Published online Apr 16, 2026. doi: 10.4253/wjge.v18.i4.117976
Revised: January 11, 2026
Accepted: March 3, 2026
Published online: April 16, 2026
Processing time: 114 Days and 17.4 Hours
Endoscopic ultrasound (EUS) remains operator-dependent with notable diag
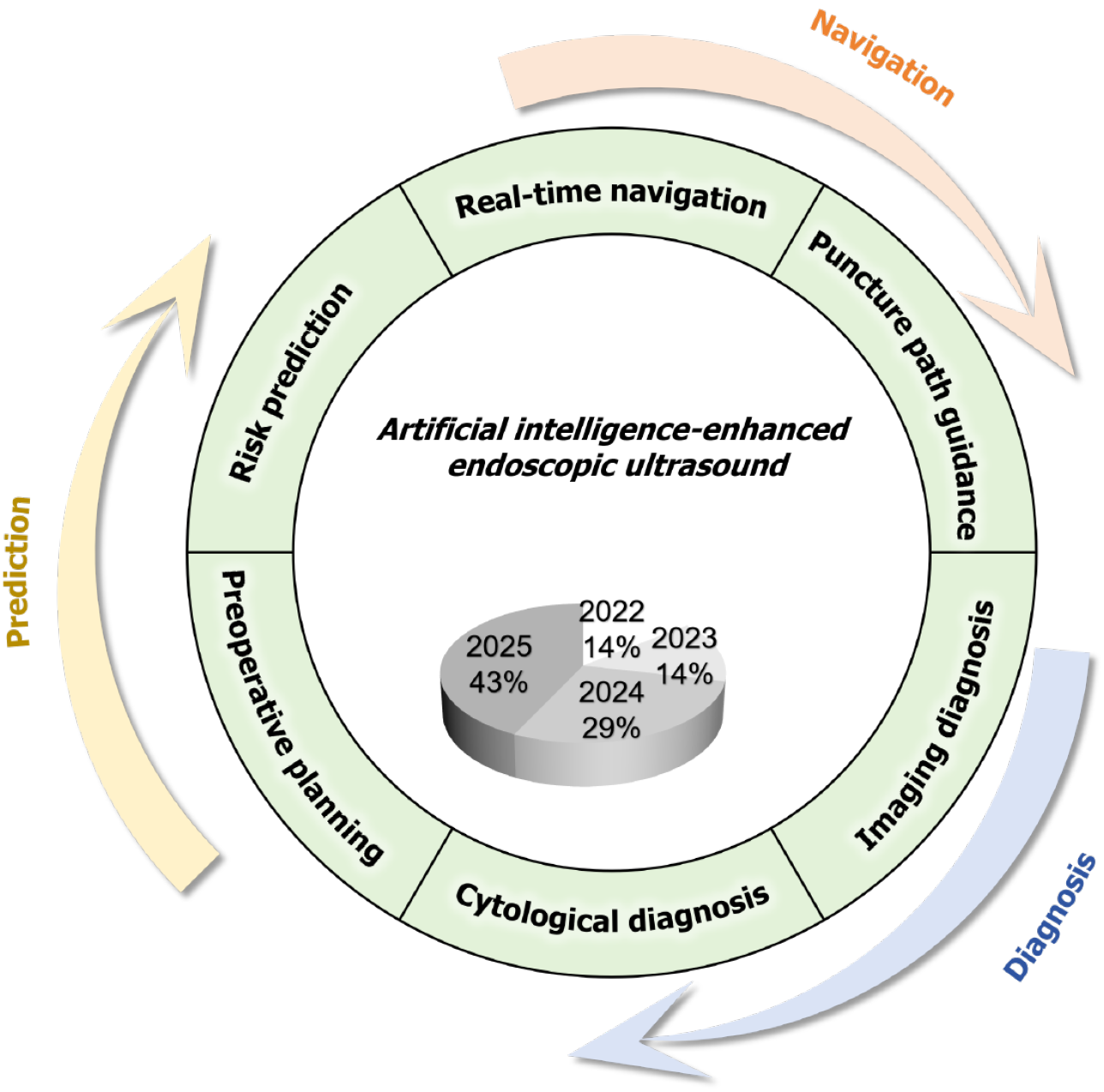
Core Tip: This article proposes an artificial intelligence-powered “Prediction-Navigation-Diagnosis” framework for endoscopic ultrasound. It highlights how artificial intelligence improves preoperative risk stratification, reduces intraoperative missed detections (about 10%) via real-time navigation, and enhances diagnostic accuracy. Key challenges for clinical adoption - data heterogeneity, high costs, and unclear regulations - are summarized, with future success hinging on multimodal integration and adaptive policies.
- Citation: Chen ZY, Wang YQ, Tan XZ, Liu P, Peng Y. Artificial intelligence in endoscopic ultrasound: Clinical translation of a prediction, navigation, and diagnosis framework. World J Gastrointest Endosc 2026; 18(4): 117976
- URL: https://www.wjgnet.com/1948-5190/full/v18/i4/117976.htm
- DOI: https://dx.doi.org/10.4253/wjge.v18.i4.117976
Endoscopic ultrasound (EUS) represents a cornerstone technique for diagnosing and treating digestive diseases, particularly valuable in early pancreatic cancer detection, differentiation of submucosal tumors, and guidance of fine-needle aspiration (FNA)[1]. However, the effectiveness of EUS remains highly operator-dependent, resulting in significant diagnostic variability, subjective image interpretation, and a high missed detection rate in anatomical blind zones. These limitations collectively increase procedural risks and reduce diagnostic consistency. Artificial intelligence (AI) has em
PubMed, EMBASE, IEEE Xplore, and China National Knowledge Infrastructure.
EUS combined with AI/deep learning and Computer-Aided Diagnosis/Real-Time Navigation/FNA.
Inclusion criteria: Original studies on EUS-AI models published between 2022 and 2025.
Exclusion criteria: Studies lacking clinical validation or containing duplicate data.
The risk of bias in included studies was assessed using the Prediction model Risk Of Bias Assessment Tool. Most current evidence originates from single-center, retrospective case series with limited sample sizes and without prospective external validation. Consequently, the overall risk of bias is high, primarily due to non-representative populations, lack of blinding, and absence of validation in independent cohorts. These limitations signify that the reported performance metrics may not generalize to broader clinical settings. It must be acknowledged that the field remains in a relatively early translational stage; high-quality evidence from large-scale, prospective, multicenter trials is still scarce. Therefore, AI-enhanced EUS should currently be regarded as an assistive, rather than a replacement, technology in clinical practice.
The diagnostic accuracy of EUS depends significantly on the operator, resulting in considerable inter-observer variability in lesion interpretation. For example, the diagnostic accuracy of conventional EUS for gastric stromal tumors ranges from only 45% to 67%[3,4]. Additionally, EUS imaging alone has inherent limitations in differentiating specific lesion types. Specifically, distinguishing benign from malignant solid pancreatic masses based solely on EUS images has limited specificity (50% to 60%), increasing the risk of misdiagnosis[5]. Variations in operator experience also cause procedural blind spots and missed lesions. Critical anatomical regions, such as the pancreatic head and the pancreatobiliary junction, are frequently overlooked, with missed lesion rates ranging from 70% to 93% among operators[6].
Several procedure-related complications limit the safety of EUS-guided FNA (EUS-FNA). Post-procedural acute pancreatitis occurs in approximately 1.8% of cases, often due to iatrogenic injury of the pancreatic ductal system[7]. Perforation is less common (around 0.4%), influenced by operator inexperience, anatomical variations, and pre-existing conditions such as gastrointestinal strictures or diverticula[8]. Hemorrhage occurs in nearly 2% of procedures, with increased risk when targeting highly vascular organs like the pancreas[8]. Furthermore, inadequate specimen acquisition is a persistent issue, potentially reducing diagnostic yield and necessitating repeat procedures[9].
As an EUS-based modality, EUS is susceptible to imaging artifacts, such as reverberation, acoustic shadowing, and speckle noise, which complicate interpretation and require considerable expertise to identify and reduce. Although high-frequency transducers (typically 5-20 MHz) provide high spatial resolution, their penetration depth is limited to a few centimeters. Consequently, deep anatomical structures distal to the probe, such as retroperitoneal portions of large tumors or pancreatic visualization in obese patients, are often inadequately visualized, reducing diagnostic clarity.
In summary, conventional EUS faces significant limitations, including pronounced operator-dependent variability in diagnostic performance, procedure-related complications such as pancreatitis (1.8%), perforation (0.4%), hemorrhage (2%), frequent difficulties in obtaining adequate tissue samples, and fundamental imaging limitations inherent to EUS physics.
EUS can detect biliary sludge and microlithiasis, known risk factors for acute pancreatitis[10]. Given its invasiveness and cost, proper patient selection is essential to avoid both overuse and underdiagnosis. Sirtl et al[11] developed a machine learning model using data from 218 acute pancreatitis patients across three tertiary centers. The model incorporated age, liver enzymes, lipids, and urinalysis to predict the risk of biliary sludge/microlithiasis. It achieved 97.9% sensitivity, capturing nearly all sludge-positive cases, and a negative predictive value of 78.6%, thereby reducing unnecessary EUS in low-risk patients. A web-based tool was also created for EUS triage based on patient metrics[12]. Despite certain limi
Endoscopic resection is the first-line minimally invasive treatment for subepithelial lesions (SELs), highlighting the need for precise preoperative risk assessment[13]. Using EUS data from over 1000 patients across eight hospitals in Shandong, Chen et al[14] developed the SEL Endoscopic Resection Predictor. Based on lesion location, size, morphology, and echogenicity, SEL Endoscopic Resection Predictor predicts four clinical outcomes: Ulcer management (standard vs advanced closure), procedure time > 30 minutes, hospital stay ≥ 5 days, and medication costs exceeding hospital averages. Although rare outcomes such as perforation were excluded due to low incidence, larger studies may further expand their predictive capabilities. Such tools support personalized planning, resource allocation, and preoperative counseling.
EUS is crucial for diagnosing digestive tract diseases[15-17]. However, its effectiveness varies widely among operators, with miss rates ranging from 70% to 93% due to anatomical blind spots[6]. Several AI-powered systems have been developed to improve scanning standardization and procedural guidance. Wu et al[18] created EUS-intelligent and real-time endoscopy analytical device, a real-time AI assistant that identifies eight standard scanning positions and 24 an
In summary, these systems exemplify a new generation of AI-augmented EUS platforms: EUS-intelligent and real-time endoscopy analytical device serves as a scanning supervisor, EUS-automatic image report system as an automated re
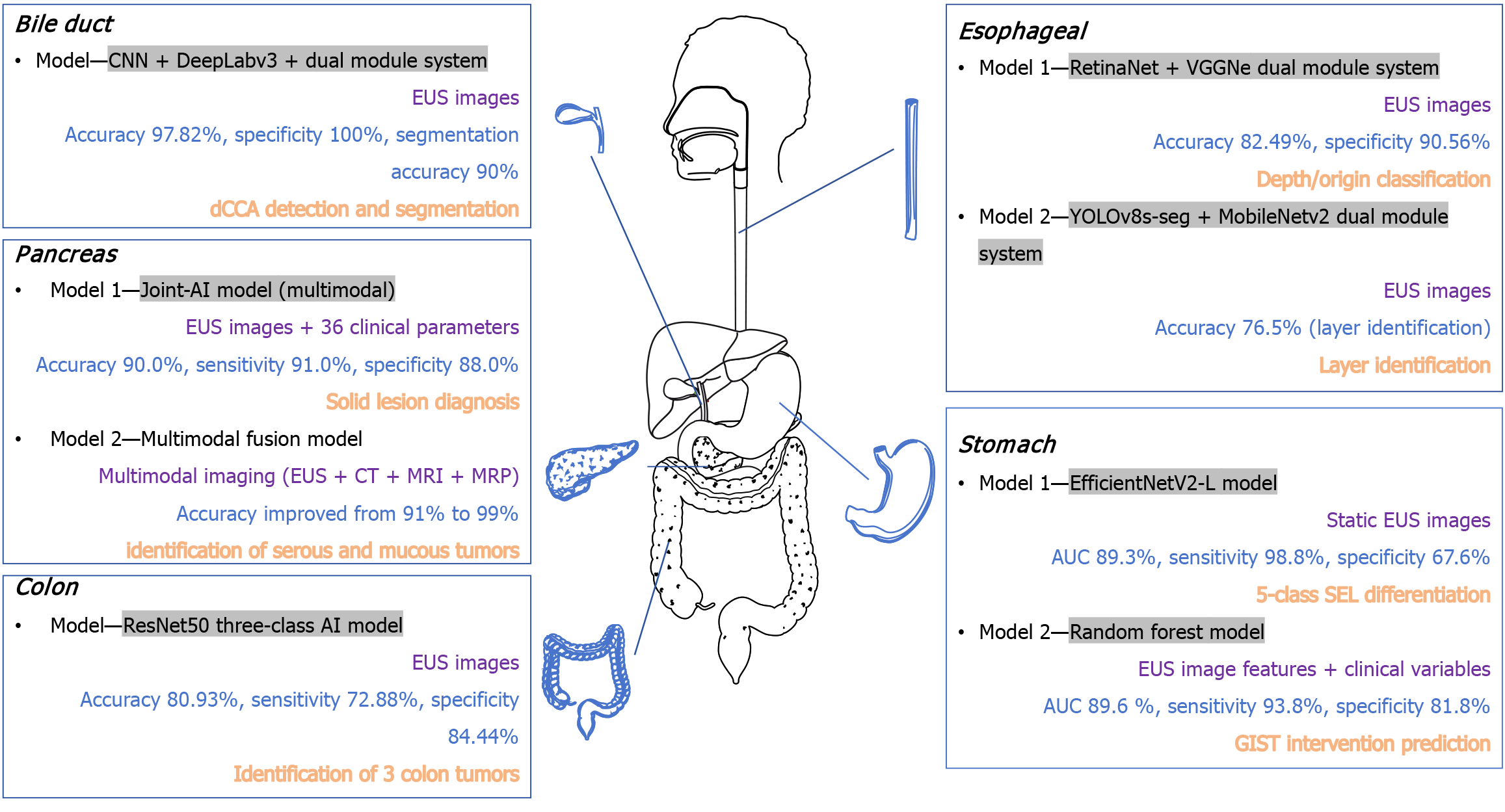
Imaging-driven AI diagnostic models: Gastric mesenchymal tumors, such as gastrointestinal stromal tumors (GIST), leiomyomas, schwannomas, and other SELs, are commonly identified during gastroscopy[21,22]. However, their differentiation through EUS is challenging, with conventional diagnostic accuracy ranging from 45% to 67% due to high operator variability[3,4]. To address this issue, Hirai et al[23] developed a deep learning model using EfficientNetV2-L trained on 631 pathologically confirmed SEL cases from 12 institutions. The model achieved an overall accuracy of 86.1% in five-class classification (GIST, leiomyoma, schwannoma, neuroendocrine tumor, and ectopic pancreas) and a sensitivity of 98.8% for GIST detection. However, accurately distinguishing schwannomas remains challenging. A limitation of this model is its current use of static images, without incorporating dynamic EUS features (e.g., elastography, Doppler flow) or clinical metadata. In contrast, Joo et al[24] developed a multimodal fusion model integrating EUS image features with clinical variables (patient age, tumor location) using machine learning (random forest). This model achieved an area under the curve of 0.896 and a sensitivity of 93.8%, significantly improving the identification of GIST cases requiring surgery and providing robust clinical decision support. Together, these developments highlight the potential of unimodal and multimodal AI approaches to enhance diagnostic accuracy and standardize SEL evaluations using EUS.
Subepithelial esophageal lesions, including leiomyomas and GIST, typically follow a benign course, though some carry malignant potential. Considering China’s high esophageal cancer incidence (five-year survival: 15%-25%)[25-27], EUS is crucial for localizing lesions (mucosal, submucosal, muscularis propria)[28-30]. However, EUS diagnostic accuracy depends significantly on the operator[31]. To address this issue, Liu et al[32] developed a dual-module deep learning system (RetinaNet-based locator, VGGNet-based classifier). It achieved 82.49% accuracy, 90.56% specificity, and 99.3% accuracy for determining tumor origin, significantly outperforming conventional methods. Similarly, Zhang et al[33] developed a lightweight “detection-stratification” model (YOLOv8s-seg for lesion contouring, MobileNetv2 for layer identification). This system achieved 92.2% precision in lesion contouring and 76.5% accuracy in distinguishing superficial from deep lesions, comparable to senior endoscopists (74.9%-79.8%) and outperforming junior practitioners (65.6%). Despite promising results, both models require multi-center validation, real-time video analytics, and mul
Solid pancreatic lesions present diagnostic challenges due to ambiguous etiology (benign vs malignant)[34-37]. Although EUS is standard for morphological evaluation, conventional interpretation based solely on EUS imaging features has limited specificity (50%-60%)[38]. Cui et al[39] developed a multimodal AI system (Joint-AI) combining deep learning-based EUS image analysis with 36 clinical parameters (age, symptoms, lab markers). In a cross-validation trial involving 12 endoscopists, this model improved junior endoscopists’ diagnostic accuracy by 21% (69%-90%) and sensitivity by 30% (61%-91%). Senior endoscopists initially skeptical of AI assistance showed increased adoption when given interpretable decision support (visual highlights, explanatory annotations). Future implementation requires addressing clinical integration challenges and establishing clear accountability for AI-assisted diagnostics.
Distal cholangiocarcinoma is a rare but highly aggressive malignancy with nonspecific early symptoms and poor five-year survival (10%-40%)[40,41]. Orzan et al[42] developed the first EUS-based AI model specifically for biliary tract evaluation (non-pancreatic), using data from 112 distal cholangiocarcinoma patients and 1248 EUS images. The dual-module system included a customized convolutional neural network for classification (97.82% accuracy, 100% specificity) and a segmentation module (DeepLabv3+, ResNet50 backbones), achieving 90% accuracy in bile duct segmentation and improved tumor margin identification. Although significant, further large-scale, multi-center validation and real-time video integration are needed for clinical adoption.
Colorectal cancer is the third most diagnosed malignancy worldwide[43]. Unlike colonoscopy, which is limited to superficial evaluation, EUS allows detailed visualization of deeper tumor structures (adenomas vs carcinomas)[44,45]. In a multicenter study involving 554 patients and 8738 EUS images, Men et al[46] created an AI model based on the ResNet50 architecture for triple-classification (carcinoma, adenoma, benign tumors). The model achieved 80.93% accuracy, 72.88% sensitivity, and 84.44% specificity for malignant tumors, outperforming senior endoscopists (κ = 0.674 vs κ = 0.557) and substantially surpassing junior endoscopists (κ = 0.284). This demonstrates AI’s potential to standardize colorectal tumor classification and reduce experience-based variability.
In summary, AI-augmented EUS significantly improves diagnostic accuracy across various gastrointestinal conditions (gastric tumors, esophageal lesions, pancreaticobiliary diseases, colorectal neoplasms; Figure 2). Currently, most AI models face the following common challenges in real-world clinical application: Model performance is highly dependent on the quality of training data and annotation consistency, and diagnostic accuracy may significantly decrease in cases of poor image quality, atypical lesion presentation, or rare diseases. Particularly noteworthy is that static image-based analysis methods fail to fully integrate the incremental information provided by dynamic EUS imaging modalities, such as elastography and doppler flow imaging. Furthermore, the generalizability of models still requires systematic validation in real-world clinical environments involving multicenter, prospective designs, diverse acquisition devices, and operators. Future research should focus on developing AI systems capable of integrating dynamic imaging se
Cytology-driven AI diagnostic models: Although imaging-based AI has significantly improved diagnostic precision, histopathological assessment via EUS-FNA remains the gold standard[47-49]. Its utility, however, is limited by a scarcity of pathologists, as nearly 80% of institutions cannot perform rapid on-site evaluation. This shortage results in diagnostic inconsistencies, especially in equivocal cases such as distinguishing inflammatory changes from early malignancies[50,51]. To address these challenges, Fujii et al[52] developed a transformer-based rapid on-site evaluation-AI system trained on 4059 cytological images, achieving an accuracy of 88.2%, a sensitivity of 91.4%, and an area under the curve of 0.954 through geometric transformation-based augmentation. Fang et al[53] proposed a complementary semi-supervised convolutional neural network, reducing manual annotation by 80% while attaining an internal accuracy of 95.1% and a Kappa of 0.853 for diagnosing challenging cases. Despite these advances, both systems require further validation through multicenter studies and improved clinical integration. Future efforts should aim to integrate cytology-based AI models with EUS imaging to develop unified systems that support puncture planning and real-time diagnostic feedback, thereby decreasing reliance on specialized pathology resources.
AI performance in complex clinical scenarios: While existing AI models excel in typical cases, their generalizability in complex scenarios - such as co-existing lesions, rare tumors, and discrimination between inflammation and malignancy - remains to be validated. For instance, identifying early pancreatic cancer against a background of chronic pancreatitis[39] often leads to a significant drop in AI sensitivity. Furthermore, current models struggle to distinguish between histologic subtypes with similar EUS features, such as neuroendocrine tumors G1 vs G2[23]. Future efforts should focus on constructing multicenter databases of complex cases and developing few-shot learning and uncertainty quantification methods to enhance AI robustness in edge-case scenarios.
Post-EUS-FNA pancreatitis occurs in approximately 1.8% of procedures, with a similar incidence (2%) reported for hemorrhage[7,8]. Current risk prediction primarily relies on clinical scoring systems, such as those provided by the American Society for Gastrointestinal Endoscopy and European Society of Gastrointestinal Endoscopy guidelines, which offer limited granularity for personalized assessments[54,55]. Although no dedicated AI prediction model specifically targets EUS-FNA-related complications, Brenner et al[56] developed a machine learning tool for predicting post-endoscopic retrograde cholangiopancreatography pancreatitis. This model includes 20 variables, such as patient age and procedural complexity[56], and provides real-time risk stratification while simulating the effects of preventive interventions (non-steroidal anti-inflammatory drugs, hydration, stenting) on individual risk reduction. Future research could integrate procedural metrics (number of needle passes, needle type), patient biomarkers (baseline amylase levels), and AI-quantified imaging characteristics (vascularity patterns) to develop accurate prediction models for complications like pancreatitis and bleeding after EUS-FNA.
In summary, AI technologies now support the entire EUS workflow within a multidimensional framework. To facilitate comparative evaluation, Table 1 systematically summarizes representative AI models across four key domains: Preoperative planning, intraoperative navigation, imaging-based diagnosis, and cytology-based diagnosis, including details on algorithm architecture, dataset size, performance metrics, feature types, and validation methodologies.
| Application field | Ref. | Algorithm type | Sample size | Validation method | Application scenarios | Accuracy (%) | Sensitivity/specificity (%) | PPV (%) | NPV (%) | F1 score | AUC (95%CI) | Public availability/ reproducibility |
| Pre-procedural | Sirtl et al[11], 2023 | Machine Learning | 218 patients | Multicenter retrospective | Biliary sludge risk screening | 84 | 97.9/92 | 89.5 | 98.2 | 0.93 | 0.96 (0.92-0.98) | Code: Not available; data: Not shared |
| Chen et al[14], 2024 | Predictive model | > 1000 patients | Multicenter prospective | Endoscopic resection outcome prediction | NA | NA | NA | NA | NA | NA | Model: Not released; data: Institutional | |
| Intra-procedural | Wu et al[18], 2023 | 5 DL models ensemble | 290 patients | RCT | Anatomical landmark tracking | NA | NA | NA | NA | NA | NA | Code: Not available; system: Proprietary (EUS-IREAD) |
| Li et al[19], 2025 | Automated reporting system | 114 patients | Prospective trial | Standard site documentation | 90.3 | NA | NA | NA | NA | NA | Code: Closed source; data: Not public | |
| Rizzatti et al[20], 2025 | DCNN | 550 cases | Technical validation | Real-time anatomical navigation | NA | NA | NA | NA | NA | NA | Not publicly available | |
| Image diagnosis | Hirai et al[23], 2022 | EfficientNetV2-L | 631 cases | Multicenter retrospective | 5-class SEL differentiation | 89.3 | 98.8/67.6 | 85.4 | 94.2 | 0.91 | 0.94 (0.90-0.97) | Code: Not shared; data: Multi-institutional (restricted) |
| Joo et al[24], 2024 | Random forest | NA | Single-center | GIST intervention prediction | 89.6 | 93.8/81.8 | 88.9 | 89.5 | 0.91 | 0.896 (0.86-0.93) | Code: Not available; data: Not disclosed | |
| Liu et al[32], 2022 | RetinaNet + VGGNet | NA | Single-center | Depth/origin classification | 82.5 | 80.2/90.6 | 87.1 | 85 | 0.83 | 0.88 (0.84-0.92) | Not publicly released | |
| Zhang et al[33], 2025 | YOLOv8s-seg + MobileNetv2 | NA | Single-center | Layer identification | 76.50 | NA | NA | NA | NA | NA | Code: Not available; model: Proprietary | |
| Cui et al[39], 2024 | Joint-AI | 12 physicians | RCT | Solid lesion diagnosis | 90.0 | 91.0/88.0 | 89.5 | 89.8 | 0.9 | 0.93 (0.89-0.96) | Code: Not shared; data: Anonymized, available on request | |
| Orzan et al[42], 2024 | CNN + DeepLabv3+ | 112 cases (1248 images) | Single-center | dCCA detection and segmentation | 97.8 | 100.0/94.4 | 96.8 | 100 | 0.98 | 0.99 (0.97-1.00) | Not publicly available | |
| Men et al[46], 2025 | ResNet50 | 554 cases (8738 images) | Dual-center | Identification of 3 colon tumors | 80.9 | 72.9/84.4 | 78.5 | 79.8 | 0.75 | 0.85 (0.81-0.89) | Code: Not released; data: Multi-center (restricted) | |
| Cytology diagnosis | Fujii et al[52], 2024 | Transformer | 45 cases (4059 images) | Augmentation test | Malignant cell detection | 88.2 | 91.4/85.0 | 86.7 | 90.1 | 0.89 | 0.954 (0.93-0.98) | Code: GitHub; data: Partially open |
| Fang et al[53], 2025 | SSCNN | NA | Semi-supervised framework | Cytology analysis with limited labels | 95.1 | 93.8/97.3 | 96.5 | 94.8 | 0.95 | 0.97 (0.94-0.99) | Code: Not available; model: Internal use only |
The clinical translation of EUS-AI crucially hinges on reproducibility and rigorous validation. Widespread adoption requires open-source initiatives sharing code, datasets, and model weights, supported by platforms like the Medical Imaging and Data Resource Center. A major translational bottleneck is the frequent lack of independent external validation for published models. Most studies are single-center and retrospective, limiting generalizability across diverse populations and equipment. This validation gap directly undermines real-world reliability and delays clinical integration, as performance often drops outside controlled development settings. Therefore, prioritizing transparent, multi-institutional validation frameworks is essential for building trust and facilitating deployment.
Most advanced AI systems (e.g., Endoangel) rely on specific endoscopic platforms (such as Olympus), which limits their adoption in healthcare institutions with heterogeneous equipment. To achieve practical vendor-agnostic integration, several technical pathways can be pursued: (1) Developing digital imaging and communications in medicine structured report standards-compliant modules for standardized EUS data export; (2) Implementing middleware to convert proprietary data streams into a unified format; and (3) Adopting privacy-preserving distributed learning techniques such as federated learning, enabling multi-center model training without sharing raw data. These concrete steps enhance interoperability and reduce dependency on single-vendor ecosystems.
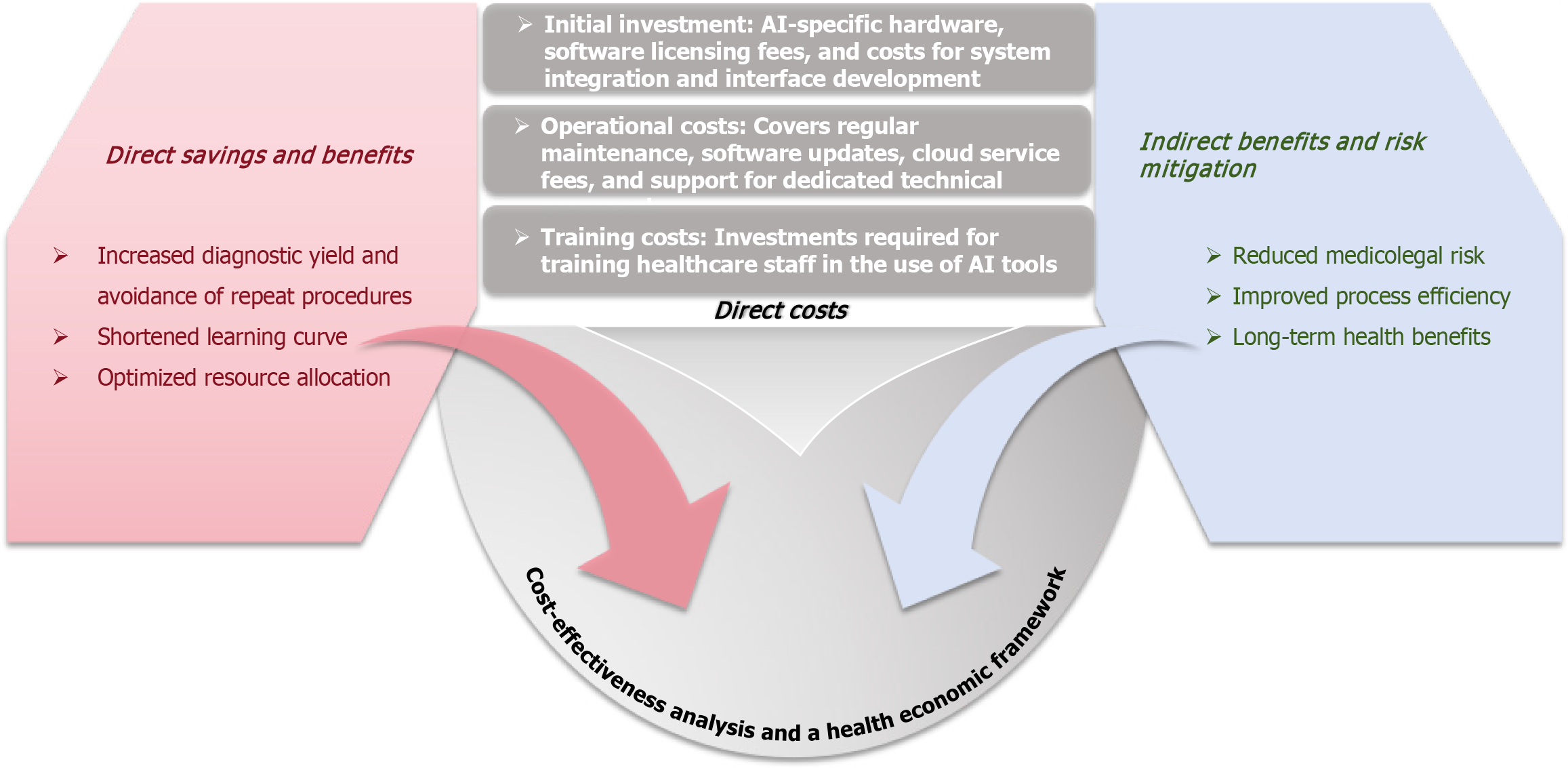
The high initial investment in AI systems (hardware, software licensing, maintenance) and potential operational complexity are major barriers to adoption in resource-limited settings. To strengthen the argument for hospital administrators, a structured health economic framework should be outlined (Figure 3). This framework should encompass not only upfront expenditures but also long-term savings from increased diagnostic yield, enhanced workflow efficiency due to shortened learning curves, and potential reductions in medicolegal risks. Widespread implementation should therefore be supported by clear cost-benefit studies demonstrating the comprehensive healthcare economic value of AI.
Embedding AI tools into existing EUS workflows may alter operational habits and increase initial procedure time. Successful integration requires optimizing human-machine interfaces to ensure that AI-assisted prompts are timely, intuitive, and do not interfere with the primary operational field of view, as well as providing training to reduce resistance to use.
Traditional EUS training involves a long learning period. AI navigation and diagnostic support systems have the potential to serve as standardized training tools, shortening the time required for novices to reach competency (e.g., reducing from over 50 procedures to fewer). However, this must be accompanied by the design and validation of effective AI-assisted training curricula and evaluation systems.
EUS data are characterized by significant heterogeneity arising from variations in probe frequency, inconsistent imaging annotation protocols, and operator-specific techniques. These discrepancies lead to data imbalance, limit model generalizability, and potentially introduce regional biases; for example, models trained primarily on European data may exhibit reduced sensitivity when applied to Asian populations[57]. To address these issues, data standardization efforts should adopt established frameworks like the European Health Data Space. This initiative promotes unified EUS data quality standards, including anonymized image-sharing protocols, to enhance model robustness and minimize bias.
A critical yet often overlooked dimension of EUS is its ability to capture dynamic physiological and mechanical properties - such as tissue stiffness, vascular flow, and contrast kinetics - which are essential for differential diagnosis. Current AI models predominantly rely on static images, thereby losing valuable functional information embedded in sequences like elastography (strain ratio), contrast-enhanced time-intensity curves, and doppler flow signatures[19]. Future standardization protocols must emphasize the inclusion and annotation of dynamic features, along with de
Beyond standardization, multimodal integration combining EUS with computed tomography, magnetic resonance imaging and clinical data represents a promising strategy to improve diagnostic accuracy[58-60]. For instance, Seza et al[61] developed a multimodal AI model incorporating EUS, contrast-enhanced computed tomography, T2-weighted magnetic resonance imaging, and magnetic resonance pancreatography (magnetic resonance pancreatography). This model increased accuracy in classifying pancreatic cystic neoplasms from 91% to 99%, enabling near-perfect discrimination between serous and mucinous tumors. Future research should prioritize optimizing multimodal fusion methods, uncovering biologically relevant correlations, and extending this diagnostic paradigm to other gastrointestinal ma
The adoption of advanced AI systems in EUS, such as Endoangel, remains limited by dependency on specific hardware platforms (e.g., Olympus endoscopes). High implementation costs further restrict accessibility, especially in resource-limited settings[62]. Another challenge is the prolonged learning curve for conventional EUS training, typically requiring over 50 supervised procedures. To improve affordability and scalability, future development should emphasize lightweight, vendor-agnostic AI architectures with reduced hardware requirements and operational expenses. Therefore, to enhance affordability and scalability, future development must prioritize the creation of lightweight, vendor-agnostic AI architectures that systematically reduce hardware reliance and operational expenditures.
In this context, hardware compatibility and integration complexity emerge as critical barriers to widespread im
In addition to cost-efficiency, AI’s fundamental promise lies in its ability to narrow experience-based performance gaps, effectively “elevating non-expert practitioners to specialist-level proficiency” while optimizing resource use through the avoidance of low-yield procedures[62]. Human-AI collaboration has already shown substantial potential. The Joint-AI model introduced by Cui et al[39] significantly improved junior endoscopists’ diagnostic accuracy (69%-90%) and sensitivity (61%-91%) for solid pancreatic lesions. Furthermore, interpretable decision support increased AI acceptance among specialists[39]. Fang et al’s semi-supervised[53] convolutional neural network demonstrated similar potential by reducing annotation dependence by 80% (using only 20% labeled data) while maintaining accuracy above 93.9% in diagnosing pancreatic ductal adenocarcinoma. However, critical challenges persist regarding conflict resolution when AI and clinician judgments diverge. Multidisciplinary consensus frameworks may provide structured mechanisms to reconcile such discrepancies. Further efficiency could be achieved through active learning strategies, where AI pro
Clarifying accountability frameworks: Integrating AI into EUS poses complex ethical challenges, particularly regarding accountability for diagnostic or navigational errors, such as missing critical lesions, where responsibility among clinicians, hospitals, and developers remains unclear. Although systems like EUS-AIRS enhance traceability by documenting procedural details (e.g., spatial relationships between needles and lesions)[19], a systematic framework for accountability has yet to be established.
Transitioning from problem to principle, the establishment of such a framework must be grounded in a clear hierarchy of duties. Clinicians, as the ultimate arbiters of clinical judgment, bear the primary responsibility for decisions made with AI assistance. This necessitates not only operational oversight but also the preserved professional autonomy to critically evaluate and, when necessary, override algorithmic recommendations. Concurrently, healthcare institutions carry an institutional duty to rigorously validate AI tools and ensure their adherence to established clinical standards. Developers, for their part, are obligated to ensure the safety, efficacy, and fundamental transparency of their algorithms, thereby furnishing the trustworthy substrate upon which clinical decisions are made. Ultimately, the crystallization of these distinct yet interdependent obligations requires robust regulatory articulation, as envisaged in instruments like the EU AI Act, which must precisely delineate the scope of liability for each stakeholder[63,64].
Enhancing algorithmic transparency and interpretability: Demystifying the “black-box” nature of AI is essential for building trust. Explainable AI techniques should be promoted to provide visual justification for AI diagnoses (e.g., heatmaps highlighting suspicious areas). Concurrently, adopting “Model Cards” -standardized documentation for each clinical AI model - can disclose details such as training data composition, performance metrics, known limitations, and target populations, analogous to drug package inserts[65,66].
Updating informed consent processes: Traditional consent forms often fail to address the specifics and risks of AI-assisted decision-making. The informed consent process should be revised to explicitly inform patients of AI’s role in their care, explaining its function (assistance rather than replacement), potential benefits (e.g., improved consistency), and uncertainties (e.g., limited judgment in rare cases). In the future, generative AI could assist in creating personalized consent documents and patient education materials, thereby improving communication efficiency and transparency[67-69].
China currently lacks a comprehensive regulatory framework for clinical AI applications. To address this, a multi-tiered oversight system should be established by drawing upon and synthesizing key strengths of international models (Table 2). This system should include: (1) Adoption of a “Predetermined Change Control” pathway, similar to the Food and Drug Administration model, allowing predefined updates (e.g., adjustments in diagnostic thresholds) without full reapproval, analogous to iterative software upgrades[70]; (2) Implementation of risk-stratified oversight aligned with the European Union’s Medical Device Regulation, requiring rigorous clinical validation for high-risk applications (e.g., AI-assisted early cancer screening) while streamlining evaluation for lower-risk tools (e.g., automated reporting systems)[71]; and (3) Continuous performance monitoring mechanisms, following frameworks developed by organizations like the Japan Gastroenterological Endoscopy Society. These mechanisms would mandate systematic documentation of discrepancies between AI recommendations and clinician decisions, alongside periodic reporting of model performance to maintain accountability throughout the AI lifecycle[72].
| Feature | United States (FDA) | European Union (MDR/IVDR + AI Act) |
| Regulatory core | Function-based (what the software does) tiered regulation | Device risk + AI system risk dual matrix regulation. |
| Classification logic | CADt → CADe → CADx → CADa, with increasing risk | Class I, IIa, IIb, III (device risk), plus the AI Act’s “high-risk” category for most medical AI |
| Primary pathways | 510(k) (substantial equivalence), De Novo (novel low-moderate risk), PMA (high risk) | Self-certification (class I low risk), notified body conformity assessment (class IIa and above) |
| Adaptability | Predetermined change control plans allow for iterative updates to cleared algorithms within defined bounds without new submission | Update processes under regulations are stricter; significant software changes typically require re-assessment/notification to the notified body |
| Clinical evidence | Emphasizes prospective, multi-center clinical trials to demonstrate safety and effectiveness | Emphasizes clinical evaluation with comprehensive technical documentation and performance validation, aligned with GDPR |
| Transparency | Requires disclosure of algorithm performance | The AI act emphasizes transparency, requiring high-risk AI systems to provide clear usage information and ensure outputs are interpretable and overseen by humans |
| Process flowcharts | AI medical device concept → determine intended use and function: CADt, CADe, CADx, or CADa → assign FDA risk class → generate clinical and technical evidence → submit to FDA for review → FDA approval clearance → post-market surveillance | Medical AI product → determine device risk class per MDR/IVDR → determine AI system risk level per AI act (typically “high-risk”) → comply with both sets of requirements → undergo conformity Assessment by a notified body (for class IIa and above) → obtain CE marking |
Although the “Prediction-Navigation-Diagnosis” framework provides a clear conceptual structure for AI-enhanced EUS, its clinical translation still faces integration barriers. To facilitate a safe and structured transition from “framework” to “workflow”, we propose a phased, stepwise integration roadmap (Table 3). This roadmap is designed to progressively validate and implement AI tools according to their risk level, ultimately enabling seamless and effective AI integration across the entire EUS workflow.
| Phase | Primary goals | Key evidence required | Regulatory milestones |
| Phase 1 (1-2 years) | Low-risk auxiliary tasks (e.g., automated measurement, structured reporting) | Feasibility studies, human-machine interaction safety data | Registration or simplified clearance (e.g., class II medical device) |
| Phase 2 (3-5 years) | Moderate-risk diagnostic support (e.g., lesion detection and classification) | Multicenter diagnostic accuracy studies, clinical utility assessment | Moderate-risk device approval (requires performance monitoring plan) |
| Phase 3 (> 5 years) | High-risk autonomous functions (e.g., puncture path planning, real-time complication alert) | Large-scale RCTs: Real-world effectiveness data | High-risk device approval (requires a full post-market surveillance system) |
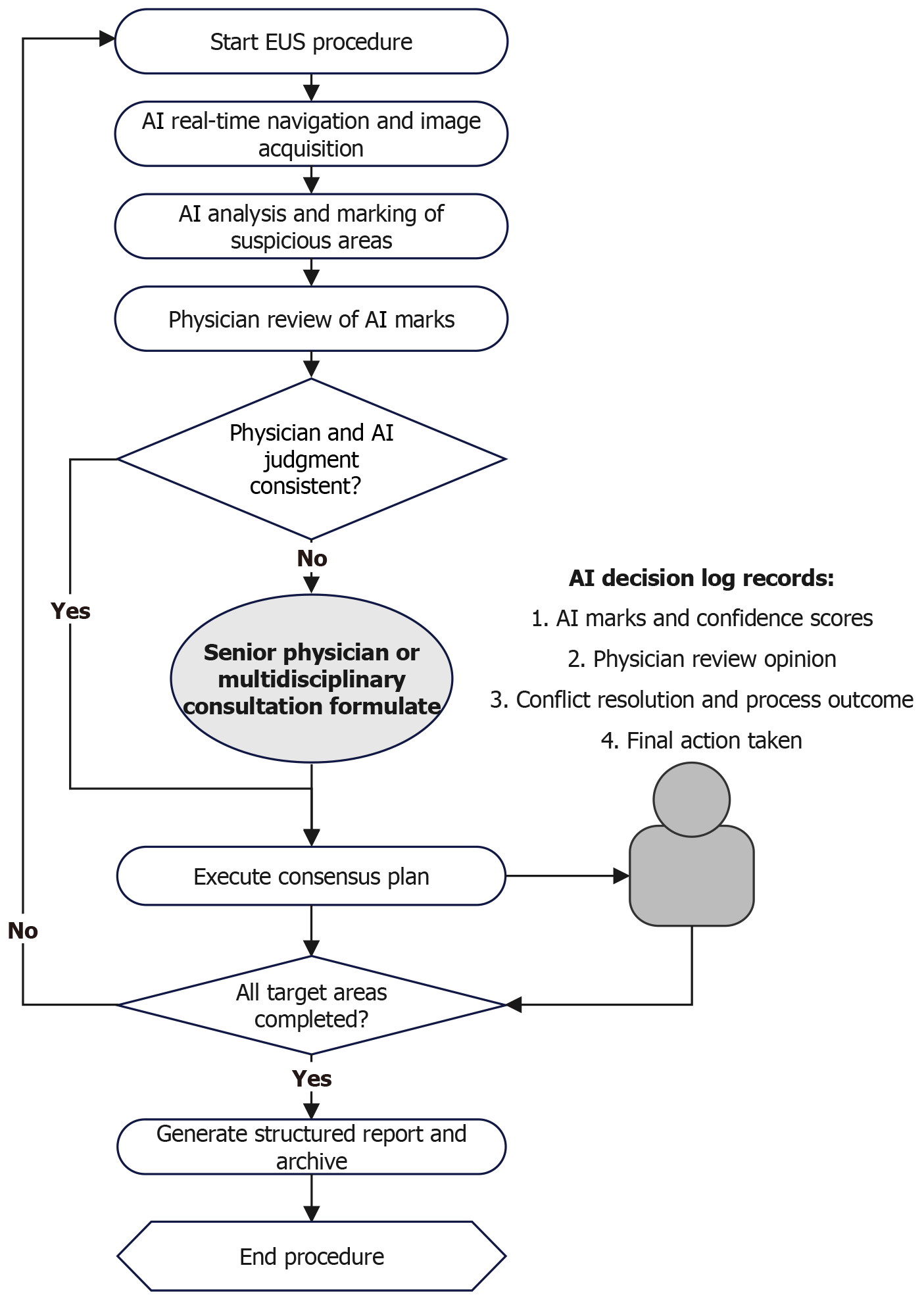
This integrated workflow is operationally outlined in Figure 4. It depicts the closed-loop, AI-augmented EUS pro
Beyond the phased integration pathway, the future intelligence of EUS will also benefit from the convergence of emerging technologies such as multimodal large language models and augmented reality. Multimodal large language models can integrate EUS images, clinical notes, pathology reports, and even genomic data to enable automated, structured reporting and provide differential diagnostic support for complex cases, thereby reducing documentation burden and improving decision consistency. Meanwhile, combining real-time AI detection with augmented reality navigation could overlay key anatomical structures, lesion contours, and planned puncture paths directly into the endoscopist’s visual field, enabling “what-you-see-is-what-you-treat” precision and further shortening the learning curve while enhancing procedural safety. These technological advances are expected to drive EUS toward a fully integrated, highly autonomous intelligent surgical platform in the long term (beyond phase 3).
In conclusion, AI demonstrates significant potential to enhance EUS through a comprehensive “Prediction-Navigation-Diagnosis” framework, addressing key limitations in operator dependency and diagnostic variability. Current applications in preoperative risk stratification, real-time guidance, and diagnostic augmentation show promising improvements in procedural consistency and accuracy. However, widespread clinical translation faces substantial challenges, including data heterogeneity, implementation costs, and unresolved ethical and regulatory considerations. Future success will depend on standardizing multimodal data integration, developing accessible and robust algorithms, establishing clear governance for human-AI collaboration, and fostering adaptive regulatory policies. By systematically addressing these barriers, AI-enhanced EUS can evolve towards a more intelligent, standardized, and accessible paradigm in precision endoscopy.
The authors would like to express their sincere gratitude to the colleagues who provided valuable insights during the conceptual development of this review. We also thank the research assistants for their support in the preliminary literature search and data organization.
| 1. | Sooklal S, Chahal P. Endoscopic Ultrasound. Surg Clin North Am. 2020;100:1133-1150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 38] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 2. | Zhang D, Wu C, Yang Z, Yin H, Liu Y, Li W, Huang H, Jin Z. The application of artificial intelligence in EUS. Endosc Ultrasound. 2024;13:65-75. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 10.5] [Reference Citation Analysis (1)] |
| 3. | Karaca C, Turner BG, Cizginer S, Forcione D, Brugge W. Accuracy of EUS in the evaluation of small gastric subepithelial lesions. Gastrointest Endosc. 2010;71:722-727. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 141] [Article Influence: 8.8] [Reference Citation Analysis (1)] |
| 4. | Lim TW, Choi CW, Kang DH, Kim HW, Park SB, Kim SJ. Endoscopic ultrasound without tissue acquisition has poor accuracy for diagnosing gastric subepithelial tumors. Medicine (Baltimore). 2016;95:e5246. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 37] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 5. | Giovannini M. The place of endoscopic ultrasound in bilio-pancreatic pathology. Gastroenterol Clin Biol. 2010;34:436-445. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 23] [Article Influence: 1.4] [Reference Citation Analysis (3)] |
| 6. | Giljaca V, Gurusamy KS, Takwoingi Y, Higgie D, Poropat G, Štimac D, Davidson BR. Endoscopic ultrasound versus magnetic resonance cholangiopancreatography for common bile duct stones. Cochrane Database Syst Rev. 2015;2015:CD011549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 74] [Article Influence: 6.7] [Reference Citation Analysis (6)] |
| 7. | Ribeiro A, Goel A. The Risk Factors for Acute Pancreatitis after Endoscopic Ultrasound Guided Biopsy. Korean J Gastroenterol. 2018;72:135-140. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 12] [Article Influence: 1.5] [Reference Citation Analysis (1)] |
| 8. | Hamada T, Yasunaga H, Nakai Y, Isayama H, Horiguchi H, Matsuda S, Fushimi K, Koike K. Rarity of severe bleeding and perforation in endoscopic ultrasound-guided fine needle aspiration for submucosal tumors. Dig Dis Sci. 2013;58:2634-2638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 21] [Article Influence: 1.6] [Reference Citation Analysis (1)] |
| 9. | Sugimoto M, Irie H, Takagi T, Suzuki R, Konno N, Asama H, Sato Y, Nakamura J, Takasumi M, Hashimoto M, Kato T, Kobashi R, Kobayashi Y, Hashimoto Y, Hikichi T, Ohira H. Efficacy of EUS-guided FNB using a Franseen needle for tissue acquisition and microsatellite instability evaluation in unresectable pancreatic lesions. BMC Cancer. 2020;20:1094. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 10. | Umans DS, Rangkuti CK, Sperna Weiland CJ, Timmerhuis HC, Bouwense SAW, Fockens P, Besselink MG, Verdonk RC, van Hooft JE; Dutch Pancreatitis Study Group. Endoscopic ultrasonography can detect a cause in the majority of patients with idiopathic acute pancreatitis: a systematic review and meta-analysis. Endoscopy. 2020;52:955-964. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 23] [Article Influence: 3.8] [Reference Citation Analysis (1)] |
| 11. | Sirtl S, Żorniak M, Hohmann E, Beyer G, Dibos M, Wandel A, Phillip V, Ammer-Herrmenau C, Neesse A, Schulz C, Schirra J, Mayerle J, Mahajan UM. Machine learning-based decision tool for selecting patients with idiopathic acute pancreatitis for endosonography to exclude a biliary aetiology. World J Gastroenterol. 2023;29:5138-5153. [PubMed] [DOI] [Full Text] |
| 12. | Lv XH, Wang CH, Xie Y. Efficacy and safety of submucosal tunneling endoscopic resection for upper gastrointestinal submucosal tumors: a systematic review and meta-analysis. Surg Endosc. 2017;31:49-63. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 86] [Article Influence: 9.6] [Reference Citation Analysis (1)] |
| 13. | Deprez PH, Moons LMG, OʼToole D, Gincul R, Seicean A, Pimentel-Nunes P, Fernández-Esparrach G, Polkowski M, Vieth M, Borbath I, Moreels TG, Nieveen van Dijkum E, Blay JY, van Hooft JE. Endoscopic management of subepithelial lesions including neuroendocrine neoplasms: European Society of Gastrointestinal Endoscopy (ESGE) Guideline. Endoscopy. 2022;54:412-429. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 325] [Cited by in RCA: 260] [Article Influence: 65.0] [Reference Citation Analysis (2)] |
| 14. | Chen X, Zhou J, Wang P, Wang P, Wang L, Mu L, Lang C, Mu Y, Wang X, Shang R, Li Q, Lv H, Wu K, Shi N, Jia X, Lai Y, Zhang Y, Li Z, Zhong N. Endoscopic ultrasound-based application system for predicting endoscopic resection-related outcomes and diagnosing subepithelial lesions: Multicenter prospective study. Dig Endosc. 2024;36:141-151. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (1)] |
| 15. | Singhi AD, Koay EJ, Chari ST, Maitra A. Early Detection of Pancreatic Cancer: Opportunities and Challenges. Gastroenterology. 2019;156:2024-2040. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 691] [Cited by in RCA: 610] [Article Influence: 87.1] [Reference Citation Analysis (5)] |
| 16. | Meeralam Y, Al-Shammari K, Yaghoobi M. Diagnostic accuracy of EUS compared with MRCP in detecting choledocholithiasis: a meta-analysis of diagnostic test accuracy in head-to-head studies. Gastrointest Endosc. 2017;86:986-993. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 109] [Article Influence: 12.1] [Reference Citation Analysis (2)] |
| 17. | Han C, Nie C, Shen X, Xu T, Liu J, Ding Z, Hou X. Exploration of an effective training system for the diagnosis of pancreatobiliary diseases with EUS: A prospective study. Endosc Ultrasound. 2020;9:308-318. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 1.2] [Reference Citation Analysis (1)] |
| 18. | Wu HL, Yao LW, Shi HY, Wu LL, Li X, Zhang CX, Chen BR, Zhang J, Tan W, Cui N, Zhou W, Zhang JX, Xiao B, Gong RR, Ding Z, Yu HG. Validation of a real-time biliopancreatic endoscopic ultrasonography analytical device in China: a prospective, single-centre, randomised, controlled trial. Lancet Digit Health. 2023;5:e812-e820. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (1)] |
| 19. | Li X, Yao L, Wu H, Tan W, Zhou W, Zhang J, Dong Z, Ding X, Yu H. A deep learning-based, real-time image report system for linear EUS. Gastrointest Endosc. 2025;101:1166-1173.e11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 20. | Rizzatti G, Tripodi G, De Lucia SS, Pellegrino A, Boskoski I, Larghi A, Spada C. Artificial intelligence system for EUS navigation and anatomical landmark recognition. VideoGIE. 2025;10:358-363. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (1)] |
| 21. | Hedenbro JL, Ekelund M, Wetterberg P. Endoscopic diagnosis of submucosal gastric lesions. The results after routine endoscopy. Surg Endosc. 1991;5:20-23. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 196] [Cited by in RCA: 214] [Article Influence: 6.1] [Reference Citation Analysis (4)] |
| 22. | Park CH, Kim EH, Jung DH, Chung H, Park JC, Shin SK, Lee YC, Kim H, Lee SK. Impact of periodic endoscopy on incidentally diagnosed gastric gastrointestinal stromal tumors: findings in surgically resected and confirmed lesions. Ann Surg Oncol. 2015;22:2933-2939. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 25] [Article Influence: 2.3] [Reference Citation Analysis (4)] |
| 23. | Hirai K, Kuwahara T, Furukawa K, Kakushima N, Furune S, Yamamoto H, Marukawa T, Asai H, Matsui K, Sasaki Y, Sakai D, Yamada K, Nishikawa T, Hayashi D, Obayashi T, Komiyama T, Ishikawa E, Sawada T, Maeda K, Yamamura T, Ishikawa T, Ohno E, Nakamura M, Kawashima H, Ishigami M, Fujishiro M. Artificial intelligence-based diagnosis of upper gastrointestinal subepithelial lesions on endoscopic ultrasonography images. Gastric Cancer. 2022;25:382-391. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 60] [Article Influence: 15.0] [Reference Citation Analysis (1)] |
| 24. | Joo DC, Kim GH, Lee MW, Lee BE, Kim JW, Kim KB. Artificial Intelligence-Based Diagnosis of Gastric Mesenchymal Tumors Using Digital Endosonography Image Analysis. J Clin Med. 2024;13:3725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (2)] |
| 25. | Domper Arnal MJ, Ferrández Arenas Á, Lanas Arbeloa Á. Esophageal cancer: Risk factors, screening and endoscopic treatment in Western and Eastern countries. World J Gastroenterol. 2015;21:7933-7943. [PubMed] [DOI] [Full Text] |
| 26. | Zhang Y. Epidemiology of esophageal cancer. World J Gastroenterol. 2013;19:5598-5606. [PubMed] [DOI] [Full Text] |
| 27. | Lin Y, Totsuka Y, He Y, Kikuchi S, Qiao Y, Ueda J, Wei W, Inoue M, Tanaka H. Epidemiology of esophageal cancer in Japan and China. J Epidemiol. 2013;23:233-242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 400] [Cited by in RCA: 439] [Article Influence: 33.8] [Reference Citation Analysis (3)] |
| 28. | Rana SS, Sharma R, Gupta R. High-frequency miniprobe endoscopic ultrasonography for evaluation of indeterminate esophageal strictures. Ann Gastroenterol. 2018;31:680-684. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 29. | Xu GQ, Qian JJ, Chen MH, Ren GP, Chen HT. Endoscopic ultrasonography for the diagnosis and selecting treatment of esophageal leiomyoma. J Gastroenterol Hepatol. 2012;27:521-525. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 17] [Article Influence: 1.2] [Reference Citation Analysis (1)] |
| 30. | Sun LJ, Chen X, Dai YN, Xu CF, Ji F, Chen LH, Chen HT, Chen CX. Endoscopic Ultrasonography in the Diagnosis and Treatment Strategy Choice of Esophageal Leiomyoma. Clinics (Sao Paulo). 2017;72:197-201. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 12] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 31. | Mortensen MB. Combined pretherapeutic endoscopic and laparoscopic ultrasonography may predict survival of patients with upper gastrointestinal tract cancer. Surg Endosc. 2012;26:891-892. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 32. | Liu GS, Huang PY, Wen ML, Zhuang SS, Hua J, He XP. Application of endoscopic ultrasonography for detecting esophageal lesions based on convolutional neural network. World J Gastroenterol. 2022;28:2457-2467. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 6] [Reference Citation Analysis (2)] |
| 33. | Zhang AM, Jiang DM, Wang SP, Liu W, Sun BB, Wang Z, Zhou GY, Wu YF, Cai QY, Guo JT, Sun SY. Artificial intelligence-assisted endoscopic ultrasound diagnosis of esophageal subepithelial lesions. Surg Endosc. 2025;39:3821-3831. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 34. | Vincent A, Herman J, Schulick R, Hruban RH, Goggins M. Pancreatic cancer. Lancet. 2011;378:607-620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2265] [Cited by in RCA: 2176] [Article Influence: 145.1] [Reference Citation Analysis (5)] |
| 35. | Kitano M, Yoshida T, Itonaga M, Tamura T, Hatamaru K, Yamashita Y. Impact of endoscopic ultrasonography on diagnosis of pancreatic cancer. J Gastroenterol. 2019;54:19-32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 265] [Cited by in RCA: 224] [Article Influence: 32.0] [Reference Citation Analysis (8)] |
| 36. | Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer Statistics, 2021. CA Cancer J Clin. 2021;71:7-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12117] [Cited by in RCA: 12194] [Article Influence: 2438.8] [Reference Citation Analysis (7)] |
| 37. | Santo E, Bar-Yishay I. Pancreatic solid incidentalomas. Endosc Ultrasound. 2017;6:S99-S103. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 21] [Article Influence: 2.3] [Reference Citation Analysis (1)] |
| 38. | Teshima CW, Sandha GS. Endoscopic ultrasound in the diagnosis and treatment of pancreatic disease. World J Gastroenterol. 2014;20:9976-9989. [PubMed] [DOI] [Full Text] |
| 39. | Cui H, Zhao Y, Xiong S, Feng Y, Li P, Lv Y, Chen Q, Wang R, Xie P, Luo Z, Cheng S, Wang W, Li X, Xiong D, Cao X, Bai S, Yang A, Cheng B. Diagnosing Solid Lesions in the Pancreas With Multimodal Artificial Intelligence: A Randomized Crossover Trial. JAMA Netw Open. 2024;7:e2422454. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 31] [Article Influence: 15.5] [Reference Citation Analysis (1)] |
| 40. | Krampitz GW, Aloia TA. Staging of Biliary and Primary Liver Tumors: Current Recommendations and Workup. Surg Oncol Clin N Am. 2019;28:663-683. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 10] [Article Influence: 1.4] [Reference Citation Analysis (1)] |
| 41. | Razumilava N, Gores GJ. Cholangiocarcinoma. Lancet. 2014;383:2168-2179. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1509] [Cited by in RCA: 1477] [Article Influence: 123.1] [Reference Citation Analysis (6)] |
| 42. | Orzan RI, Santa D, Lorenzovici N, Zareczky TA, Pojoga C, Agoston R, Dulf EH, Seicean A. Deep Learning in Endoscopic Ultrasound: A Breakthrough in Detecting Distal Cholangiocarcinoma. Cancers (Basel). 2024;16:3792. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 43. | Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76817] [Cited by in RCA: 69714] [Article Influence: 13942.8] [Reference Citation Analysis (46)] |
| 44. | Kou S, Thakur S, Eltahir A, Nie H, Zhang Y, Song A, Hunt SR, Mutch MG, Chapman WC Jr, Zhu Q. A portable photoacoustic microscopy and ultrasound system for rectal cancer imaging. Photoacoustics. 2024;39:100640. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 10] [Reference Citation Analysis (1)] |
| 45. | Chao G, Ye F, Li T, Gong W, Zhang S. Estimation of invasion depth of early colorectal cancer using EUS and NBI-ME: a meta-analysis. Tech Coloproctol. 2019;23:821-830. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 11] [Article Influence: 1.6] [Reference Citation Analysis (1)] |
| 46. | Men H, Yan C, Peng X, Jin SQ, Du YH, Tang ZS, Li H, Ou-Yang T, Zhang S, Ding LS, Deng J, Xu Z, Li GB, Luo HY, Li Z, Xie F, Han S. Validation of multiple deep learning models for colorectal tumor differentiation with endoscopic ultrasound images: a dual-center study. J Gastrointest Oncol. 2025;16:435-452. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (5)] |
| 47. | Facciorusso A, Arvanitakis M, Crinò SF, Fabbri C, Fornelli A, Leeds J, Archibugi L, Carrara S, Dhar J, Gkolfakis P, Haugk B, Iglesias Garcia J, Napoleon B, Papanikolaou IS, Seicean A, Stassen PMC, Vilmann P, Tham TC, Fuccio L. Endoscopic ultrasound-guided tissue sampling: European Society of Gastrointestinal Endoscopy (ESGE) Technical and Technology Review. Endoscopy. 2025;57:390-418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 39] [Article Influence: 39.0] [Reference Citation Analysis (1)] |
| 48. | Yang F, Liu E, Sun S. Rapid on-site evaluation (ROSE) with EUS-FNA: The ROSE looks beautiful. Endosc Ultrasound. 2019;8:283-287. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 29] [Article Influence: 4.1] [Reference Citation Analysis (1)] |
| 49. | Iglesias-Garcia J, Lariño-Noia J, Abdulkader I, Domínguez-Muñoz JE. Rapid on-site evaluation of endoscopic-ultrasound-guided fine-needle aspiration diagnosis of pancreatic masses. World J Gastroenterol. 2014;20:9451-9457. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 43] [Cited by in RCA: 63] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 50. | Schmidt RL, Walker BS, Howard K, Layfield LJ, Adler DG. Rapid on-site evaluation reduces needle passes in endoscopic ultrasound-guided fine-needle aspiration for solid pancreatic lesions: a risk-benefit analysis. Dig Dis Sci. 2013;58:3280-3286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 36] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 51. | Matynia AP, Schmidt RL, Barraza G, Layfield LJ, Siddiqui AA, Adler DG. Impact of rapid on-site evaluation on the adequacy of endoscopic-ultrasound guided fine-needle aspiration of solid pancreatic lesions: a systematic review and meta-analysis. J Gastroenterol Hepatol. 2014;29:697-705. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 102] [Article Influence: 8.5] [Reference Citation Analysis (1)] |
| 52. | Fujii Y, Uchida D, Sato R, Obata T, Akihiro M, Miyamoto K, Morimoto K, Terasawa H, Yamazaki T, Matsumoto K, Horiguchi S, Tsutsumi K, Kato H, Inoue H, Cho T, Tanimoto T, Ohto A, Kawahara Y, Otsuka M. Effectiveness of data-augmentation on deep learning in evaluating rapid on-site cytopathology at endoscopic ultrasound-guided fine needle aspiration. Sci Rep. 2024;14:22441. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 16] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 53. | Fang D, Huang Y, Li S, Shi C, Bao J, Du D, Xuan L, Ye L, Zhang Y, Zhu C, Zheng H, Shi Z, Mei Q, Wang H. A semi-supervised convolutional neural network for diagnosis of pancreatic ductal adenocarcinoma based on EUS-FNA cytological images. BMC Cancer. 2025;25:495. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 5] [Reference Citation Analysis (1)] |
| 54. | Adler DG, Jacobson BC, Davila RE, Hirota WK, Leighton JA, Qureshi WA, Rajan E, Zuckerman MJ, Fanelli RD, Baron TH, Faigel DO; ASGE. ASGE guideline: complications of EUS. Gastrointest Endosc. 2005;61:8-12. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 159] [Cited by in RCA: 145] [Article Influence: 6.9] [Reference Citation Analysis (1)] |
| 55. | Polkowski M, Larghi A, Weynand B, Boustière C, Giovannini M, Pujol B, Dumonceau JM; European Society of Gastrointestinal Endoscopy (ESGE). Learning, techniques, and complications of endoscopic ultrasound (EUS)-guided sampling in gastroenterology: European Society of Gastrointestinal Endoscopy (ESGE) Technical Guideline. Endoscopy. 2012;44:190-206. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 229] [Cited by in RCA: 210] [Article Influence: 15.0] [Reference Citation Analysis (2)] |
| 56. | Brenner T, Kuo A, Sperna Weiland CJ, Kamal A, Elmunzer BJ, Luo H, Buxbaum J, Gardner TB, Mok SS, Fogel ES, Phillip V, Choi JH, Lua GW, Lin CC, Reddy DN, Lakhtakia S, Goenka MK, Kochhar R, Khashab MA, van Geenen EJM, Singh VK, Tomasetti C, Akshintala VS. Development and validation of a machine learning-based, point-of-care risk calculator for post-ERCP pancreatitis and prophylaxis selection. Gastrointest Endosc. 2025;101:129-138.e0. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 18] [Article Influence: 18.0] [Reference Citation Analysis (1)] |
| 57. | Dove ES. The European Health Data Space as a Case Study. Ethics Hum Res. 2024;46:29-35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 58. | Lu X, Zhang S, Ma C, Peng C, Lv Y, Zou X. The diagnostic value of EUS in pancreatic cystic neoplasms compared with CT and MRI. Endosc Ultrasound. 2015;4:324-329. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 44] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 59. | Tirkes T, Patel AA, Tahir B, Kim RC, Schmidt CM, Akisik FM. Pancreatic cystic neoplasms and post-inflammatory cysts: interobserver agreement and diagnostic performance of MRI with MRCP. Abdom Radiol (NY). 2021;46:4245-4253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 60. | Visser BC, Yeh BM, Qayyum A, Way LW, McCulloch CE, Coakley FV. Characterization of cystic pancreatic masses: relative accuracy of CT and MRI. AJR Am J Roentgenol. 2007;189:648-656. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 122] [Article Influence: 6.4] [Reference Citation Analysis (1)] |
| 61. | Seza K, Tawada K, Kobayashi A, Nakamura K. Multimodal Artificial Intelligence Using Endoscopic USG, CT, and MRI to Differentiate Between Serous and Mucinous Cystic Neoplasms. Cureus. 2025;17:e85547. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (1)] |
| 62. | Messmann H, Bisschops R, Antonelli G, Libânio D, Sinonquel P, Abdelrahim M, Ahmad OF, Areia M, Bergman JJGHM, Bhandari P, Boskoski I, Dekker E, Domagk D, Ebigbo A, Eelbode T, Eliakim R, Häfner M, Haidry RJ, Jover R, Kaminski MF, Kuvaev R, Mori Y, Palazzo M, Repici A, Rondonotti E, Rutter MD, Saito Y, Sharma P, Spada C, Spadaccini M, Veitch A, Gralnek IM, Hassan C, Dinis-Ribeiro M. Expected value of artificial intelligence in gastrointestinal endoscopy: European Society of Gastrointestinal Endoscopy (ESGE) Position Statement. Endoscopy. 2022;54:1211-1231. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 153] [Cited by in RCA: 119] [Article Influence: 29.8] [Reference Citation Analysis (2)] |
| 63. | Kalodanis K, Feretzakis G, Rizomiliotis P, Verykios VS, Papapavlou C, Skrekas A, Anagnostopoulos D. Evaluating the Impact of the EU AI Act on Medical Device Regulation. Stud Health Technol Inform. 2025;323:40-44. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 64. | Kotter E, D'Antonoli TA, Cuocolo R, Hierath M, Huisman M, Klontzas ME, Martí-Bonmatí L, May MS, Neri E, Nikolaou K, Pinto Dos Santos D, Radzina M, Shelmerdine SC, Bellemo A; European Society of Radiology (ESR). Guiding AI in radiology: ESR's recommendations for effective implementation of the European AI Act. Insights Imaging. 2025;16:33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 53] [Reference Citation Analysis (1)] |
| 65. | Wilkinson MD, Dumontier M, Aalbersberg IJ, Appleton G, Axton M, Baak A, Blomberg N, Boiten JW, da Silva Santos LB, Bourne PE, Bouwman J, Brookes AJ, Clark T, Crosas M, Dillo I, Dumon O, Edmunds S, Evelo CT, Finkers R, Gonzalez-Beltran A, Gray AJ, Groth P, Goble C, Grethe JS, Heringa J, 't Hoen PA, Hooft R, Kuhn T, Kok R, Kok J, Lusher SJ, Martone ME, Mons A, Packer AL, Persson B, Rocca-Serra P, Roos M, van Schaik R, Sansone SA, Schultes E, Sengstag T, Slater T, Strawn G, Swertz MA, Thompson M, van der Lei J, van Mulligen E, Velterop J, Waagmeester A, Wittenburg P, Wolstencroft K, Zhao J, Mons B. The FAIR Guiding Principles for scientific data management and stewardship. Sci Data. 2016;3:160018. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13458] [Cited by in RCA: 6651] [Article Influence: 665.1] [Reference Citation Analysis (5)] |
| 66. | Amith MT, Cui L, Roberts K, Tao C. Application of an ontology for model cards to generate computable artifacts for linking machine learning information from biomedical research. Proc Int World Wide Web Conf. 2023;2023:820-825. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.7] [Reference Citation Analysis (1)] |
| 67. | Azevedo CB, Martinho AS, Braga I, Nogueira-Silva C, Barroso C, Correia-Pinto J. ChatGPT-4o in Enhancing Informed Consent in Pediatric Surgical Practice. J Pediatr Surg. 2025;60:162413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (1)] |
| 68. | Kerbage A, Kassab J, El Dahdah J, Burke CA, Achkar JP, Rouphael C. Accuracy of ChatGPT in Common Gastrointestinal Diseases: Impact for Patients and Providers. Clin Gastroenterol Hepatol. 2024;22:1323-1325.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 45] [Article Influence: 22.5] [Reference Citation Analysis (1)] |
| 69. | Everett SM, Triantafyllou K, Hassan C, Mergener K, Tham TC, Almeida N, Antonelli G, Axon A, Bisschops R, Bretthauer M, Costil V, Foroutan F, Gauci J, Hritz I, Messmann H, Pellisé M, Roelandt P, Seicean A, Tziatzios G, Voiosu A, Gralnek IM. Informed consent for endoscopic procedures: European Society of Gastrointestinal Endoscopy (ESGE) Position Statement. Endoscopy. 2023;55:952-966. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 70. | Yearley AG, Goedmakers CMW, Panahi A, Doucette J, Rana A, Ranganathan K, Smith TR. FDA-approved machine learning algorithms in neuroradiology: A systematic review of the current evidence for approval. Artif Intell Med. 2023;143:102607. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (1)] |
| 71. | Santra S, Kukreja P, Saxena K, Gandhi S, Singh OV. Navigating regulatory and policy challenges for AI enabled combination devices. Front Med Technol. 2024;6:1473350. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 17] [Reference Citation Analysis (1)] |
| 72. | Mori Y, Ishihara R, Ogata H, Kutsumi H, Saito Y, Sumiyama K, Sekiguchi M, Tajiri H, Fujishiro M, Matsuda K, Yano T, Aoki R, Ishiyama M, Imagawa A, Omae M, Oda Y, Kato M, Sakamoto T, Sasabe M, Shiotani A, Suzuki S, Tamai N, Hikichi T, Hirasawa T, Makiguchi M, Misawa M, Yabuuchi Y, Yamaguchi D, Yamada M, Igarashi Y, Tanaka S. Artificial Intelligence in Gastrointestinal Endoscopy: The Japan Gastroenterological Endoscopy Society Position Statements. Dig Endosc. 2025;37:1116-1122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (1)] |