Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.114220
Revised: November 23, 2025
Accepted: January 26, 2026
Published online: April 15, 2026
Processing time: 206 Days and 16.2 Hours
Colorectal cancer (CRC) remains a prevalent malignancy with a high incidence globally. Conventional chemotherapy is limited by substantial toxicity, and the development of novel drugs faces considerable challenges. In this context, drug repurposing has emerged as a promising strategy for CRC treatment.
To explore the therapeutic potential of drug repurposing by evaluating the com
Integrated analysis of data from the SOLAD, CSSL, and SynLethDB databases was performed to identify synthetic lethal interactions involving PIK3CD, leading to the identification of HMGCR as a PIK3CD-interacting synthetic lethal partner. The anti-proliferative effects of atorvastatin (an HMGCR inhibitor) and gefitinib (an epidermal growth factor receptor-PIK3CD pathway inhibitor) were evaluated alone or in combination in SW480 and HCT116 CRC cells using the Cell Counting Kit-8 assay. Cell invasiveness was assessed by Transwell assay, and the expres
Database screening revealed a synthetic lethal relationship between HMGCR and PIK3CD. Experimentally, the combination of atorvastatin and gefitinib exerted a synergistic inhibitory effect on the proliferation and invasion of CRC cells. Mechanistically, this synthetic lethality effect was mediated through the AMPK-SREBP-1 signaling pathway. In vivo, the combined treatment significantly suppressed tumor growth in a mouse xenograft model.
The combination of atorvastatin and gefitinib induces synthetic lethality in CRC via the AMPK-SREBP1 pathway, effectively inhibiting tumor proliferation and metastasis. This drug repurposing strategy presents a novel and translatable approach for CRC therapy.
Core Tip: Integrated analysis of data from the SOLAD, CSSL, and SynLethDB databases was performed to identify HMGCR as a PIK3CD-interacting synthetic lethal partner. Atorvastatin in combination with gefitinib synergistically inhibited the proliferation and metastasis of colorectal cancer (CRC) cells through a synthetic lethal effect via the AMP-activated protein kinase-sterol regulatory element-binding protein 1 signaling pathway. This study provides new research ideas for CRC treatment.
- Citation: Huang JH, Ma JQ. HMGCR loss is synthetic lethal with PIK3CD inhibition in colorectal cancer cells. World J Gastrointest Oncol 2026; 18(4): 114220
- URL: https://www.wjgnet.com/1948-5204/full/v18/i4/114220.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i4.114220
Colorectal cancer (CRC), the third most prevalent malignancy globally, represents a major public health challenge. In recent years, its incidence has continued to rise annually[1,2]. China accounts for the highest number of new CRC cases and related deaths worldwide[3]. Studies have indicated that approximately 20% of CRC patients present with metastases at diagnosis, and the 5-year survival rate for metastatic CRC remains as low as 14%[4,5]. Although the comprehensive application of surgical resection, radiotherapy, chemotherapy, targeted therapy, and other treatment modalities has improved survival outcomes to some extent, therapeutic efficacy remains unsatisfactory for a subset of patients. Hence, there is an urgent need to develop safer and more effective precision treatment strategies.
Lipid metabolic reprogramming is closely associated with tumor growth, proliferation, and invasion during CRC progression[6,7]. Cholesterol metabolism, a key component of lipid metabolism, is critical for maintaining membrane fluidity and stability, thereby playing an important role in tumorigenesis and progression. Statins, a class of lipid-lowering agents widely used in cardiovascular disease, reduce cholesterol synthesis by inhibiting HMGCR and have attracted increasing attention due to their potential anticancer properties[8,9]. Several studies have demonstrated that statins can inhibit tumor growth by inducing cell cycle arrest, suppressing tumor angiogenesis and differentiation[10-13]. Therefore, the potential anticancer properties of statins are increasingly recognized.
Synthetic lethality has emerged as a promising anticancer strategy[14]. It refers to a genetic interaction wherein disruption of either of two genes alone is nonlethal, but simultaneous disruption of both results in cell death[15]. Drug development based on synthetic lethality has the potential to overcome limitations of conventional targeted therapies, such as systemic toxicity, drug resistance, and narrow target applicability, thereby providing a safer and more effective precision medicine approach[16]. For instance, PRMT5 inhibitors selectively eliminate MTAP-deficient tumor cells[17], while WRN inhibitors show promise for treating solid tumors with high microsatellite instability or mismatch repair deficiency[18]. Further elucidation of synthetic lethal interactions in cancer is expected to yield more effective clinical therapeutic strategies.
In this study, we analyzed synthetic lethal databases and identified a synthetic lethal interaction between HMGCR and PIK3CD. Based on this interaction, we subsequently treated CRC cells with a combination of atorvastatin (an HMGCR inhibitor) and gefitinib (an EGFR-PIK3CD signaling pathway inhibitor). Our results demonstrate that the combination of gefitinib and atorvastatin exerts synergistic antitumor effects, suppressing cell proliferation and metastasis while en
The human CRC cell lines SW480 and HCT116 were procured from Wuhan Procell Biotechnology Co., Ltd. (Hubei Province, China). Cells were maintained in Dulbecco's Modified Eagle's Medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1% penicillin, and 1% streptomycin. All cell cultures were incubated at 37 °C in a humidified atmosphere of 5% CO2. The culture medium was refreshed 24 hours after seeding. Cells in the logarithmic growth phase were used for all experiments.
SW480 and HCT116 cells were seeded into 96-well plates at a density of 2000 cells/well. After 24 hours, the cells were treated with a range of concentrations of gefitinib (0 μM, 2.5 μM, 5 μM, 10 μM, and 15 μM) or atorvastatin (0 μM, 2 μM,
Cells were evenly seeded into 6-well plates at a density of 4000 cells/well and allowed to adhere overnight. Subsequently, the culture medium was replaced with 2 mL of fresh medium containing the designated drug concentrations. The me
Cell migration was assessed using Transwell chambers (Corning, United States) placed in 24-well plates. Serum-starved cells were harvested, resuspended, and adjusted to a density of 3 × 104 cells/mL. A 200 μL aliquot of the cell suspension was added to the upper chamber, while the lower chamber was filled with 800 μL of medium containing 10% FBS as a chemoattractant. Cells were treated with gefitinib (5 μM), atorvastatin (8 μM), or their combination. After 24 hours of incubation, non-migrated cells on the upper surface of the membrane were carefully removed with a cotton swab. The migrated cells on the lower surface were washed with PBS, fixed with 4% formaldehyde for 30 minutes, and stained with 1% crystal violet for 20 minutes. The number of migrated cells was counted in five random fields per membrane under a light microscope, and ImageJ software was used for analysis. The experiment was independently repeated three times.
Following 24 hours of drug treatment, cells were collected, washed with cold PBS, and lysed on ice using RIPA lysis buffer for 40 minutes. The lysates were centrifuged at 12000 × g for 15 minutes at 4 °C, and the supernatant was collected. Protein concentration was determined using a bicinchoninic acid assay kit. Equal amounts of protein were separated by SDS-PAGE and transferred onto polyvinylidene difluoride membranes. The membranes were blocked with 5% non-fat milk for 1 hour at room temperature and then incubated with specific primary antibodies (1:1000 dilution) overnight at 4 °C. After washing with TBST, the membranes were incubated with horseradish peroxidase-conjugated secondary antibodies for 1 hour at room temperature. Protein bands were visualized using an enhanced chemiluminescence substrate.
Cell proliferation was evaluated using a 5-ethynyl-2'-deoxyuridine (EdU) assay kit (C0075S, Beyotime, China). Briefly, cells were seeded into 24-well plates (5 × 104 cells/well), treated for 24 hours, and then incubated with 200 μL of EdU working solution for 2 hours at 37 °C. The cells were fixed with 4% paraformaldehyde and processed according to the manufacturer's protocol.
Total RNA was extracted from cells after 24 hours of culture, and its concentration and purity were measured. Complementary DNA was synthesized by reverse transcription. Quantitative PCR was performed using SYBR Green Master Mix on a real-time PCR system. The relative mRNA expression levels were calculated using the 2-ΔΔCt method. All primers were synthesized by Sangon Biotech (Shanghai, China).
Paraffin-embedded tissue sections were deparaffinized, rehydrated, and stained with hematoxylin and eosin (H&E) following standard protocols. After dehydration and mounting, the sections were examined under a microscope for histological analysis.
Tissue sections were fixed, permeabilized with 0.3% Triton X-100, and blocked with bovine serum albumin. The sections were then incubated overnight at 4 °C with an anti-Ki67 primary antibody (9449T, Cell Signaling Technology, United States). After washing, the sections were incubated with a fluorescently labeled secondary antibody, counterstained with 4',6-diamidino-2-phenylindole, and mounted for imaging.
Female BALB/c nude mice (4-6 weeks old, weighing 23-25 g) were purchased from Silaike Jingda Laboratory Animal Co., Ltd. (Hunan Province, China). A single-cell suspension of HCT116 or SW480 cells was prepared at a density of 5 × 107 cells/mL in PBS. Each mouse was subcutaneously injected with 200 μL of the cell suspension into the right flank. Tumor size was measured weekly with calipers, and tumor volume was calculated using the formula: (Length × width²)/2. Four weeks post-inoculation, the mice were humanely euthanized. Euthanasia was performed by intravenous injection of 3% sodium pentobarbital, followed by cervical dislocation after the loss of consciousness and pain reflex was confirmed. The terminal tumors were excised, and their size, weight, and volume were recorded. The maximum tumor volume did not exceed 1.5 cm³. All animal experiments were approved by the Animal Care and Use Committee of the Third Hospital of Xiangya University and conducted in strict accordance with relevant guidelines and regulations, including the ARRIVE guidelines.
All statistical analyses were performed using SPSS 20.0 and GraphPad Prism 8.0 software. Quantitative data are presented as the mean ± SD. Comparisons between two groups were analyzed using the Student's t-test, while com
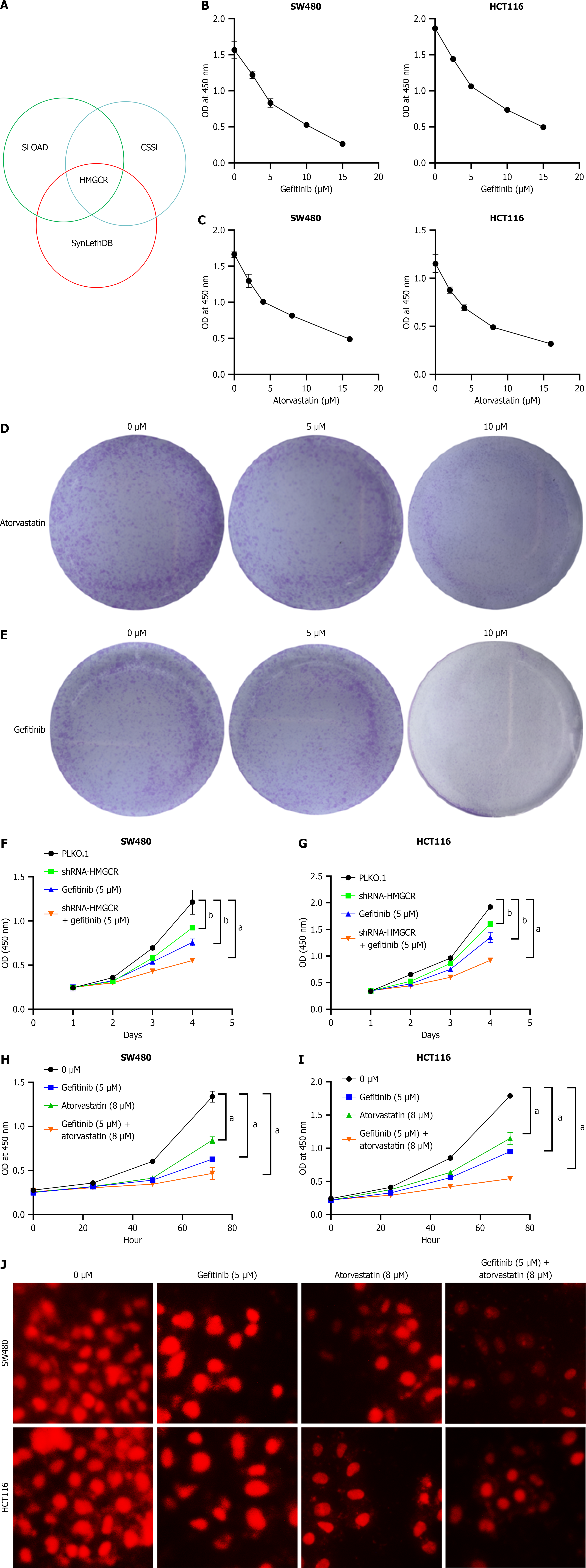
Bioinformatic analysis using the SOLAD, CSSL, and SynLethDB databases predicted a synthetic lethal interaction between HMGCR and PIK3CD in CRC (Figure 1A). To validate this prediction, SW480 and HCT116 CRC cells were treated with gefitinib and atorvastatin. CCK-8 assays demonstrated that the combination of gefitinib and atorvastatin synergistically inhibited cell viability in a concentration-dependent manner (Figure 1B and C). Consistently, colony for
The critical role of HMGCR in this synthetic lethal interaction was confirmed by knockdown experiments. HMGCR depletion significantly attenuated the antiproliferative effects of gefitinib based combination treatment in both cell lines (Figure 1F and G). Compared with single-agent treatments, the combination of atorvastatin and gefitinib exerted a superior inhibitory on the proliferation of SW480 and HCT116 cell (Figure 1H and I). This enhanced antiproliferative effect was further corroborated by EdU assays (Figure 1J). Based on the concentration-dependent inhibitory profiles, the median lethal concentrations (5 μM gefitinib and 8 μM atorvastatin) were determined and selected for subsequent combination therapy experiments.
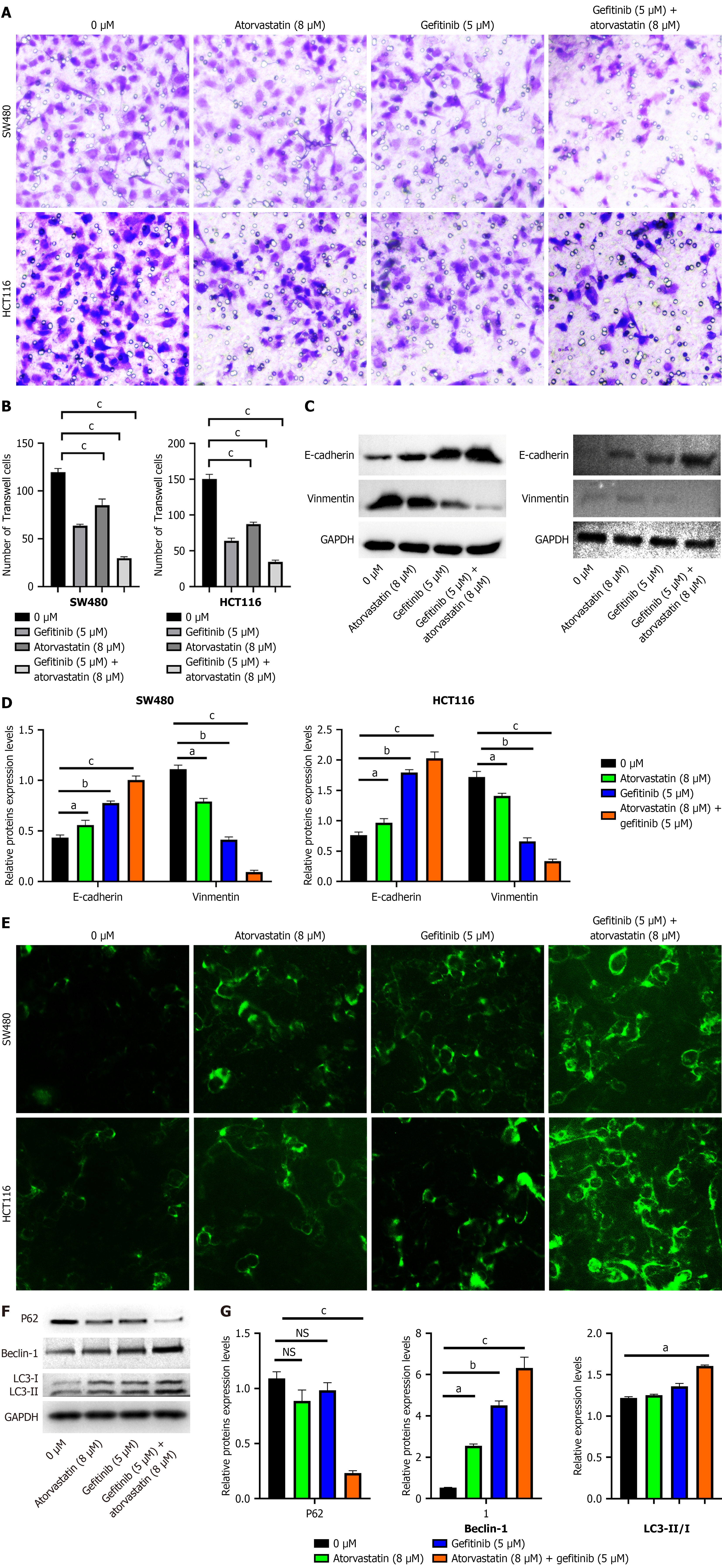
We next investigated the effect of the combination treatment on metastasis and autophagy. Transwell assays showed that the combination of atorvastatin and gefitinib inhibited the migration of SW480 and HCT116 cells than either agent alone (Figure 2A and B). Western blotting indicated that the combination treatment upregulated the epithelial marker E-cadherin and downregulated the mesenchymal marker vimentin, with effects more pronounced than those of single agent treatments (Figure 2C and D), suggesting enhanced suppression of epithelial-mesenchymal transition.
Combination therapy significantly promoted autophagy. Immunofluorescence staining revealed a greater increase in LC3 puncta formation in cells treated with both drugs, indicating enhanced autophagosome formation (Figure 2E). Western blotting of autophagy-related markers consistently showed that the combination treatment increased the LC3-II/I ratio and Beclin-1 expression, while decreasing P62 expression, to a greater extent than single-agent treatments (Figure 2F and G).
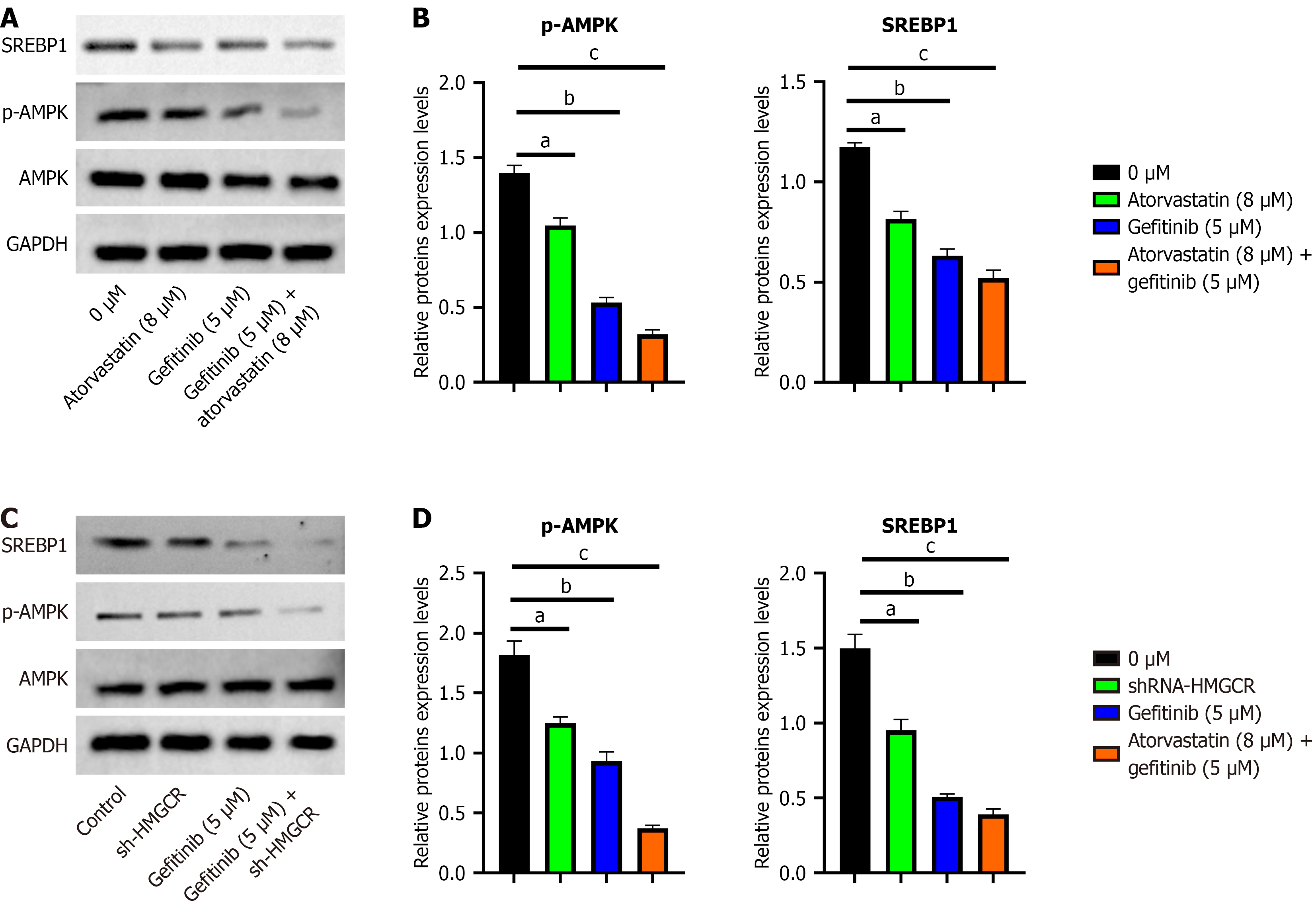
To explore the underlying mechanism, we examined the AMPK-SREBP1 pathway. Western blotting demonstrated that the combination of atorvastatin and gefitinib reduced the levels of p-AMPK and SREBP1 more effectively than either drug alone (Figure 3A and B). Similarly, the combination of HMGCR knockdown (sh-HMGCR) with gefitinib also led to a greater reduction in p-AMPK and SREBP1 levels compared with individual interventions (Figure 3C and D). These results indicated that the combination therapy inhibited the AMPK-SREBP1 signaling pathway.
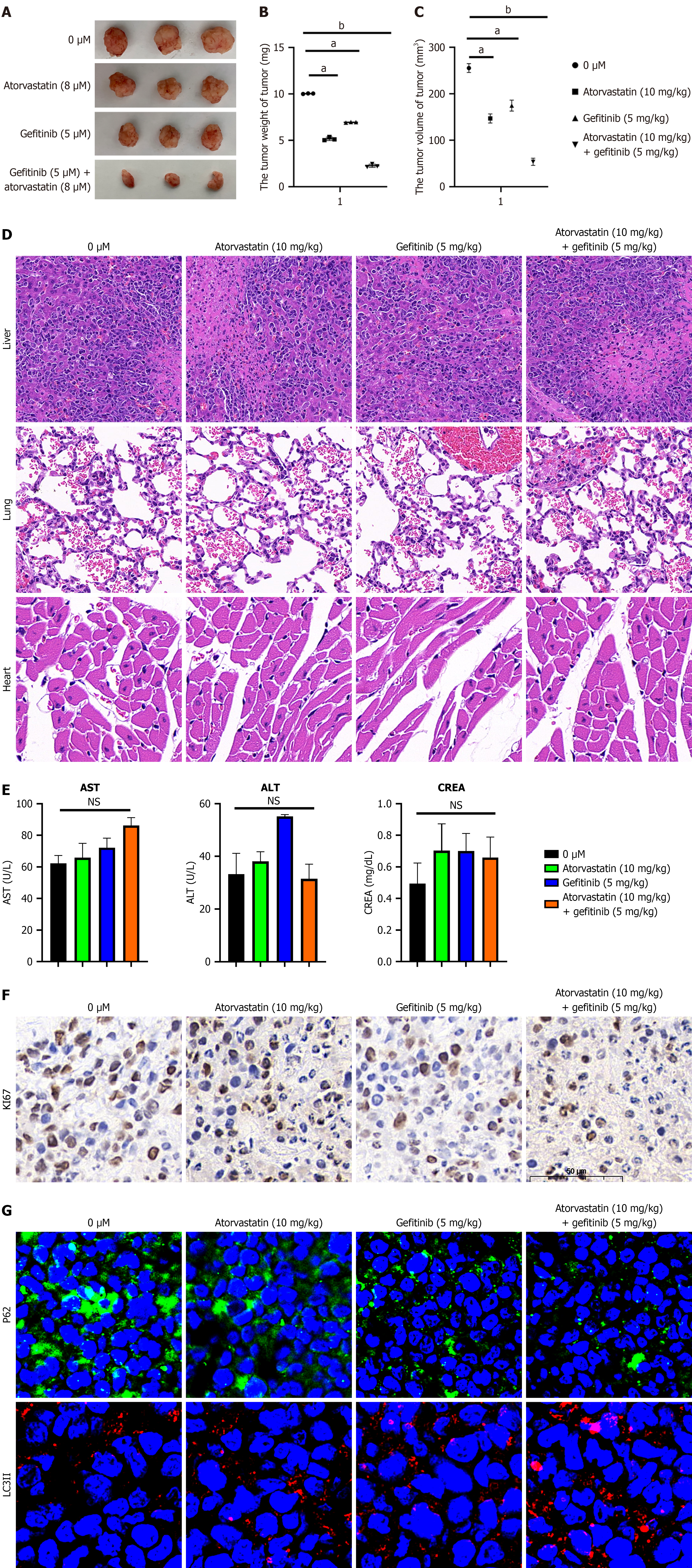
Antitumor efficacy of the combination was evaluated using a mouse xenograft model. Compared with the control or single-agent groups, cotreatment with atorvastatin and gefitinib significantly reduced tumor volume and weight (Figure 4A-C). H&E staining of major organs (heart, lungs, and liver) revealed no obvious pathological damage (Figure 4D). Serum biochemical analysis showed no significant differences in liver function markers (alanine aminotransferase, aspartate aminotransferase) or CREA levels between the control and treatment groups, confirming the favorable safety profile of the combination (Figure 4E).
Immunohistochemical staining for Ki67 showed that the combination treatment suppressed tumor cell proliferation more effectively than either monotherapy (Figure 4F). Additionally, immunofluorescence staining of tumor tissues confirmed enhanced autophagy in the combination group in vivo, as evidenced by increased LC3 puncta and decreased P62 expression (Figure 4G). Collectively, these findings illustrated that the combination atorvastatin and gefitinib exerted potent antitumor activity and clinical potential.
Cancer treatment strategies are contingent on many factors, including tumor origin (sporadic or genetic), molecular profile, patient age, and cancer stage[19,20]. Conventional modalities, such as surgery, chemotherapy, radiotherapy, comprehensive therapy, and immunotherapy remain the cornerstone of clinical management[21,22]. In addition, ad
Statins are widely used for dyslipidemia management, and are now being evaluated in clinical studies for cancer pre
Our findings regarding the synthetic lethal interaction between atorvastatin and gefitinib should be interpreted within the evolving context of statin repurposing in oncology. Previous studies have examined combinations such as ator
Gefitinib and atorvastatin individually inhibited proliferation, invasion, and autophagy induction in SW480 and HCT116 cells. However, their combination resulted in significantly enhanced inhibitory effects. This study elucidated the antitumor mechanism of the gefitinib-atorvastatin combination from an autophagic perspective and provides an experimental foundation for its further development. The combined regimen increased the sensitivity of CRC cells to gefitinib and exhibited a favorable safety profile without significant cardiac, pulmonary, or hepatic toxicity. These findings support the clinical translational potential of this treatment strategy.
| 1. | McCabe MA, Mauro AJ, Schoen RE. Novel colorectal cancer screening methods - opportunities and challenges. Nat Rev Clin Oncol. 2025;22:581-591. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 2. | Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, Jemal A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229-263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16785] [Cited by in RCA: 14733] [Article Influence: 7366.5] [Reference Citation Analysis (19)] |
| 3. | Wu Y, He S, Cao M, Teng Y, Li Q, Tan N, Wang J, Zuo T, Li T, Zheng Y, Xia C, Chen W. Comparative analysis of cancer statistics in China and the United States in 2024. Chin Med J (Engl). 2024;137:3093-3100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 110] [Article Influence: 55.0] [Reference Citation Analysis (1)] |
| 4. | Li Q, Geng S, Luo H, Wang W, Mo YQ, Luo Q, Wang L, Song GB, Sheng JP, Xu B. Signaling pathways involved in colorectal cancer: pathogenesis and targeted therapy. Signal Transduct Target Ther. 2024;9:266. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 398] [Cited by in RCA: 319] [Article Influence: 159.5] [Reference Citation Analysis (2)] |
| 5. | Nunes L, Li F, Wu M, Luo T, Hammarström K, Torell E, Ljuslinder I, Mezheyeuski A, Edqvist PH, Löfgren-Burström A, Zingmark C, Edin S, Larsson C, Mathot L, Osterman E, Osterlund E, Ljungström V, Neves I, Yacoub N, Guðnadóttir U, Birgisson H, Enblad M, Ponten F, Palmqvist R, Xu X, Uhlén M, Wu K, Glimelius B, Lin C, Sjöblom T. Prognostic genome and transcriptome signatures in colorectal cancers. Nature. 2024;633:137-146. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 113] [Cited by in RCA: 103] [Article Influence: 51.5] [Reference Citation Analysis (1)] |
| 6. | Chen L, Dai P, Liu L, Chen Y, Lu Y, Zheng L, Wang H, Yuan Q, Li X. The lipid-metabolism enzyme ECI2 reduces neutrophil extracellular traps formation for colorectal cancer suppression. Nat Commun. 2024;15:7184. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 24] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 7. | Zhao Y, Liu MJ, Zhang L, Yang Q, Sun QH, Guo JR, Lei XY, He KY, Li JQ, Yang JY, Jian YP, Xu ZX. High mobility group A1 (HMGA1) promotes the tumorigenesis of colorectal cancer by increasing lipid synthesis. Nat Commun. 2024;15:9909. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 8. | Ren YM, Zhuang ZY, Xie YH, Yang PJ, Xia TX, Xie YL, Liu ZH, Kang ZR, Leng XX, Lu SY, Zhang L, Chen JX, Xu J, Zhao EH, Wang Z, Wang M, Cui Y, Tan J, Liu Q, Jiang WH, Xiong H, Hong J, Chen YX, Chen HY, Fang JY. BCAA-producing Clostridium symbiosum promotes colorectal tumorigenesis through the modulation of host cholesterol metabolism. Cell Host Microbe. 2024;32:1519-1535.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 60] [Article Influence: 30.0] [Reference Citation Analysis (2)] |
| 9. | Zhou X, Wang G, Tian C, Du L, Prochownik EV, Li Y. Inhibition of DUSP18 impairs cholesterol biosynthesis and promotes anti-tumor immunity in colorectal cancer. Nat Commun. 2024;15:5851. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 26] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 10. | Gabitova-Cornell L, Surumbayeva A, Peri S, Franco-Barraza J, Restifo D, Weitz N, Ogier C, Goldman AR, Hartman TR, Francescone R, Tan Y, Nicolas E, Shah N, Handorf EA, Cai KQ, O'Reilly AM, Sloma I, Chiaverelli R, Moffitt RA, Khazak V, Fang CY, Golemis EA, Cukierman E, Astsaturov I. Cholesterol Pathway Inhibition Induces TGF-β Signaling to Promote Basal Differentiation in Pancreatic Cancer. Cancer Cell. 2020;38:567-583.e11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 177] [Cited by in RCA: 160] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 11. | Stouth DW, Lebeau PF, Austin RC. Repurposing Two Old Friends to Fight Cancer: Caffeine and Statins. Cancer Res. 2023;83:2091-2092. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 12. | Hillis AL, Martin TD, Manchester HE, Högström J, Zhang N, Lecky E, Kozlova N, Lee J, Persky NS, Root DE, Brown M, Cichowski K, Elledge SJ, Muranen T, Fruman DA, Barry ST, Clohessy JG, Madsen RR, Toker A. Targeting Cholesterol Biosynthesis with Statins Synergizes with AKT Inhibitors in Triple-Negative Breast Cancer. Cancer Res. 2024;84:3250-3266. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 21] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 13. | Maier CR, Hartmann O, Prieto-Garcia C, Al-Shami KM, Schlicker L, Vogel FCE, Haid S, Klann K, Buck V, Münch C, Schmitz W, Einig E, Krenz B, Calzado MA, Eilers M, Popov N, Rosenfeldt MT, Diefenbacher ME, Schulze A. USP28 controls SREBP2 and the mevalonate pathway to drive tumour growth in squamous cancer. Cell Death Differ. 2023;30:1710-1725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 27] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 14. | Prindle V, Richardson AE, Sher KR, Kongpachith S, Kentala K, Petiwala S, Cheng D, Widomski D, Le P, Torrent M, Chen A, Walker S, Palczewski MB, Mitra D, Manaves V, Shi X, Lu C, Sandoval S, Dezso Z, Buchanan FG, Verduzco D, Bierie B, Meulbroek JA, Pappano WN, Plotnik JP. Synthetic lethality of mRNA quality control complexes in cancer. Nature. 2025;638:1095-1103. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 15. | Ryan CJ, Devakumar LPS, Pettitt SJ, Lord CJ. Complex synthetic lethality in cancer. Nat Genet. 2023;55:2039-2048. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 48] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 16. | Li S, Topatana W, Juengpanich S, Cao J, Hu J, Zhang B, Ma D, Cai X, Chen M. Development of synthetic lethality in cancer: molecular and cellular classification. Signal Transduct Target Ther. 2020;5:241. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 103] [Cited by in RCA: 86] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 17. | Strobl CD, Schaffer S, Haug T, Völkl S, Peter K, Singer K, Böttcher M, Mougiakakos D, Mackensen A, Aigner M. Selective PRMT5 Inhibitors Suppress Human CD8(+) T Cells by Upregulation of p53 and Impairment of the AKT Pathway Similar to the Tumor Metabolite MTA. Mol Cancer Ther. 2020;19:409-419. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 18. | Maio M, Ascierto PA, Manzyuk L, Motola-Kuba D, Penel N, Cassier PA, Bariani GM, De Jesus Acosta A, Doi T, Longo F, Miller WH, Oh DY, Gottfried M, Xu L, Jin F, Norwood K, Marabelle A. Pembrolizumab in microsatellite instability high or mismatch repair deficient cancers: updated analysis from the phase II KEYNOTE-158 study. Ann Oncol. 2022;33:929-938. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 388] [Cited by in RCA: 313] [Article Influence: 78.3] [Reference Citation Analysis (8)] |
| 19. | Jaber D, Zhang J, Godley LA. Detecting likely germline variants during tumor-based molecular profiling. J Clin Invest. 2025;135:e190264. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 20. | Eslinger C, Seddighzadeh B, Yee C, Elsabbagh Z, Pai R, Hartley C, Starr J, Bekaii-Saab T, Halfdanarson TR, Sonbol MB. Clinical Outcomes and Molecular Profiling of Pancreatic Acinar Cell Carcinoma: A Retrospective Study. JCO Precis Oncol. 2025;9:e2400450. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 21. | Lei G, Zhuang L, Gan B. The roles of ferroptosis in cancer: Tumor suppression, tumor microenvironment, and therapeutic interventions. Cancer Cell. 2024;42:513-534. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 462] [Cited by in RCA: 382] [Article Influence: 191.0] [Reference Citation Analysis (1)] |
| 22. | Tarannum M, Ding X, Barisa M, Hu S, Anderson J, Romee R, Zhang J. Engineering innate immune cells for cancer immunotherapy. Nat Biotechnol. 2025;43:516-533. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 26] [Article Influence: 26.0] [Reference Citation Analysis (0)] |
| 23. | Kath J, Franke C, Drosdek V, Du W, Glaser V, Fuster-Garcia C, Stein M, Zittel T, Schulenberg S, Porter CE, Andersch L, Künkele A, Alcaniz J, Hoffmann J, Abken H, Abou-El-Enein M, Pruß A, Suzuki M, Cathomen T, Stripecke R, Volk HD, Reinke P, Schmueck-Henneresse M, Wagner DL. Integration of ζ-deficient CARs into the CD3ζ gene conveys potent cytotoxicity in T and NK cells. Blood. 2024;143:2599-2611. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 32] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 24. | Hu Y, Zhang M, Yang T, Mo Z, Wei G, Jing R, Zhao H, Chen R, Zu C, Gu T, Xiao P, Hong R, Feng J, Fu S, Kong D, Xu H, Cui J, Huang S, Liang B, Yuan X, Cui Q, Guo H, Yu Y, Feng Y, Jin C, Ren J, Chang AH, Wang D, Huang H. Sequential CD7 CAR T-Cell Therapy and Allogeneic HSCT without GVHD Prophylaxis. N Engl J Med. 2024;390:1467-1480. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 104] [Cited by in RCA: 83] [Article Influence: 41.5] [Reference Citation Analysis (0)] |
| 25. | Lin MT, Zheng G, Rodriguez E, Tseng LH, Parini V, Xian R, Zou Y, Gocke CD, Eshleman JR. Double PIK3CA Alterations and Parallel Evolution in Colorectal Cancers. Am J Clin Pathol. 2022;157:244-251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 26. | Farin HF, Mosa MH, Ndreshkjana B, Grebbin BM, Ritter B, Menche C, Kennel KB, Ziegler PK, Szabó L, Bollrath J, Rieder D, Michels BE, Kress A, Bozlar M, Darvishi T, Stier S, Kur IM, Bankov K, Kesselring R, Fichtner-Feigl S, Brüne B, Goetze TO, Al-Batran SE, Brandts CH, Bechstein WO, Wild PJ, Weigert A, Müller S, Knapp S, Trajanoski Z, Greten FR. Colorectal Cancer Organoid-Stroma Biobank Allows Subtype-Specific Assessment of Individualized Therapy Responses. Cancer Discov. 2023;13:2192-2211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 80] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 27. | Zheng H, Liu H, Li H, Dou W, Wang J, Zhang J, Liu T, Wu Y, Liu Y, Wang X. Characterization of stem cell landscape and identification of stemness-relevant prognostic gene signature to aid immunotherapy in colorectal cancer. Stem Cell Res Ther. 2022;13:244. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 120] [Cited by in RCA: 113] [Article Influence: 28.3] [Reference Citation Analysis (0)] |
| 28. | Chen Z, Cao H, Zhang J, Zhong W, Teng X. The impact of preoperative treatment on mismatch repair protein and HER2 expression in colorectal cancer: an analysis of paired samples. BMC Cancer. 2024;24:1407. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 29. | Li L, Wang H, Zhang S, Gao S, Lu X, Pan Y, Tang W, Huang R, Qiao K, Ning S. Statins inhibit paclitaxel-induced PD-L1 expression and increase CD8+ T cytotoxicity for better prognosis in breast cancer. Int J Surg. 2024;110:4716-4726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 16] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 30. | Han JX, Tao ZH, Wang JL, Zhang L, Yu CY, Kang ZR, Xie Y, Li J, Lu S, Cui Y, Xu J, Zhao E, Wang M, Chen J, Wang Z, Liu Q, Chen HM, Su W, Zou TH, Zhou CB, Hong J, Chen H, Xiong H, Chen YX, Fang JY. Microbiota-derived tryptophan catabolites mediate the chemopreventive effects of statins on colorectal cancer. Nat Microbiol. 2023;8:919-933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 158] [Article Influence: 52.7] [Reference Citation Analysis (1)] |
| 31. | Gash KJ, Chambers AC, Cotton DE, Williams AC, Thomas MG. Potentiating the effects of radiotherapy in rectal cancer: the role of aspirin, statins and metformin as adjuncts to therapy. Br J Cancer. 2017;117:210-219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 32. | Ridker PM, Bhatt DL, Pradhan AD, Glynn RJ, MacFadyen JG, Nissen SE; PROMINENT, REDUCE-IT, and STRENGTH Investigators. Inflammation and cholesterol as predictors of cardiovascular events among patients receiving statin therapy: a collaborative analysis of three randomised trials. Lancet. 2023;401:1293-1301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 540] [Cited by in RCA: 462] [Article Influence: 154.0] [Reference Citation Analysis (0)] |
| 33. | Xu XT, Chen J, Ren X, Ma YR, Wang X, Ma YY, Zhao DG, Zhou RP, Zhang K, Goodin S, Li DL, Zheng X. Effects of atorvastatin in combination with celecoxib and tipifarnib on proliferation and apoptosis in pancreatic cancer sphere-forming cells. Eur J Pharmacol. 2021;893:173840. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 34. | Zhang QS, Deater M, Phan N, Marcogliese A, Major A, Guinan EC, Grompe M. Combination therapy with atorvastatin and celecoxib delays tumor formation in a Fanconi anemia mouse model. Pediatr Blood Cancer. 2019;66:e27460. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 7] [Article Influence: 1.0] [Reference Citation Analysis (0)] |