Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.114547
Revised: December 15, 2025
Accepted: January 26, 2026
Published online: April 15, 2026
Processing time: 195 Days and 1.6 Hours
CD161 in hepatocellular carcinoma (HCC) research remains very limited, despite recent evidence that has implicated it as an immune checkpoint and a potential immunotherapeutic target in cancers such as glioma and breast cancer.
To examine the CD161, CD8, and programmed death receptor 1 (PD-1) expression profiles in patients with hepatitis B virus-associated chronic liver disease and HCC and to evaluate the effects of PD-1 inhibitors on the expression of these molecules in the peripheral blood of patients with HCC.
The CD161, CD8, and PD-1 expression levels in the peripheral blood of individuals with chronic hepatitis B (CHB), decompensated cirrhosis, or HCC and in liver cancer tissues obtained from the HCC group were analyzed using bioinformatics, flow cytometry, qPCR, and immunohistochemistry.
The proportion of CD161+CD8+ T cells was significantly elevated in the peripheral blood of patients with HCC. This result suggested a potential correlation between T-cell exhaustion and chronic inflammation. The bioinformatics analysis revealed a co-expression relationship between CD161 and PD-1, and we found that PD-1 inhibitors effectively decreased the proportion of PD-1+ cells in the peripheral blood but did not significantly affect the CD161 expression. The CD161 immunohistochemical staining intensity was significantly greater in tumor tissues than in both adjacent nontumorous liver tissues and liver tissues from patients with CHB.
CD161+CD8+ T cells are PD-1-independent biomarkers of T-cell exhaustion in patients with HCC, as these cells are present at elevated levels in peripheral blood and exhibit increased tissue expression. PD-1 inhibitors effectively reduce PD-1+ cells, but they fail to modulate CD161 expression. This highlights its distinct regulatory mechanism and potential as a novel therapeutic target.
Core Tip: This study reveals elevated CD161+CD8+ T cells in peripheral blood of hepatitis B virus (HBV)-associated hepatocellular carcinoma (HCC) patients, correlating with disease progression. Programmed death receptor 1 (PD-1) inhibitors reduced PD-1+ subsets but did not alter CD161 expression, indicating CD161 operates independently of PD-1 pathways. Critically, CD161 protein overexpression in HCC tissues—exceeding paracancerous and chronic hepatitis B tis
- Citation: Liu CR, Zhang M, Wang MQ, Zhang W, Li J, Shi GZ, Liang FF, Li YP, Huang N. CD161+CD8+ T cells in patients with hepatitis B virus-associated hepatocellular carcinoma. World J Gastrointest Oncol 2026; 18(4): 114547
- URL: https://www.wjgnet.com/1948-5204/full/v18/i4/114547.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i4.114547
Tumor immunotherapy has rapidly advanced in recent years and has attracted extensive attention to the pivotal role of T cells in antitumor immunity. In particular, the use of immune checkpoint inhibitors has markedly improved the prognosis of a variety of solid malignancies. However, the functionality of tumor-infiltrating lymphocytes within different tumor microenvironments (TMEs) can be suppressed by immunosuppressive signals, and this leads to a diminished antitumor immune response[1,2]. Under this context, the CD161 molecule functions as a potential immunosuppressive receptor, and it has emerged as a research focal point. In 2021, Mathewson et al[3] reported in a study published in the Cell a high CD161 expression in T cells from glioma patients using single-cell sequencing techniques. They reported that the blocking of the CD161 expression markedly increased the cytotoxic efficacy of T cells against tumor cells[3]. Similar results were obtained in studies that had used human breast cancer cell lines[4]. CD161 is encoded by the KLRB1 gene. It is broadly expressed across natural killer (NK) cells and T cells and is preferentially distributed on memory T cells[5-7]. Upon binding to CD161, its ligand, the LLT1, can inhibit the cytotoxicity of NK cells and suppress interferon-γ (IFN-γ) secretion[8-11]. This immunomodulatory mechanism parallels the role of the programmed death receptor 1 (PD-1)/programmed death ligand-1 axis, and it positions CD161 as a promising target for immunotherapeutic strategies[12-14].
CD161 expression patterns exhibit variations across diverse malignancies, but research regarding its role in hepatocellular carcinoma (HCC) remains scarce. Sun et al[15] reported that CD161 and IL-7R overexpression on tumor-infiltrating CD8+ T cells in patients with recurrent HCC correlated with reduced cytotoxicity and a decreased proliferative capacity. This result suggested a role for these molecules in tumor-associated T-cell exhaustion[15]. Another study demonstrated that CD8+PD-1+CD161+ T cells exhibited enhanced proliferative potential and cytotoxicity in patients with HCC. In addition, high levels of these cells were significantly correlated with extended recurrence-free survival (RFS)[16]. Further, studies have found that the blocking the LLT1/CD161 interaction may reverse the functional suppression of NK cells and boost their anti-tumor activity. This would restore the anti-tumor cytotoxicity of NK cells in primary tumors while enhancing their control over metastatic foci[14]. However, the findings from these studies varied across different tumor types, and the precise mechanism of action of CD161 in HCC remains unclear. Thus, CD161 function and its associated immune molecules in HCC warrants further investigation, particularly among patients who receive immunotherapy. CD161 has a potential immunosuppressive role in liver cancer; hence, an investigation of its utility as an immunotherapeutic target for HCC treatment is imperative. In our present study, we quantified the CD161, CD8, and PD-1 expressions in the peripheral blood mononuclear cells (PBMCs) from patients with liver cancer using flow cytometry. In addition, immunohistochemical staining was conducted on resected liver cancer tissues and the corresponding paracancerous tissues to determine the CD161, CD8, and PD-1 molecular expression levels. This study was designed to elucidate the expression patterns of CD161 in hepatitis B virus (HBV)-associated liver cancer and to evaluate its impact on the prognosis.
Analysis of the RNA-seq data from The Cancer Genome Atlas: The RNA-seq data and corresponding clinical information of patients with HCC were retrieved from The Cancer Genome Atlas (TCGA)-Liver Hepatocellular Carcinoma (LIHC) dataset. The KLRB1 (CD161), CD8A (CD8), and PDCD1 (PD-1) gene expression levels in tumors and paracancerous tissues were determined using R software, and the differences in expressions between these two groups were compared.
Survival analysis using the Kaplan-Meier plotter: The Kaplan-Meier plotter database was used to examine the associations of the target gene expressions with the overall survival (OS), the progression-free survival (PFS), and the RFS in patients with HCC. All of the hepatitis virus-associated HCC samples were screened for the survival analyses.
Co-expression and functional enrichment analyses: Genes co-expressed with KLRB1 were identified using the cBioPortal database (correlation coefficient r > 0.55). Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were conducted using the DAVID 6.8 tool. The top five functional enrichments were visualized using bubble maps generated using the R package “ggplot2”.
Sample source and collection: Peripheral blood samples were obtained from patients who were diagnosed with chronic hepatitis B (CHB), HBV-associated decompensated cirrhosis (DCC), or HBV-associated HCC and were hospitalized in the Department of Infectious Diseases at the Second Affiliated Hospital of Xi’an Jiaotong University from July 2022 to March 2023. All of the liver cancer tissues and corresponding paracancerous tissues were harvested from surgical inpatients at the Second Affiliated Hospital of Xi’an Jiaotong University within the same timeframe.
Patients aged 18 years or older who fulfilled the diagnostic criteria for CHB, DCC, or HCC were eligible for inclusion. Patients who were pregnant, had hematologic disorders and/or other concurrent viral infections, or were receiving immunosuppressive therapy were excluded from the study. In addition, HCC patients who received PD-1 inhibitor therapy were required to meet the following criteria: (1) Had received PD-1 inhibitors for a minimum of 12 weeks; and (2) The immunohistochemical staining samples were derived from pathologically confirmed HCC. Those who had previously undergone PD-1 inhibitor treatment were excluded.
Reagents and equipment: The peripheral blood lymphocyte separation medium (ApplyGen, Beijing, China), red blood cell lysis buffer (Boster BioTech, Wuhan, China), immunofluorescent antibodies (CD8A, Alexa Fluor; CD279 PE and CD161 APC, BD Biosciences, United States), an RNA rapid extraction kit (Mei5Bio, Beijing, China), a cDNA synthesis kit (RR047A; TaKaRa, Japan), real-time qPCR reagents (RR820A, TaKaRa, Japan), a flow cytometer (BD Biosciences, United States), a cryogenic centrifuge (Eppendorf AG, Germany), an ultraviolet spectrophotometer (NanoDrop One; Thermo, United States), a standard PCR machine (Bio-Rad, United States), an ABI 7500 Fast real-time PCR system (Applied Biosystems, United States), paraffin embedding medium and a microtome (Leica Microsystems, Germany), and an upright microscope (E100, Nikon Corporation, Japan) were used in this study. The qPCR primers were synthesized by Sangon Biotech Co., Ltd. (Shanghai, China), and their sequences are provided in Supplementary Table 1.
Isolation of the PBMCs and flow cytometry: The isolated PBMCs were resuscitated and then subjected to immunofluorescence staining. Flow cytometry was performed to analyze the lymphocyte subsets, specifically the CD8+ T cells and the PD-1+ and CD161+ subsets. In addition, CD8+PD-1+CD161+ T cell distribution was further analyzed.
Real-time-qPCR: Total RNA was extracted from liver cancer and paracancerous tissues and quantified using a NanoDrop One ultraviolet spectrophotometer. Reverse transcription and quantitative fluorescence detection were performed according to the manufacturer’s instructions. The PCR conditions were as follows: Initial denaturation at 95 °C for 30 seconds (1 cycle), followed by denaturation at 95 °C for 5 seconds and annealing at 60 °C for 30 seconds (40 cycles).
Immunohistochemical staining: The tissue sections were dewaxed and then dehydrated through an ethanol gradient. After antigen retrieval using a citrate buffer, the sections were thoroughly rinsed with phosphate-buffered saline (PBS). Antibodies specific to PD-1, CD8A, and CD161 were separately added, and the sections were incubated overnight at 4 °C. Following the addition of the secondary antibodies, the sections were incubated for 30 minutes at room temperature, rinsed again with PBS, and developed using 3,3'-diaminobenzidine (DAB). Hematoxylin was subsequently applied for counterstaining. The immunostaining images were processed using ImageJ software. The positively stained areas were separated in “H-DAB” mode. The images were transformed into grayscale images, with the thresholds adjusted accordingly. Ultimately, the average optical density (OD) values of the target molecules were calculated, and a higher OD value represented a higher expression level of the target molecule.
Statistical analysis was conducted using IBM SPSS® Statistics 25.0 software. The average OD values of the target molecules in cancer tissues and the corresponding paracancerous tissues in the immunohistochemical analysis were paired for comparison. The normally distributed measurement data were expressed as the means ± SD, whereas the nonnormally distributed measurement data were expressed as the medians (M) with the interquartile range (P25-P75). An analysis of variance (ANOVA) was used to compare the normally distributed measurement data across multiple groups, while a Student’s t test was used for comparisons between two groups. A paired t test was used for the paired data. For the nonnormally distributed measurement data, the Kruskal-Wallis test was applied for comparisons across multiple groups, and the Mann-Whitney U test was used for comparisons between two groups. All of the tests were two-tailed, with a P value of less than 0.05 considered to be statistically significant.
This study was approved by the Ethics Committee of the Second Affiliated Hospital of Xi’an Jiaotong University (Approval No. 2022110).
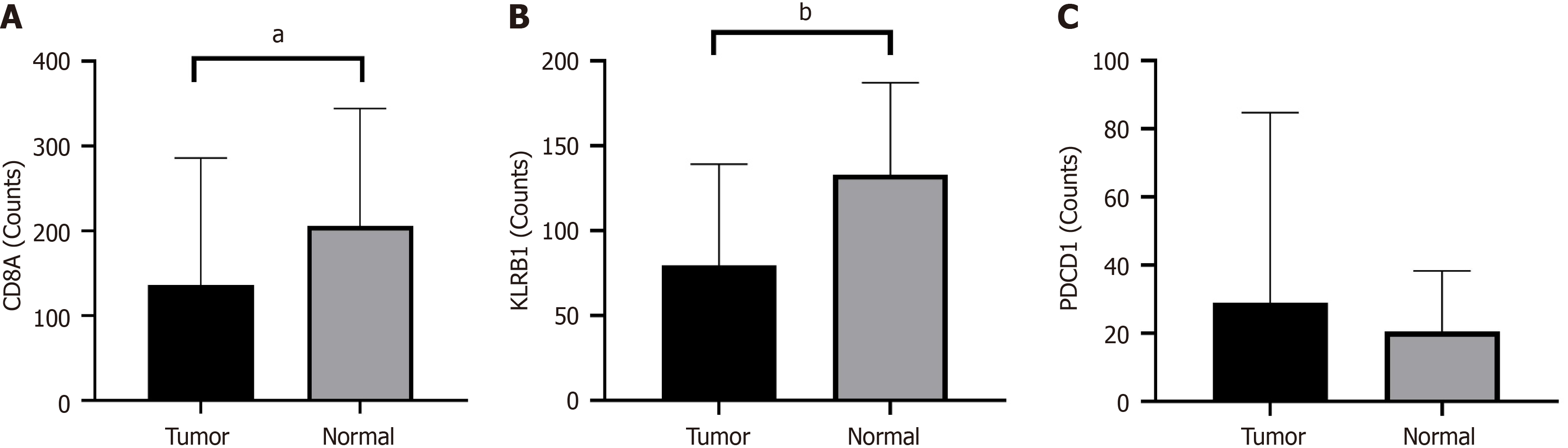
CD8 and CD161 mRNA expression levels were lower in the HCC tissues than in paracancerous tissues: RNA-seq data from the TCGA-LIHC cohort were extracted using R software. The results yielded expression levels of 59427 genes from 374 HCC tissues and 50 paracancerous tissues. The median expression levels of CD8A [118 (53-283) counts vs 206 (112.25-344) counts, P < 0.05; Figure 1A] and KLRB1 [79 (37-139) counts vs 133 (81-187) counts, P < 0.001; Figure 1B] were significantly lower in the HCC tissues than in the paracancerous tissues. PDCD1 expression was higher in the HCC tissues than in paracancerous tissues [30 (12-88) counts vs 21 (12-38) counts, P = 0.102], but the difference was not statistically significant (Figure 1C).
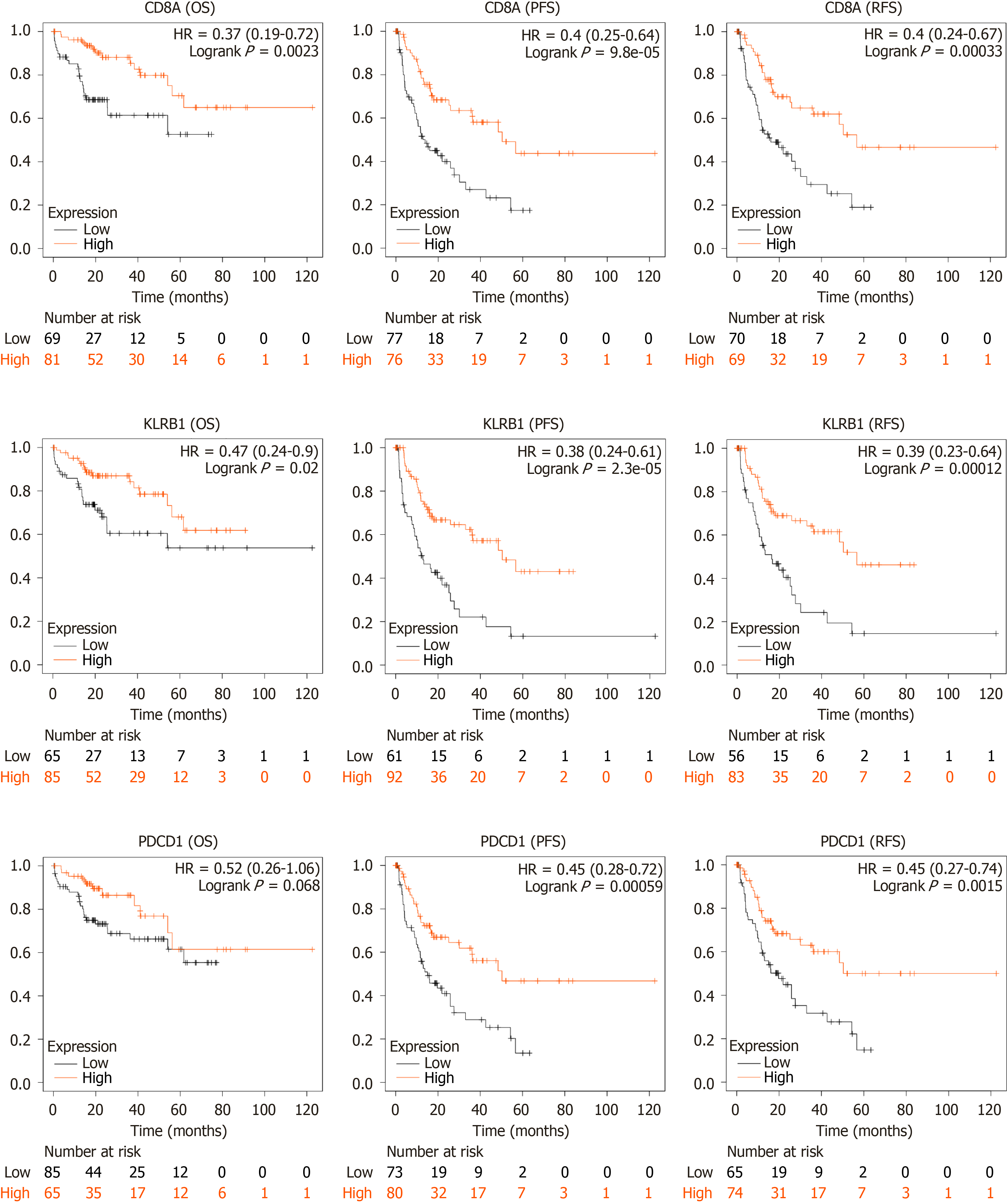
High CD8 and CD161 mRNA expression levels were closely associated with a favorable prognosis in patients with HCC: A total of 153 patients with HBV-associated HCC were identified after conditional screening. A total of 150 patients were used for OS analysis, 138 patients for RFS analysis, and 152 patients for PFS analysis. The results indicated that compared with the low CD8A expression group, the high CD8A expression group had significantly longer OS values (upper quartile: 56.2 months vs 14.2 months; cutoff = 108 counts), PFS values (median: 50.3 months vs 12.87 months; cutoff = 121 counts), and RFS values (median: 56.67 months vs 15.97 months; cutoff = 121 counts). The PDCD1 expression level did not affect OS. The OS values (upper quartile: 54.1 months vs 14.4 months; cutoff = 86 counts), PFS values (median: 54.1 months vs 14.4 months; cutoff = 70 counts), and RFS values (median: 56.67 months vs 16.37 months; cutoff = 70 counts) were significantly longer in the high-KLRB1 expression group than those of the low-KLRB1 expression group. Thus, high mRNA expression levels of CD8 and CD161 were closely correlated with improved prognoses in patients with HBV-associated HCC (Figure 2).
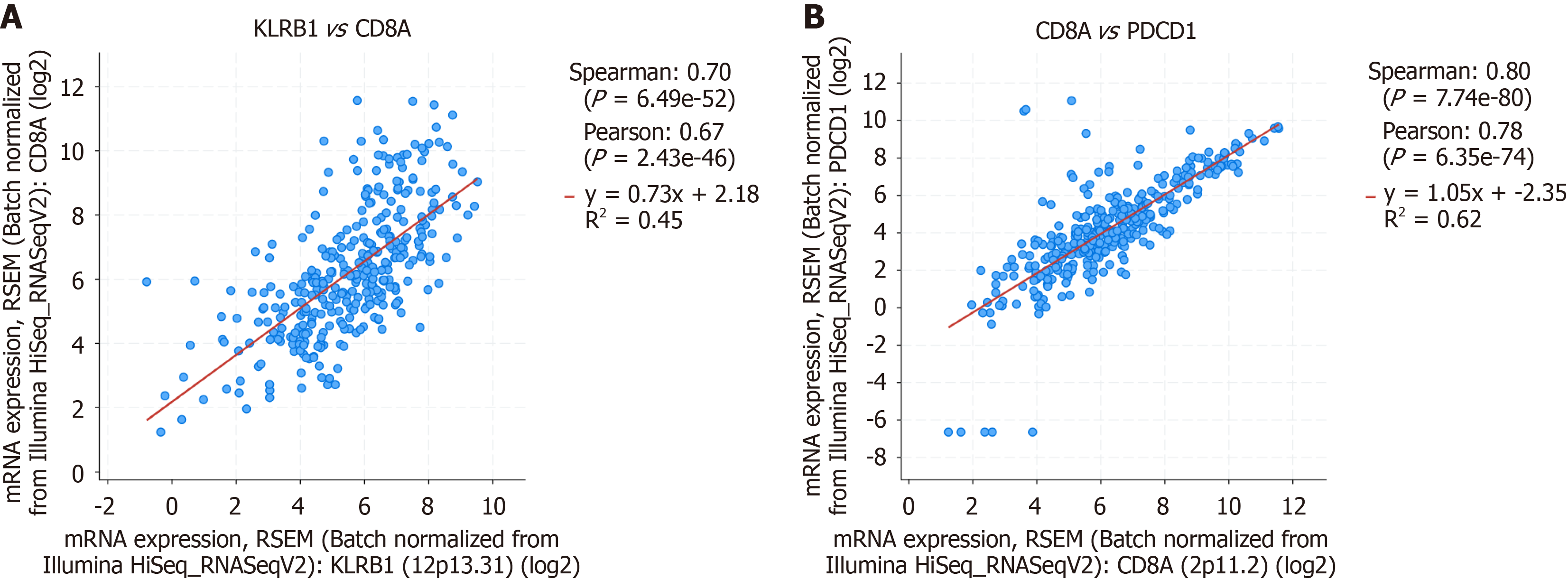
A significant correlation existed between KLRB1 and CD8A gene expression: A total of 400 genes were co-expressed with KLRB1 (r > 0.55, P value < 0.05), as analyzed using the cBioPortal database in the TCGA-LIHC dataset. The five genes with the highest correlation coefficients were CD3E, CD6, CD247, ACAP1, and CD48 (all r > 0.8; Supplementary Table 2). KLRB1 was co-expressed with both CD8A (r = 0.7, P < 0.001) and PDCD1 (r = 0.58, P < 0.001; Figure 3).
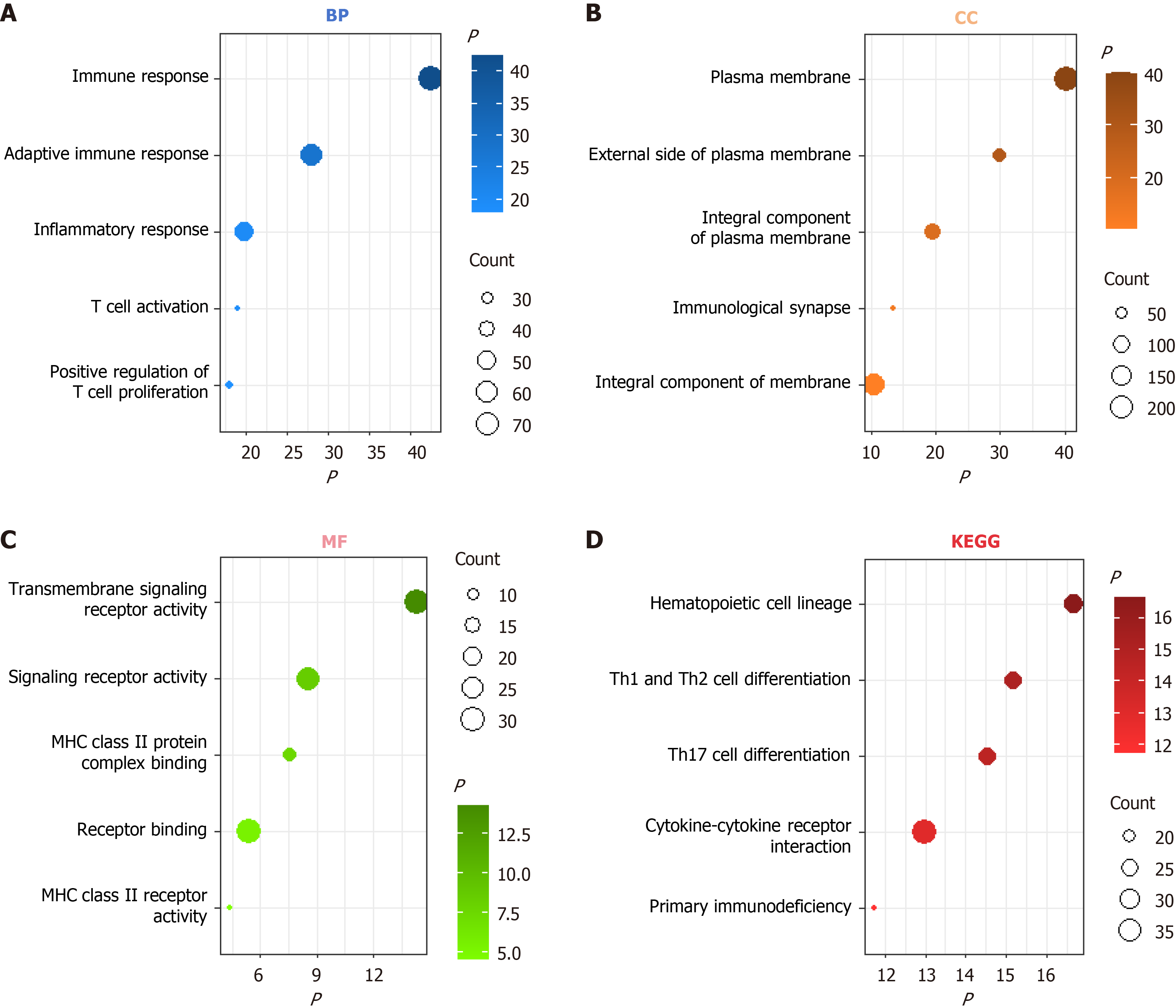
KLRB1 in HCC was related to immunoregulation, as indicated by the functional enrichment analysis: The biological functions of KLRB1 were explored using the DAVID v 6.8. The genes co-expressed with KLRB1 were identified using Spearman correlation analyses within the TCGA-LIHC dataset accessed via the cBioPortal database. GO and KEGG analyses were subsequently conducted based on the identified gene sets. In the TCGA-LIHC dataset, the most relevant biological processes associated with KLRB1 included the immune response, adaptive immune response, inflammatory response, and T-cell activation (Figure 4A). Further, the cell membrane was the most relevant cellular component associated with KLRB1 (Figure 4B). The molecular functions predominantly linked to KLRB1 included transmembrane signaling receptor activity and signaling receptor activity (Figure 4C). The KEGG analysis revealed that the most relevant signaling pathways associated with KLRB1 were those involved in the hematopoietic cell lineage and Th1 and Th2 cell differentiation (Figure 4D). Thus, as a key effector within the T-cell microenvironment, CD161 may play a pivotal role in the immune response and immune regulation in HCC.
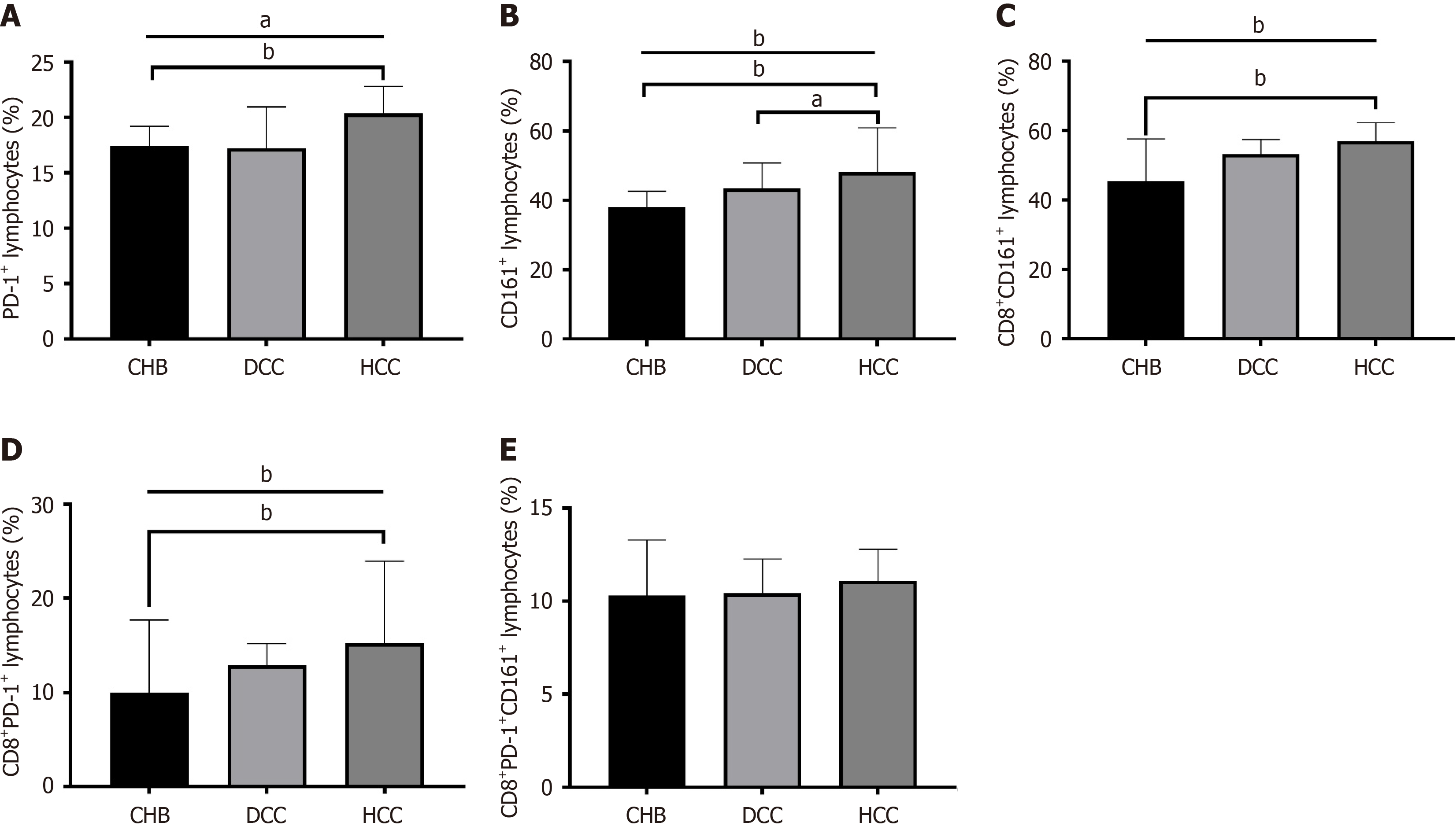
CD161, CD8, and PD-1 expression in patients with CHB, DCC, and HCC displayed the following flow cytometric findings: There were 15 patients with CHB, 18 patients with DCC, and 30 patients with non-PD-1 inhibitor-treated HCC (non-PD-1-HCC) were included in the flow cytometry analysis. The CD161, CD8, and PD-1 expression level comparison revealed that the proportion of PD-1+ lymphocytes in the peripheral blood of patients with CHB was significantly lower than that in the peripheral blood of the patients without PD-1-HCC [17.40% (6.19%-19.21%) vs 20.36% (17.94%-22.81%), P < 0.01]. However, no significant differences were observed between the patients with CHB and the patients with DCC or between the patients with DCC and the patients without PD-1-HCC (Figure 5A). In addition, the proportion of CD161+ lymphocytes in the peripheral blood was significantly lower in the patients with CHB [37.98% (32.06%-42.62%)] and the patients with DCC [43.45% (36.10%-50.78%)] than in the patients without PD-1-HCC [48.15% (44.39%-60.92%)], with no significant difference between the patients with CHB and the patients with DCC (Figure 5B). Further, we compared this result with that in the patients without PD-1-HCC, and the proportion of CD161+CD8+ T cells in the patients with CHB was significantly lower than that in the patients without PD-1-HCC [45.48% (40.57%-57.63%) vs 56.92% (52.00%-62.28%), P < 0.01]. However, the results were not significantly different between the patients with DCC and the patients with CHB or between the patients with DCC and the patients with HCC (Figure 5C). The proportion of CD8+PD-1+ T cells in the peripheral blood was significantly lower than that in the patients with CHB [10.00% (8.68%-17.69%)] and the patients with DCC [12.85% (8.17%-15.19%)] than in the patients without PD-1-HCC [15.23% (12.17%-23.97%)]. In addition, the difference observed between the patients with CHB and the patients with DCC was not statistically significant (Figure 5D). Finally, there was no statistically significant difference in the proportion of CD8+PD-1+CD161+ T cells among these three groups (Figure 5E). Thus, the proportions of PD-1+ lymphocytes, CD161+ lymphocytes, and CD8+PD-1+/CD161+CD8+ T cells in the peripheral blood of patients with CHB and patients with DCC were significantly lower than those in the patients without PD-1-HCC. This result suggested that the proportions of immunosuppressive T cells markedly increase as the disease progressed, and this may contribute to the pathogenesis and immune evasion of liver cancer.
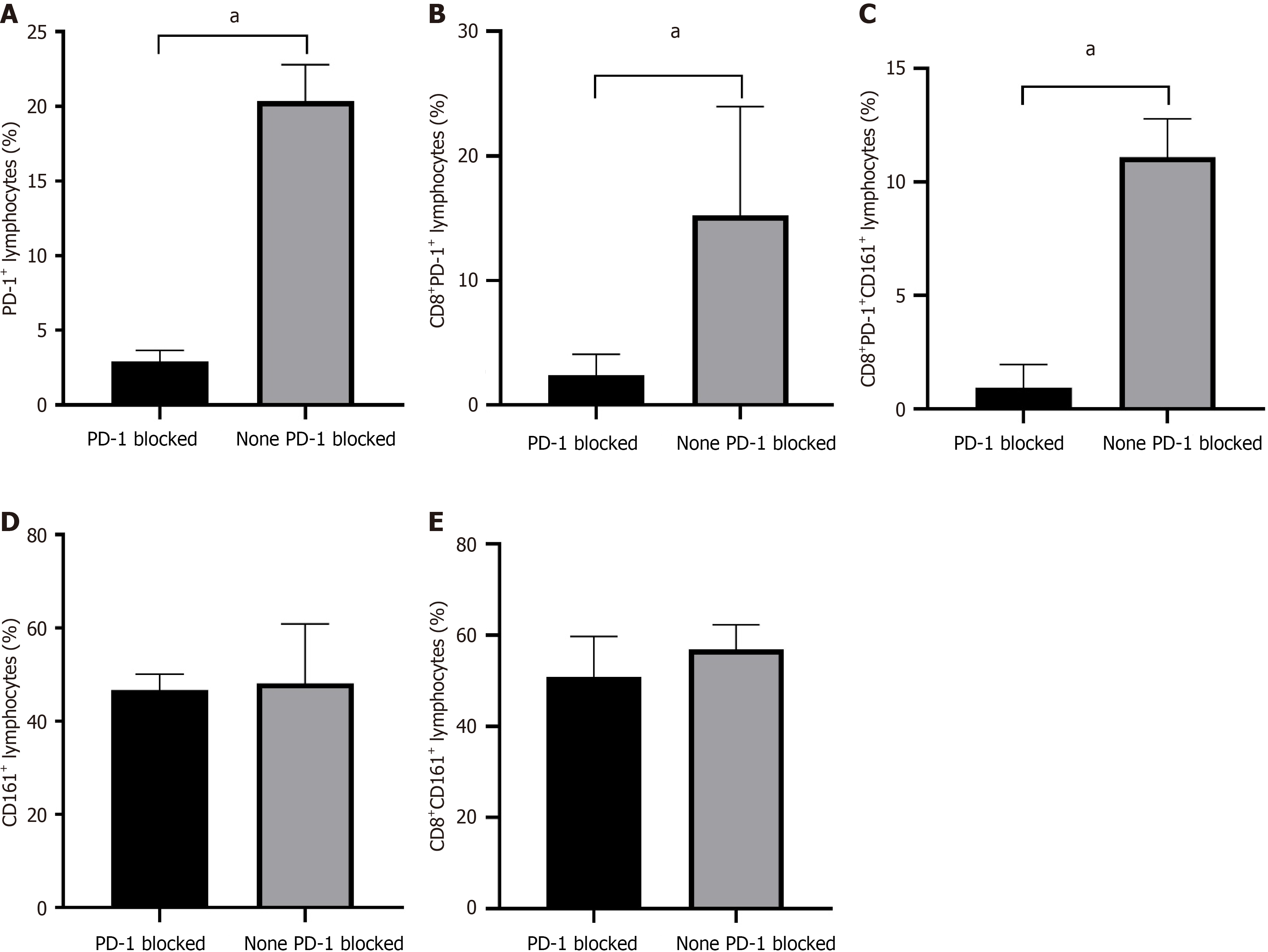
Effects of PD-1 inhibitors on the CD161, CD8, and PD-1 expressions in the peripheral blood: An additional cohort of 19 patients with HBV-associated HCC who had received PD-1 inhibitor therapy was compared with 30 patients with HCC without PD-1 inhibitor exposure to investigate the effects of the PD-1 blockade on the peripheral blood CD8+, PD-1, and CD161 levels. The peripheral blood proportions of PD-1+ lymphocytes [2.93% (1.80%-3.67%) vs 20.36% (17.94%-22.81%), P < 0.001], CD8+PD-1+ T cells [2.40% (1.51%-4.07%) vs 15.23% (12.17%-23.97%), P < 0.001], and CD8+PD-1+CD161+ T cells [0.94% (0.61%-1.96%) vs 11.09% (10.02%-12.79%), P < 0.001] were significantly lower in the PD-1-HCC group than in the non-PD-1-HCC group (Figure 6A-C). No significant difference in the proportions of CD161+ lymphocytes or CD161+CD8+ T cells was detected between these two groups (Figure 6D and E). Thus, PD-1 inhibitors substantially affected the proportions of PD-1-associated cell subsets in the peripheral blood of patients with HCC. The significantly lower proportions of PD-1+ lymphocytes, CD8+ PD-1+ T cells, and CD8+PD-1+CD161+ T cells in patients with HCC treated with PD-1 inhibitors suggested that PD-1 inhibitors predominantly stimulated antitumor immune responses by decreasing the proportion of PD-1+ cells. However, the PD-1 blockade did not significantly influence the CD161 expression in specific T-cell subsets.
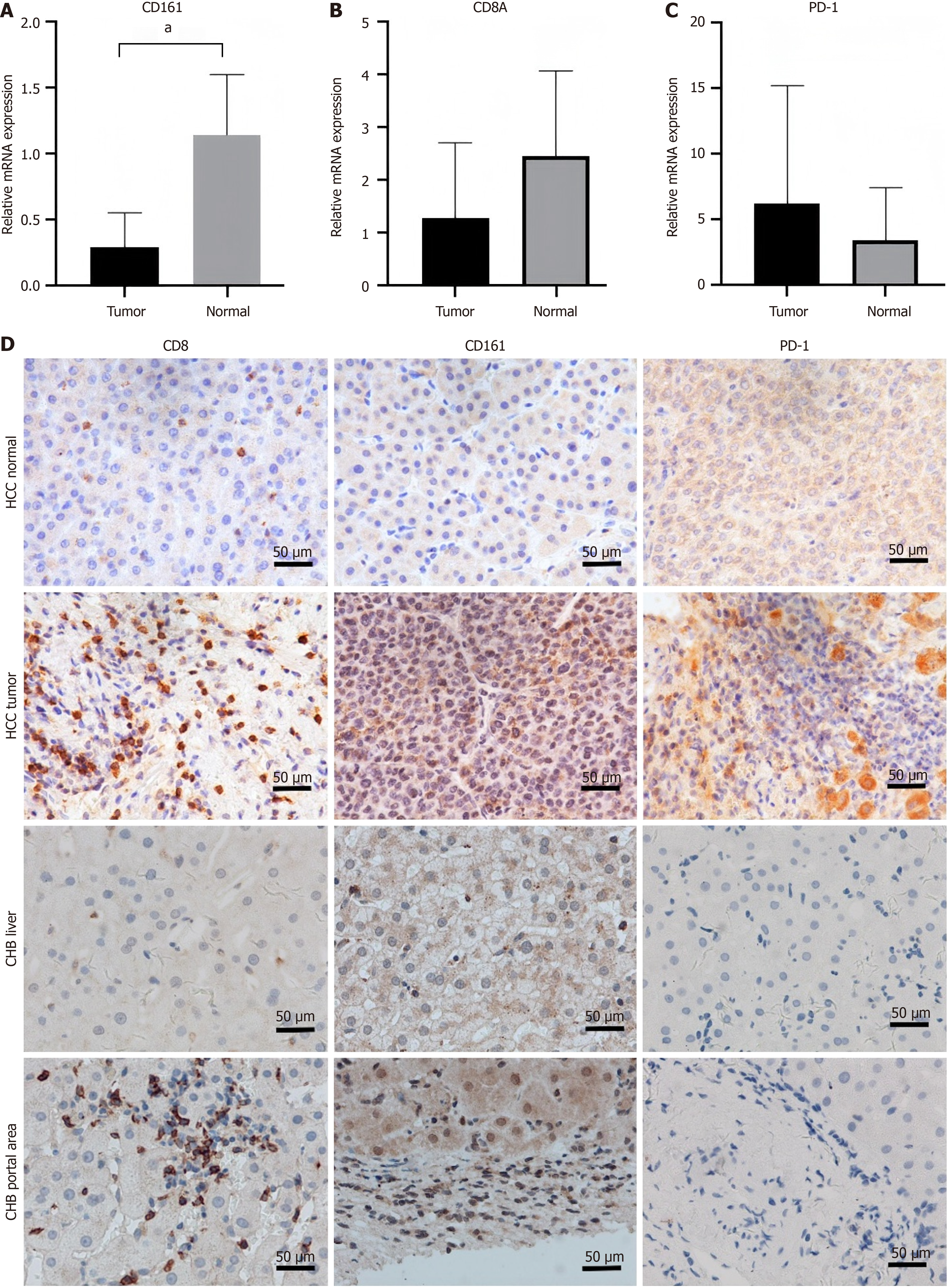
CD8A, CD161, and PDCD1 gene expressions in HCC and paracancerous tissues: The real-time qPCR results revealed that the mRNA level of CD161 in HCC tissues was significantly lower than that in paracancerous tissues (0.29 ± 0.26 vs 1.15 ± 0.45, P < 0.001; Figure 7A). The CD8A mRNA level was also lower in the HCC tissues than in paracancerous tissues (2.47 ± 1.6 vs 1.28 ± 1.24, P < 0.087), although this difference did not reach statistical significance (Figure 7B). Finally, the PD-1 mRNA level was greater in the HCC tissue samples than in the paracancerous tissue samples (7.1 ± 9.3 vs 2.8 ± 3.8, P < 0.44), but this difference was not statistically significant (Figure 7C).
Immunohistochemical staining of the CD161, PD-1, and CD8 expression levels in liver tissues from patients diagnosed with HCC and CHB: HCC and paracancerous tissues were collected from 10 patients with HBV-associated HCC. Liver tissues and portal area tissues from another 10 patients with CHB were collected. A total of 231 CD8-stained sections (76 HCC tissues, 75 paracancerous tissues, 40 CHB liver tissues, and 40 CHB portal area tissues), 240 PD-1-stained sections (80 HCC tissues, 80 paracancerous tissues, 40 CHB liver tissues, and 40 CHB portal area tissues), and 238 CD161-stained sections (80 HCC tissues, 78 paracancerous tissues, 40 CHB liver tissues, and 40 CHB portal area tissues) were analyzed (Figure 7D).
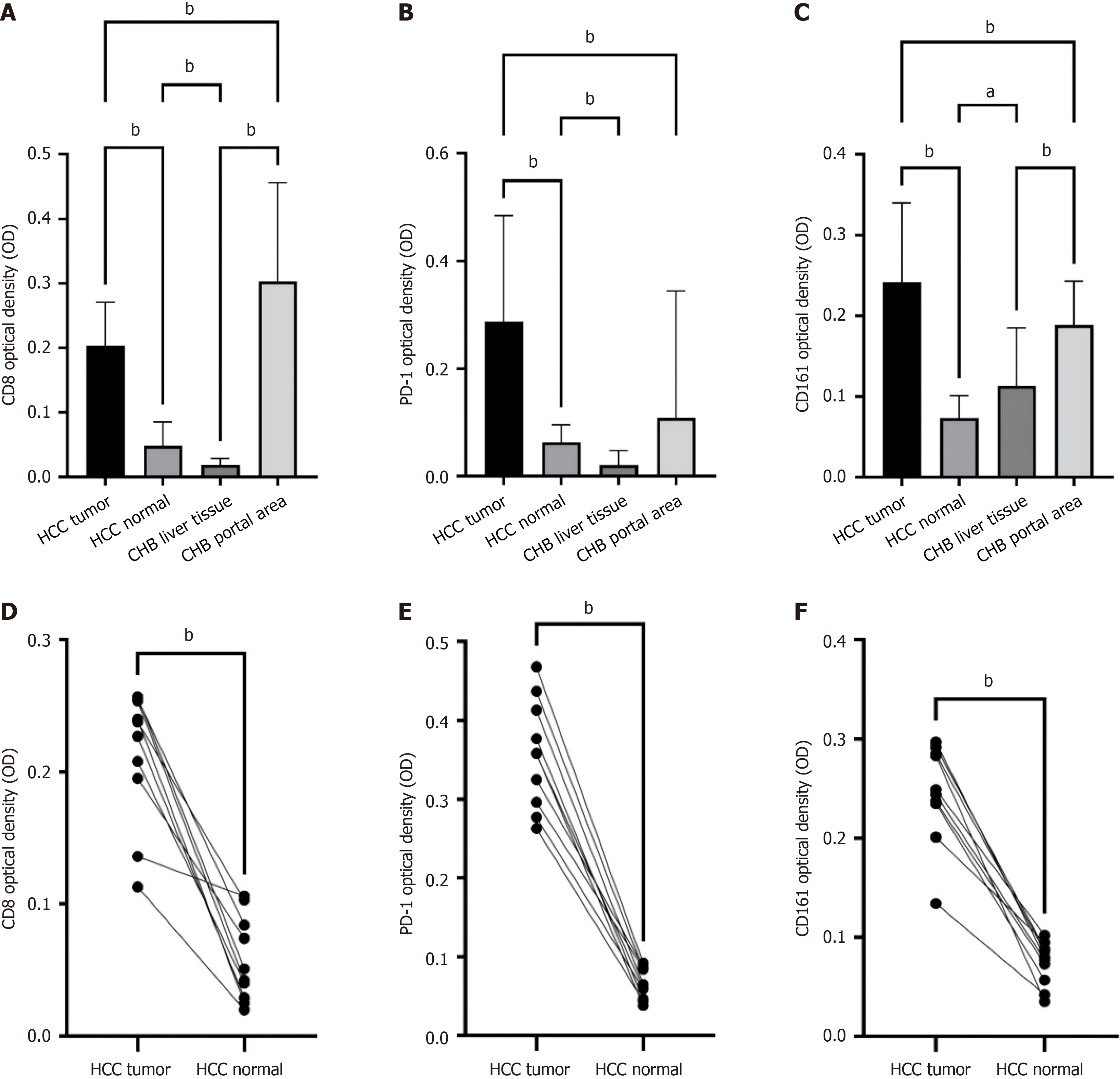
The OD values were measured after the immunohistochemical staining images were converted to grayscale. A semiquantitative method was used to compare the differences in the CD161, CD8, and PD-1 expression levels among the different groups. The expression levels of CD8 (OD = 0.191 ± 0.066 vs 0.049 ± 0.038, P < 0.0001; Figure 8A), PD-1 (OD = 0.410 ± 0.118 vs 0.062 ± 0.033, P < 0.0001; Figure 8B), and CD161 (OD = 0.231 ± 0.111 vs 0.066 ± 0.027, P < 0.0001; Figure 8C) were significantly greater in the HCC tissues than in paracancerous tissues. Thus, the OD values of CD161, CD8, and PD-1 were significantly increased in the HCC tissues, as evidenced by densitometry. An unpaired analysis using the Mann-Whitney U test and t test, as well as paired t tests, revealed that the differences in the expression levels of these three markers (CD8, PD-1, and CD161) remained statistically significant between the HCC tissues and paracancerous tissues. Specifically, the expression levels of CD8 (OD = 0.212 ± 0.051 vs 0.057 ± 0.032, P < 0.001; Figure 8D), PD-1 (OD = 0.357 ± 0.069 vs 0.066 ± 0.020, P < 0.001; Figure 8E), and CD161 (OD = 0.246 ± 0.050 vs 0.074 ± 0.022, P < 0.001; Figure 8F) remained significantly greater in the HBV-associated HCC tissues. Thus, the infiltration of CD8+ T cells, the immune checkpoint role of PD-1, and the immunomodulatory function of CD161 may collectively contribute to the immune microenvironment of HBV-associated HCC.
In addition, we compared the immunohistochemical staining levels of CD161, CD8, and PD-1 between liver tissues and portal areas in patients with CHB. The results demonstrated that the CD8 (OD = 0.303 ± 0.153 vs 0.019 ± 0.010, P < 0.0001; Figure 8A) and CD161 (OD = 0.188 ± 0.054 vs 0.113 ± 0.071, P < 0.0001; Figure 8B) levels in the portal tract areas were significantly greater than those in the liver tissues of the patients with CHB, whereas no significant difference was observed in the PD-1 levels between the two groups (OD = 0.108 ± 0.235 OD vs 0.021 ± 0.026, P = 0.138; Figure 8C). To further evaluate the differences in the expression of these molecules between patients with HCC and CHB, we compared the CD161, CD8, and PD-1 levels in liver tissues from patients with CHB with those in adjacent nontumor liver tissues from patients with HCC. Our analysis revealed that the CD8 (OD = 0.049 ± 0.038 vs 0.019 ± 0.010, P < 0.001; Figure 8A) and PD-1 (OD = 0.062 ± 0.033 vs 0.021 ± 0.026, P < 0.0001; Figure 8B) expression levels were significantly elevated in the nontumor liver tissues of patients with HCC compared with those in CHB liver tissues, whereas the CD161 level (OD = 0.066 ± 0.027 vs 0.113 ± 0.072, P < 0.05; Figure 8C) was markedly lower in the nontumor HCC tissues than in the CHB liver tissues. These findings indicated that the infiltration of CD8- and PD-1-positive cells was greater in nontumor liver tissues from patients with HCC than in those from patients with CHB; however, CD161-positive cell infiltration was significantly reduced.
In the present study, the CD8, PD-1, and CD161 expression profiles in patients with CHB, DCC, and HCC were comprehensively analyzed using flow cytometry and immunohistochemical staining. Further, the effects of PD-1 inhibitors on the expression of these molecules in the peripheral blood of patients with HCC were assessed. CD8, PD-1, and CD161 expression varied significantly among patients with HBV-associated chronic liver disease at different pathological stages, and there were also significant differences between liver cancer tissues and paracancerous tissues in patients with HCC.
The numbers of CD161+ and PD-1+ lymphocytes among the PBMCs of patients with HCC were significantly greater than those among the PBMCs of CHB. In addition, the proportions of CD161+CD8+ T cells, CD8+PD-1+ T cells were significantly greater. This observation may indicate T-cell exhaustion, and this a response to chronic antigen stimulation during HBV infection. Evidence suggests that CD161 is closely associated with T-cell functional loss, and its upregulation may contribute to T cell dysfunction under the context of chronic viral infections[17]. Further, Liu et al[18] reported that CHB, CD161+CD8+ T cells were not only were enriched in peripheral blood but also markedly infiltrated the liver. This result suggested that CD161 may participate in the inhibitory regulation of T cells during chronic HBV infection. This could potentially accelerate disease progression[18]. However, this T-cell subset increase may be associated with inflammation and pathological fibrosis. CD161+ T cells can induce hepatic stellate cell activation through the IFN-γ/IL-23/IL-17 pathway, thus exacerbating liver fibrosis[19-21]. In agreement with the findings of Sun et al[15], the enhanced infiltration of CD161+CD8+ T cells in patients with HCC was accompanied by functional impairments. In addition, functional defects in CD161+ T cells may be directly related to a poor prognosis. Glioma research has also revealed similar trends, where the upregulated expression of CD161 was found to suppress the cytotoxic activity of tumor-infiltrating CD8+ T cells. This blocks CD161 expression and could significantly bolster the antitumor function of CD8+ T cells[3]. Therefore, CD161 may play a pivotal role in the immune evasion mechanisms of HBV-associated liver diseases. Future therapeutic strategies, particularly those aimed at reversing T-cell exhaustion, may benefit from targeting CD161 to reinvigorate T-cell function.
In the present study, patients with HCC had reduced CD8+ T cells in their peripheral blood and increased CD8+ T-cell infiltration in tumor tissues. These results correlated with elevated PD-1 and CD161 expressions, suggesting that patients with HCC may exhibit CD8+ T-cell exhaustion in peripheral blood and T-cell dysfunction within the TME. Our results indicated that patients with HCC might have CD8+ T-cell exhaustion in peripheral tissue as well as CD8+ T-cell infiltration in cancer tissues, and the elevated expressions of PD-1 and CD161 in cancer tissues might suggest that these infiltrating CD8+ T cells are likely to be in a state of functional exhaustion. In a previous study, the sequencing of HCC-infiltrating CD8+ T cells revealed that these cells were predominantly exhausted and characterized by high expressions of immunosuppressive molecules, such as PD-1 and TIM-3[22]. Immunosuppressive molecules within the TME not only directly affect the function of T cells but also further weaken antitumor immunity through the recruitment of more immunosuppressive cells, such as regulatory T cells (Tregs) or myeloid-derived suppressor cells[23]. In addition, metastases within the liver can recruit peripheral CD8+ T cells into the liver, and this would lead to apoptosis and functional exhaustion[24,25]. These processes may explain the CD8+ T cell reductions in peripheral blood and the increased infiltration of CD8+ T cells in cancer tissues in patients with liver cancer.
Bioinformatics analysis revealed that CD161 and CD8 mRNA were under expressed in cancer tissues, and this was inversely correlated with the results obtained by the flow cytometry and immunohistochemistry (IHC). In fact, immunohistochemical staining was used to assess protein expression, and TCGA-LIHC data were used to examine mRNA levels. The results suggested that complex posttranscriptional and epigenetic regulation occurred between mRNA and protein translation, during which the proteins are the primary molecules that exert biological effects. It was therefore assumed that CD161, CD8, and PD-1 assessments should focus on protein expression levels. Further, if the regulatory hub of the pathway resides at the mRNA level, mRNA may exert a positive regulatory effect on the protein product; i.e., a higher mRNA level leads to increased protein translation. Conversely, if the regulatory hub lies at the protein level, the protein may exhibit a negative feedback effect on the mRNA, with higher protein levels correlated with lower mRNA levels. As shown in our study, CD8 and CD161 may negatively regulate mRNA expression at the protein level, and this may explain the more favorable prognosis in individuals with high KLRB1 mRNA expressions according to the results of the Kaplan-Meier survival analysis. Further, to validate the accuracy of the IHC staining results in this study, we referred to the Human Protein Atlas (HPA) database (https://www.proteinatlas.org), which documents CD8A and KLRB1 expression in HCC. The database includes IHC staining data from 12 patients with HCC among which 6 cases demonstrated moderate to high intensities of CD8A and KLRB1 staining in tumor tissues. In addition, the HPA data revealed a notable discrepancy between mRNA expression (RNA expression and nTPM) and protein expression (protein expression score) of the KLRB1 gene in both liver and HCC tissues, with mRNA levels being substantially lower than protein levels[26]. This finding further suggested significant heterogeneity in KLRB1 expression in HCC and indicated the presence of robust post-transcriptional regulatory mechanisms. However, the specific regulatory mechanisms warrant further exploration.
The results of this study revealed that the proportions of PD-1+, CD8+PD-1+, and CD8+PD-1+CD161+ T cells in the peripheral blood of patients with HCC treated with PD-1 inhibitors were significantly lower than those in patients with HCC who did not receive PD-1 inhibitors. In contrast, the proportions of CD161+ and CD161+CD8+ T cells did not significantly differ between these two groups. Thus, PD-1 inhibitors did not significantly affect the peripheral CD161 expression but did influence the PD-1 expression. In other words, alterations in PD-1 levels did not significantly affect the peripheral CD161 expression. This result suggested that the regulatory mechanisms of PD-1 and CD161 were independent. However, one study did indicate that the CD161 receptor signaling pathway might be involved in the regulation of PD-1 expression and that the inhibition of CD161 expression could lead to the upregulation of PD-1 expression in tumor-infiltrating T cells[3]. Therefore, CD161 may have a regulatory effect on PD-1, whereas PD-1 may have a weak reciprocal regulatory effect on CD161. The increased expression of CD161 observed in patients with chronic HBV infection and HCC may be associated with T-cell exhaustion and immunosuppression. Studies have shown that CD8+CD161+ T lymphocytes exhibit paradoxical characteristics that maintain substantial amounts of intracellular stores of cytotoxic mediators (granzyme B and perforin) while demonstrating functional impairment in target cell elimination and IFN-γ production[6,7]. Current studies have demonstrated heterogeneous CD161 expression patterns across malignancies, and its functional role exhibits tumor-type specificity. While high CD161 expression is associated with improved prognosis in patients with HCC[27], conflicting evidence has suggested opposing correlations in certain other cancer types. The dual regulatory role of CD161 in HBV infection and HCC pathogenesis needs to be fully elucidated. Therefore, a systematic investigation into its context-dependent immunomodulatory functions is required. Future studies should also investigate the interplay between CD161 and the PD-1 pathway to elucidate the specific mechanisms that contribute to immune evasion.
In this study, CD161 expression in patients with HBV-associated liver disease was explored using bioinformatics, flow cytometry, IHC, and PCR techniques. Both bioinformatics analysis and PCR results revealed lower CD161 expressions in HCC tissues than in adjacent nontumorous tissues. However, flow cytometry and IHC demonstrated higher CD161 protein expressions in HCC tissues than in both adjacent nontumorous tissues and in patients with chronic hepatitis. This discrepancy between the gene and protein expression levels suggested that CD161 expression is regulated by mechanisms other than transcription. CD161 may serve as a crucial immunosuppressive molecule and a potential therapeutic target for immunotherapy in HBV-associated HCC.
We thank all patients who agreed to participate in this study and all staff involved.
| 1. | Zhang Y, Zhang Z. The history and advances in cancer immunotherapy: understanding the characteristics of tumor-infiltrating immune cells and their therapeutic implications. Cell Mol Immunol. 2020;17:807-821. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2469] [Cited by in RCA: 2118] [Article Influence: 353.0] [Reference Citation Analysis (1)] |
| 2. | Cerella C, Dicato M, Diederich M. Enhancing personalized immune checkpoint therapy by immune archetyping and pharmacological targeting. Pharmacol Res. 2023;196:106914. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 15] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 3. | Mathewson ND, Ashenberg O, Tirosh I, Gritsch S, Perez EM, Marx S, Jerby-Arnon L, Chanoch-Myers R, Hara T, Richman AR, Ito Y, Pyrdol J, Friedrich M, Schumann K, Poitras MJ, Gokhale PC, Gonzalez Castro LN, Shore ME, Hebert CM, Shaw B, Cahill HL, Drummond M, Zhang W, Olawoyin O, Wakimoto H, Rozenblatt-Rosen O, Brastianos PK, Liu XS, Jones PS, Cahill DP, Frosch MP, Louis DN, Freeman GJ, Ligon KL, Marson A, Chiocca EA, Reardon DA, Regev A, Suvà ML, Wucherpfennig KW. Inhibitory CD161 receptor identified in glioma-infiltrating T cells by single-cell analysis. Cell. 2021;184:1281-1298. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 375] [Cited by in RCA: 331] [Article Influence: 66.2] [Reference Citation Analysis (0)] |
| 4. | Marrufo AM, Mathew SO, Chaudhary P, Malaer JD, Vishwanatha JK, Mathew PA. Blocking LLT1 (CLEC2D, OCIL)-NKRP1A (CD161) interaction enhances natural killer cell-mediated lysis of triple-negative breast cancer cells. Am J Cancer Res. 2018;8:1050-1063. [PubMed] |
| 5. | Wyrożemski Ł, Qiao SW. Immunobiology and conflicting roles of the human CD161 receptor in T cells. Scand J Immunol. 2021;94:e13090. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 22] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 6. | Takahashi T, Dejbakhsh-Jones S, Strober S. Expression of CD161 (NKR-P1A) defines subsets of human CD4 and CD8 T cells with different functional activities. J Immunol. 2006;176:211-216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 126] [Cited by in RCA: 119] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 7. | Konduri V, Oyewole-Said D, Vazquez-Perez J, Weldon SA, Halpert MM, Levitt JM, Decker WK. CD8(+)CD161(+) T-Cells: Cytotoxic Memory Cells With High Therapeutic Potential. Front Immunol. 2020;11:613204. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 81] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 8. | Oxendine S, O'Connor KC. Brain tumor T cells inhibited by their natural KLR(B1) instinct. Sci Immunol. 2021;6:eabi7302. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 9. | Braud VM, Meghraoui-Kheddar A, Elaldi R, Petti L, Germain C, Anjuère F. LLT1-CD161 Interaction in Cancer: Promises and Challenges. Front Immunol. 2022;13:847576. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 37] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 10. | Buckle I, Guillerey C. Inhibitory Receptors and Immune Checkpoints Regulating Natural Killer Cell Responses to Cancer. Cancers (Basel). 2021;13:4263. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 64] [Cited by in RCA: 56] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 11. | Pesenacker AM, Bending D, Ursu S, Wu Q, Nistala K, Wedderburn LR. CD161 defines the subset of FoxP3+ T cells capable of producing proinflammatory cytokines. Blood. 2013;121:2647-2658. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 142] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 12. | Hu X, Dong Y, Xie S, Song Y, Yu C, He Y, Wang Z, Hu Q, Ni Y, Ding L. Immune checkpoint CD161/LLT1-associated immunological landscape and diagnostic value in oral squamous cell carcinoma. J Pathol Clin Res. 2024;10:e353. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 8] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 13. | Anderson AC, Joller N, Kuchroo VK. Lag-3, Tim-3, and TIGIT: Co-inhibitory Receptors with Specialized Functions in Immune Regulation. Immunity. 2016;44:989-1004. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1859] [Cited by in RCA: 1695] [Article Influence: 169.5] [Reference Citation Analysis (1)] |
| 14. | López-Soto A, Gonzalez S, Smyth MJ, Galluzzi L. Control of Metastasis by NK Cells. Cancer Cell. 2017;32:135-154. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 670] [Cited by in RCA: 585] [Article Influence: 65.0] [Reference Citation Analysis (1)] |
| 15. | Sun Y, Wu L, Zhong Y, Zhou K, Hou Y, Wang Z, Zhang Z, Xie J, Wang C, Chen D, Huang Y, Wei X, Shi Y, Zhao Z, Li Y, Guo Z, Yu Q, Xu L, Volpe G, Qiu S, Zhou J, Ward C, Sun H, Yin Y, Xu X, Wang X, Esteban MA, Yang H, Wang J, Dean M, Zhang Y, Liu S, Yang X, Fan J. Single-cell landscape of the ecosystem in early-relapse hepatocellular carcinoma. Cell. 2021;184:404-421. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 716] [Cited by in RCA: 622] [Article Influence: 124.4] [Reference Citation Analysis (1)] |
| 16. | Li Z, Zheng B, Qiu X, Wu R, Wu T, Yang S, Zhu Y, Wu X, Wang S, Gu Z, Shen S, Wu M, Wang H, Chen L. The identification and functional analysis of CD8+PD-1+CD161+ T cells in hepatocellular carcinoma. NPJ Precis Oncol. 2020;4:28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 17. | Diniz MO, Schurich A, Chinnakannan SK, Duriez M, Stegmann KA, Davies J, Kucykowicz S, Suveizdyte K, Amin OE, Alcock F, Cargill T, Barnes E, Maini MK. NK cells limit therapeutic vaccine-induced CD8(+)T cell immunity in a PD-L1-dependent manner. Sci Transl Med. 2022;14:eabi4670. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 33] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 18. | Liu Y, Wang W, Zhu P, Cheng X, Wu M, Zhang H, Chen Y, Chen Y, Liang Z, Wu X, Weng X. Increased Non-MAIT CD161(+)CD8(+) T Cells Display Pathogenic Potential in Chronic HBV Infection. Cell Mol Gastroenterol Hepatol. 2023;15:1181-1198. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 9] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 19. | Li J, Cheng L, Jia H, Liu C, Wang S, Liu Y, Shen Y, Wu S, Meng F, Zheng B, Yang C, Jiang W. IFN-γ facilitates liver fibrogenesis by CD161(+)CD4(+) T cells through a regenerative IL-23/IL-17 axis in chronic hepatitis B virus infection. Clin Transl Immunology. 2021;10:e1353. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 20. | Abdelnabi MN, Hassan GS, Shoukry NH. Role of the type 3 cytokines IL-17 and IL-22 in modulating metabolic dysfunction-associated steatotic liver disease. Front Immunol. 2024;15:1437046. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 21. | Weiskirchen R, Tacke F. Cellular and molecular functions of hepatic stellate cells in inflammatory responses and liver immunology. Hepatobiliary Surg Nutr. 2014;3:344-363. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 124] [Reference Citation Analysis (2)] |
| 22. | Zheng C, Zheng L, Yoo JK, Guo H, Zhang Y, Guo X, Kang B, Hu R, Huang JY, Zhang Q, Liu Z, Dong M, Hu X, Ouyang W, Peng J, Zhang Z. Landscape of Infiltrating T Cells in Liver Cancer Revealed by Single-Cell Sequencing. Cell. 2017;169:1342-1356.e16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1867] [Cited by in RCA: 1615] [Article Influence: 179.4] [Reference Citation Analysis (2)] |
| 23. | Kondoh N, Mizuno-Kamiya M. The Role of Immune Modulatory Cytokines in the Tumor Microenvironments of Head and Neck Squamous Cell Carcinomas. Cancers (Basel). 2022;14:2884. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 38] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 24. | Yu J, Green MD, Li S, Sun Y, Journey SN, Choi JE, Rizvi SM, Qin A, Waninger JJ, Lang X, Chopra Z, El Naqa I, Zhou J, Bian Y, Jiang L, Tezel A, Skvarce J, Achar RK, Sitto M, Rosen BS, Su F, Narayanan SP, Cao X, Wei S, Szeliga W, Vatan L, Mayo C, Morgan MA, Schonewolf CA, Cuneo K, Kryczek I, Ma VT, Lao CD, Lawrence TS, Ramnath N, Wen F, Chinnaiyan AM, Cieslik M, Alva A, Zou W. Liver metastasis restrains immunotherapy efficacy via macrophage-mediated T cell elimination. Nat Med. 2021;27:152-164. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 870] [Cited by in RCA: 799] [Article Influence: 159.8] [Reference Citation Analysis (1)] |
| 25. | Li F, Tian Z. The liver works as a school to educate regulatory immune cells. Cell Mol Immunol. 2013;10:292-302. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 80] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 26. | The Human Protein Atlas. KLRB1. [cited 25 September 2025]. Available from: https://www.proteinatlas.org/ENSG00000111796-KLRB1/tissue. |
| 27. | Billerbeck E, Kang YH, Walker L, Lockstone H, Grafmueller S, Fleming V, Flint J, Willberg CB, Bengsch B, Seigel B, Ramamurthy N, Zitzmann N, Barnes EJ, Thevanayagam J, Bhagwanani A, Leslie A, Oo YH, Kollnberger S, Bowness P, Drognitz O, Adams DH, Blum HE, Thimme R, Klenerman P. Analysis of CD161 expression on human CD8+ T cells defines a distinct functional subset with tissue-homing properties. Proc Natl Acad Sci U S A. 2010;107:3006-3011. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 326] [Cited by in RCA: 301] [Article Influence: 18.8] [Reference Citation Analysis (0)] |