017;38:2369-2377. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 19] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 18. | Yu L, Zhu B, Wu S, Zhou L, Song W, Gong X, Wang D. Evaluation of the correlation of vasculogenic mimicry, ALDH1, KiSS-1, and MACC1 in the prediction of metastasis and prognosis in ovarian carcinoma. Diagn Pathol. 2017;12:23. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 53] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 19. | Wang G, Fu Z, Li D. MACC1 overexpression and survival in solid tumors: a meta-analysis. Tumour Biol. 2015;36:1055-1065. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 20. | Wang G, Kang MX, Lu WJ, Chen Y, Zhang B, Wu YL. MACC1: A potential molecule associated with pancreatic cancer metastasis and chemoresistance. Oncol Lett. 2012;4:783-791. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 58] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 21. | Schöpe PC, Torke S, Kobelt D, Kortüm B, Treese C, Dumbani M, Güllü N, Walther W, Stein U. MACC1 revisited - an in-depth review of a master of metastasis. Biomark Res. 2024;12:146. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 22. | Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18589] [Cited by in RCA: 21024] [Article Influence: 1911.3] [Reference Citation Analysis (5)] |
| 23. | Kohler BA, Ward E, McCarthy BJ, Schymura MJ, Ries LA, Eheman C, Jemal A, Anderson RN, Ajani UA, Edwards BK. Annual report to the nation on the status of cancer, 1975-2007, featuring tumors of the brain and other nervous system. J Natl Cancer Inst. 2011;103:714-736. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 513] [Cited by in RCA: 476] [Article Influence: 31.7] [Reference Citation Analysis (0)] |
| 24. | Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, Reichert M, Beatty GL, Rustgi AK, Vonderheide RH, Leach SD, Stanger BZ. EMT and dissemination precede pancreatic tumor formation. Cell. 2012;148:349-361. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1824] [Cited by in RCA: 1727] [Article Influence: 123.4] [Reference Citation Analysis (4)] |
| 25. | Rhim AD, Oberstein PE, Thomas DH, Mirek ET, Palermo CF, Sastra SA, Dekleva EN, Saunders T, Becerra CP, Tattersall IW, Westphalen CB, Kitajewski J, Fernandez-Barrena MG, Fernandez-Zapico ME, Iacobuzio-Donahue C, Olive KP, Stanger BZ. Stromal elements act to restrain, rather than support, pancreatic ductal adenocarcinoma. Cancer Cell. 2014;25:735-747. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1792] [Cited by in RCA: 1691] [Article Influence: 140.9] [Reference Citation Analysis (5)] |
| 26. | Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016;388:73-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1909] [Cited by in RCA: 1799] [Article Influence: 179.9] [Reference Citation Analysis (3)] |
| 27. | Urbach L, Wieland L, Penz F, Samuel RD, Küffer S, Klein L, Lenz C, Sax U, Ghadimi M, Schulz-Heddergott R, Hessmann E, Ellenrieder V, Dusetti N, Singh SK. TP53 missense-specific transcriptional plasticity drives resistance against cell cycle inhibitors in pancreatic cancer. Sci Adv. 2025;11:eadu2339. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 28. | Kim MP, Li X, Deng J, Zhang Y, Dai B, Allton KL, Hughes TG, Siangco C, Augustine JJ, Kang Y, McDaniel JM, Xiong S, Koay EJ, McAllister F, Bristow CA, Heffernan TP, Maitra A, Liu B, Barton MC, Wasylishen AR, Fleming JB, Lozano G. Oncogenic KRAS Recruits an Expansive Transcriptional Network through Mutant p53 to Drive Pancreatic Cancer Metastasis. Cancer Discov. 2021;11:2094-2111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 131] [Article Influence: 26.2] [Reference Citation Analysis (0)] |
| 29. | Hingorani SR, Wang L, Multani AS, Combs C, Deramaudt TB, Hruban RH, Rustgi AK, Chang S, Tuveson DA. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell. 2005;7:469-483. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2293] [Cited by in RCA: 2055] [Article Influence: 97.9] [Reference Citation Analysis (3)] |
| 30. | Avila JL, Kissil JL. Notch signaling in pancreatic cancer: oncogene or tumor suppressor? Trends Mol Med. 2013;19:320-327. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 103] [Article Influence: 7.9] [Reference Citation Analysis (1)] |
| 31. | Wang YY, Wang ZX, Hu YD, Wang L, Li N, Zhang B, Han W, Jiang JM. Current status of pathway analysis in genome-wide association study. Yi Chuan. 2017;39:707-716. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |