Published online Apr 28, 2026. doi: 10.3748/wjg.v32.i16.115969
Revised: December 15, 2025
Accepted: January 26, 2026
Published online: April 28, 2026
Processing time: 169 Days and 16.6 Hours
Aging is the primary risk factor for numerous chronic diseases, cognitive deterio
To study the gut microbiota profiles of age-related changes during aging in both human and rat cohorts, with particular focus on microbiota involved in lipid me
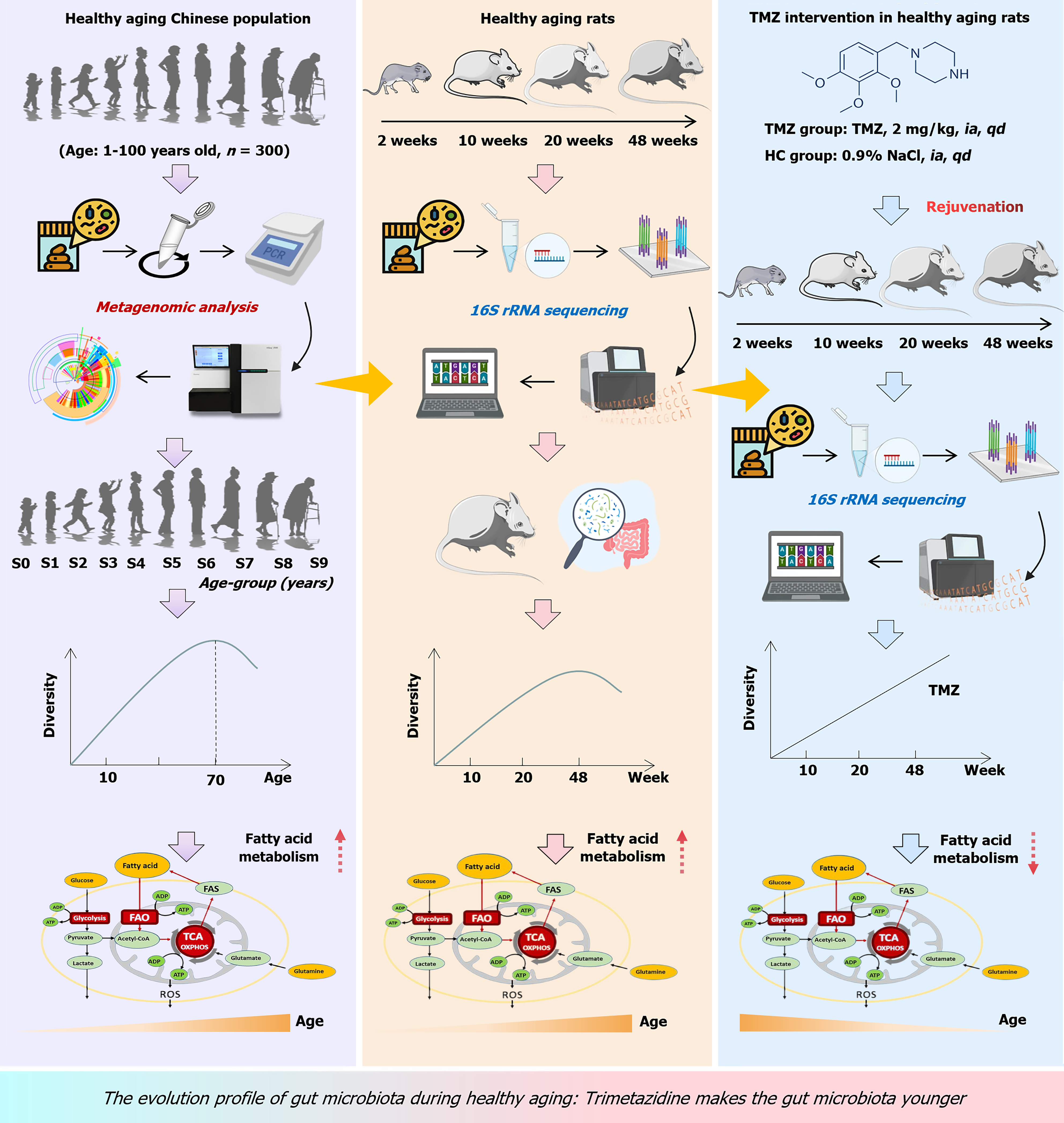
The metagenomic sequencing and 16S rRNA sequencing were performed to detect gut microbiota in 300 individuals of different age groups and rats of different weeks, mapping the evolutionary profiles of microbiota during aging across all age groups and focusing on analyzing significantly changed gut microbiota and their biological functions. Based on clues obtained from the above analysis that microbiota with fatty acid oxidation function was closely related to aging, and validation was conducted using fatty acid oxidation inhibitor TMZ in aging animal models.
Multicohort analysis revealed that gut microbial diversity follows a nonlinear trajectory, initially increasing until age 10, remaining relatively stable until approximately 70 years and declining thereafter. Functional enrichment analysis demonstrated a significant, age-associated increase in lipid metabolism pathways in both human and animal models, which was consistent with marked changes in abundance in the gut microbiota involved in lipid metabolism such as Luteipulveratus (P < 0.05). TMZ, a fatty acid oxidation inhibitor, reshaped the gut microbiota structure and suppressed the abundance of lipid metabolism-associated gut microbiota in aging rats. Further molecular validation confirmed that the TMZ inhibited fatty acid β-oxidation and significantly downregulated the expression levels of key senescence marker proteins and genes.
In conclusion, gut microbiota undergoes age-dependent remodeling, with significant enrichment of the fatty acid oxidation-related microbiome. The fatty acid oxidation inhibitor TMZ may attenuate aging phenotype through the dual modulation of gut microbial composition and lipid metabolism and may provide an antiaging strategy.
Core Tip: This study reveals that gut microbiota alterations during aging follow a nonlinear trajectory across human and rat cohorts. The key finding demonstrates increased fatty acid oxidation-related microbiota in aged individuals. Functional enrichment analysis revealed a possible association between the gut microbiota and lipid metabolism. Trimetazidine (TMZ), a fatty acid oxidation inhibitor, successfully modulated aging-associated gut microbiota and suppressed senescence markers P21 in rats. These findings establish fatty acid oxidation maybe as a crucial mechanistic link between gut microbiota and aging, positioning TMZ as a potential therapeutic intervention for healthy aging through microbiome regulation.
- Citation: Wang XM, Rao BC, Su GY, Wang HY, Zhang GZ, Ge FL, Yu ZJ, Ren ZG, Liang HX. Age-related gut microbiome profile and reversal of microbial imbalance by trimetazidine intervention. World J Gastroenterol 2026; 32(16): 115969
- URL: https://www.wjgnet.com/1007-9327/full/v32/i16/115969.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i16.115969
Aging is the primary risk factor for numerous chronic diseases, cognitive deterioration, and mortality. With the accelerating aging of the population, age-related disorders pose substantial socioeconomic burdens globally. Healthy aging has become a crucial public health objective[1]. To date, this imperative has spurred intensive research into biomarkers of aging and effective interventions to extend a healthy lifespan.
The interplay between the microbiota and host metabolism plays an important role in health. Growing evidence suggests that gut microbiota dysbiosis is a key driver of age-related physiological decline[2]. The dynamic interplay between gut microbial communities and host aging processes may offer a promising novel therapeutic strategy for delaying senescence and treating age-related diseases[3]. Notably, the composition of the gut microbiota significantly altered in an age-dependent manner[4,5]. According to previous preclinical studies, the administration of Lactobacillus plantarum GKM3 can extend lifespan and attenuate age-related cognitive decline in accelerated aging SAMP8 mice[6]. Moreover, the nematode Caenorhabditis elegans was found to inhibit bacterial folate synthesis and modulate bacterial metabolism in vivo, thus extending host lifespan[7]. These findings collectively establish the gut microbiome as a regulator of age-related pathophysiology[8]. Therefore, elucidating the characteristics of the microbiota during aging may provide a strategy for developing early diagnostic markers and establishing targeted interventions.
Emerging evidence indicates that lipid metabolism, which is closely related to the gut microbiota, plays a critical role in the aging process. As main components of lipids, fatty acids undergo significant age-associated alterations[9]. A cohort study on aging in Italy demonstrated that lipid metabolism and intestinal microbiota were involved in pivotal regulatory processes related to in human longevity[10]. Moreover, mitochondrial fatty acid oxidation (FAO) drives senescence[11]. Oxylipins, which are oxidized metabolites of polyunsaturated fatty acids, are increased in senescent cells, among which dihomo-15-deoxy-Δ-12,14-prostaglandin J2 can be used as a biomarker to assess the antiaging efficacy[11]. Previous animal studies have shown that activation of the UGT8 a-mediated glycolipid synthesis pathway in male mouse kidneys associated with inflammation and age-related kidney diseases[12]. Mechanistically, the aging process driven by mitochondrial FAO may be attributed to the attenuated protective effect of omega-3 fatty acids in preserving telomerase activity[11,13]. Trimetazidine (TMZ) is a well-established FAO inhibitor that functions by blocking FAO, thus shifting cellular metabolism toward aerobic glucose oxidation and improving adenosine triphosphate synthesis efficiency, ultimately exerting cytoprotective effects through the attenuation of oxidative stress and inflammation[14]. Consequently, the modulation of lipid metabolism-associated gut microbiota may offer a promising intervention strategy for promoting healthy aging.
In this study, we mapped the age-related alterations in the gut microbiota during aging in human cross-sectional study and rat longitudinal cohorts, focusing specifically on the microbiota involved in lipid metabolism. The FAO inhibitor TMZ was subsequently used to intervene in the aging process. These findings may facilitate the development of microbiota-based early warning biomarkers and intervention strategies, thus establishing a scientific foundation for promoting healthy aging in humans.
This study was designed according to principles of retrospective blinded evaluation and prospective sample collection and was carried out following the guidelines of the Rules of Good Clinical Practice and Helsinki Declaration[15,16]. The Research and Clinical Trial Ethics Review Committee of the First Affiliated Hospital of Zhengzhou University approved this research protocol (Approval No. 2021-KY-0716-003). The rats used in our study were housed under specific pathogen-free conditions. These animal studies were approved by the Animal Welfare and Ethics Committee of the Laboratory Animal Center of Zhengzhou University, No. ZZU-LAC2020101304].
Fecal samples from the healthy natural aging population (hereinafter referred to as the HP group) were obtained from the Chinese Healthy Integrative-Omics Project at the First Affiliated Hospital of Zhengzhou University (n = 300) using the fecal collection kit. The inclusion and exclusion criteria are presented in the Supplementary material. Sex, age, medical history and clinical laboratory test results were obtained from the hospital medical records system. Samples were categorized into ten subgroups on the basis of age. Fecal samples were processed and subjected to metagenomic sequencing analysis to determine human gut microbial profiles during the aging process (Figure 1).
Male specific-pathogen free male Sprague-Dawley rats aged one week were purchased from Beijing Huafukang Biotechnology Co., Ltd. [Production License No. SYXK (Beijing) 2019-0030, Beijing, China]. The rats were randomly divided into two cohorts: (1) In the healthy rat cohort (hereinafter referred to as the HR cohort, n = 48), fecal samples were collected at 2, 10, 20, and 48 weeks (named HR2, HR10, HR20, and HR48, respectively) and subjected to 16S rRNA gene sequencing analysis; and (2) The FAO inhibitor TMZ was utilized to treat natural aging rats (hereafter referred to as the TMZ cohort). In the TMZ cohort, healthy control (HC) groups were given 0.5 mL of saline daily (n = 12), and the TMZ group was administered 2 mg/kg TMZ (formula: C14H22N2O3; No. sc-357801A; Santa Cruz, United States) daily by intragastric administration (n = 18). Feces were collected at different life points (2, 10, 20 and 48 weeks), and then processed for 16S rRNA amplicon sequencing. At the study endpoints, the rats were euthanized under isoflurane (R510-22; Revoard, China) inhalation anesthesia, and the tissues (heart, liver, spleen, kidneys, and lungs), in accordance with American Veterinary Medical Association Guidelines for the Euthanasia of Animals (2020). A flowchart of the research design is shown in Figure 1. Detailed weight measurements for each experimental group are included in the Supplementary material. All animal experiments were performed following the ARRIVE guidelines to ensure transparent and comprehensive reporting of our in vivo studies. The specific location of this animal experiment was the Experimental Animal Center of the First Affiliated Hospital of Zhengzhou University.
All the samples were subjected to the same procedures for DNA extraction and polymerase chain reaction (PCR) amplification by the same laboratory staff[17]. The specific details are included in the Supplementary material.
Raw reads were processed using USEARCH (v11.0.667). Quality-filtered sequences were clustered into unique sequences and sorted in order of decreasing abundance to identify representative sequences using USEARCH according to the UPARSE operational taxonomic units (OTUs) analysis pipeline, and singletons were omitted in this step. OTUs were classified on the basis of 97% similarity after chimeric sequences were removed using UPARSE (version 7.1; http://drive5.com/uparse/) and were annotated using the SILVA reference database (SSU138) in qiime2-2020.11. Alpha diversity metrics (ACE estimator, Chao 1 estimator, Shannon-Wiener diversity index and Simpson diversity index) were assessed by using Mothur v1.42.1. The nonparametric Mann-Whitney U test (R 3.6.0 package stats) was used to test for significant differences between two groups. Comparisons of multiple groups were performed using a nonparametric Kruskal-Wallis test. False discovery rate correction was applied. Bray-Curtis and weighted and unweighted UniFrac dissimilarity were calculated in QIIME (v1.9.1). Principal coordinate analysis (PCoA) plots and permutation multivariate analysis of variance, which were used to test for statistical significance between the groups using 10000 permutations, were generated in the R (version 3.6.0) package vegan 2.5-7. Linear discriminant analysis (LDA) effect size (LEfSe) was used to detect taxa with differential abundance among groups (lefse 1.1, https://github.com/SegataLab/Lefse). PICRUSt2 v2.4.1 (https://github.com/picrust/picrust2/wiki) was used to predict functional abundances on the basis of 16S rRNA gene sequences[18] (Supplementary material). Raw sequencing data and accompanying information are available in the Sequence Read Archive database under accession number PRJNA1285086.
Genomic DNA was extracted from each sample by using E.Z.N.A. Stool DNA Kit (Omega Bio-tek, Inc., GA, United States) and then fragmented (approximately 350 bp, SCIENTZ08-III). The DNA library was constructed according to the instructions of the Hieff next-generation sequencing Ultima Pro DNA Library Prep Kit for Illumina V2 (Yeasen Biotechnology Co., Ltd., Shanghai, China), bridge PCR and sequencing were performed on the Nova Seq 6000 platform (Illumina Inc., United States).
Quality control on the raw reads and obtain high-quality reads for each sample. The reads were assembled, genes were predicted using Megahit and Prodigal, a nonredundant gene set was constructed using CD-HIT and Salmon software, and the gene abundance in each sample was calculated. The nonredundant gene set was blasted against kraken2, such as the CARD and VFDB databases, to obtain taxonomic and functional annotations, and the Clusters of Orthologous Groups (COG), Kyoto Encyclopedia of Genes and Genomes (KEGG) Ortholog (KO), pathway, module and Carbohydrate-Active enzymes (CAZy), information corresponding to the genes was obtained.
Alpha diversity was used to analyze the richness and diversity (ACE index, Chao index, Shannon index and Simpson index) of the intestinal flora. PCoA based on Bray-Curtis and Jaccard-Binary distances was used to determine the differences in community structure between groups. Permutation multivariate analysis of variance (also known as Adonis analysis) was used to test the statistical significance of differences between groups. R software was used to generate a bar chart of the bacterial composition in each sample from the phylum level to the species level, and the relative abundances of bacteria between groups were analyzed to determine the differential taxa. The differences between the two groups were analyzed by the nonparametric Mann-Whitney U test, and the differences between multiple groups were analyzed by the Kruskal-Wallis test, with P < 0.05 indicating a significant difference[19]. LEfSe was used to identify key taxa that differed between groups at the phylum to species levels. Functional groups (KO, KEGG pathway, CAZy, CARD, VFDB, etc.) that differed between groups were identified using LEfSe analysis[20].
The relative protein expression level was quantified by western blotting[21], and quantified with ImageJ software (v1.52a, NIH, Bethesda, MD, United States). Antibody were used with anti-glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (Proteintech, Wuhan, Hubei Province, China, 60004-1-Ig), HADHB polyclonal antibody (Proteintech, Wuhan, Hubei Province, China, Cat No. 29091-1-AP) and P21 polyclonal antibody (Proteintech, Wuhan, Hubei Province, China, Cat No. 28248-1-AP). The analysis of protein expression was conducted using ImageJ software.
The relative expression level of messenger RNA of was quantified using quantitative real-time-PCR, with 2-ΔΔCt method using GAPDH as a reference control[21]. The specific primers designed for the target genes are listed in the Supplementary material.
Paraffin-embedded rat liver tissue sections were deparaffinized in xylene and rehydrated through a graded ethanol series. Antigen retrieval was performed using ethylenediaminetetraacetic acid buffer (potential of hydrogen = 9.0, diluted 1:50 in distilled water) in a microwave oven at medium power for 8 minutes until boiling, followed by 7 minutes of heat preservation and an additional 8 minutes at medium-low power. The sections were cooled to room temperature and washed three times with phosphate buffered saline with tween (PBST) for 5 minutes each on a shaker. Endogenous peroxidase activity was blocked with 3% hydrogen peroxide solution for 25 minutes at room temperature in the dark. After they were washed with PBST (3 × 5 minutes), the sections were blocked with normal serum for 50 minutes at room temperature. Primary antibodies against P21 (1:200 dilution; ABclonal, A19094) andHADHB (1:200 dilution, Proteintech, 29091-1-AP, Wuhan, Hubei Province, China) were applied overnight at 4 °C in a humidified chamber. Following PBST washes, the sections were incubated with a horseradish peroxidase-conjugated goat anti-rabbit secondary antibody for 1 hour at 37 °C. Immunoreactive signals were visualized using 3,3’-diaminobenzidine substrate for approximately 8 minutes at room temperature. The sections were counterstained with hematoxylin for 1 minute, dehydrated, and mounted with neutral resin. A scanner (HAMAMATSU, NanoZoomer S60) was used to scan the images. Positive staining appeared as brown-yellow coloration, while the nuclei were stained blue.
The data are expressed as the means ± SD. Statistical analyses were performed using GraphPad Prism version 9.0. Differences between groups were evaluated using an unpaired two-tailed Student’s t test for comparisons between two groups or one-way analysis of variance followed by Tukey’s multiple comparison test for multigroup analyses. A probability value of P < 0.05 indicated statistical significance.
In this study, we employed metagenomic sequencing to characterize gut microbiota dynamics across the human lifespan in a cohort of healthy individuals (hereinafter referred to as the HP group; n = 300; age range: 0-100 years). The HP group was categorized into ten subgroups based on age (hereafter referred to as the S0, S1, to S9 groups; n = 30 for each group). Supplementary Table 1 summarizes key demographic and biochemical parameters stratified by age. The evaluated clinical parameters, complete blood count, a comprehensive metabolic panel including liver function tests, fasting lipid profile (triglycerides, total cholesterol, and low-density lipoprotein cholesterol), and fasting plasma glucose levels, were within normal reference ranges.
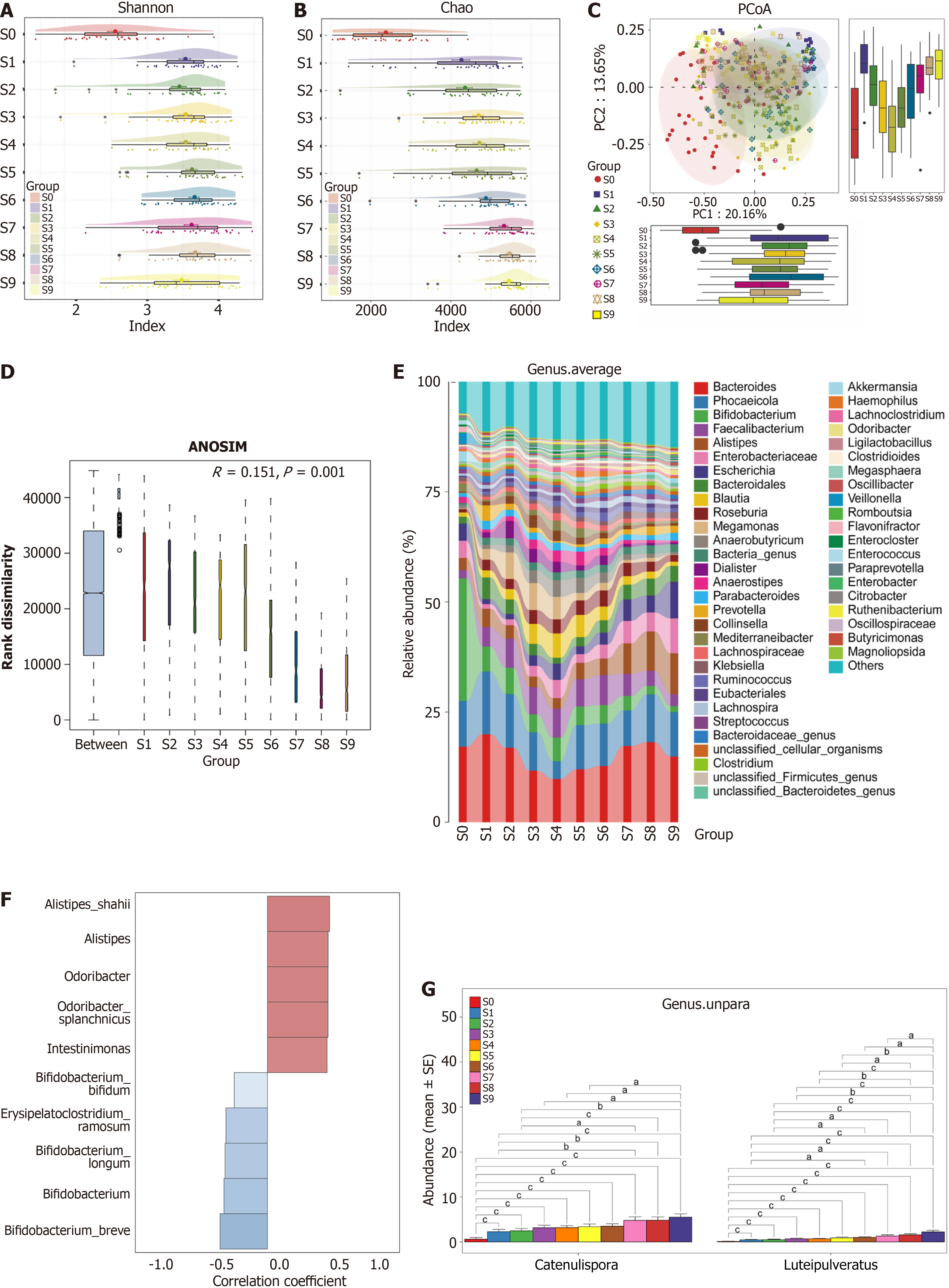
Accumulating evidence has indicated that the structural and functional composition of the gut microbiota significantly changes during aging[22,23]. To characterize the gut microbial features associated with aging, we performed a metagenomic analysis on ten incremental age groups within the HP group. Alpha diversity, as assessed by the Shannon index (Figure 2A), Chao index (Figure 2B), ACE index (Supplementary Figure 1A), and Simpson index (Supplementary Figure 1B), revealed notable variations across these groups. The most pronounced shift in microbial diversity was observed between the S0 and S1 groups. From S2 to S7, diversity remained relatively stable; however, a decline was evident starting from 70 years of age (S7 group). PCoA was employed to visualize dissimilarities in microbial communities, and beta diversity analysis indicated significant separation between groups (Figure 2C). The statistical significance of these differences was further confirmed using analysis of similarities (ANOSIM) (Figure 2D). The average compositional profiles and relative abundances of the gut microbiota at the genus (Figure 2E) and species (Supplementary Figure 2) levels demonstrated that Megamonas, Akkermansia, and Ligilactobacillus remained relatively stable throughout the aging process. Additionally, we applied LEfSe to identify species significantly enriched in each group and to evaluate their effect magnitudes. Several group-specific microbial signatures were identified at the species level (LDA > 3.0, P < 0.05), suggesting their potential as biomarkers for healthy aging (Supplementary Figure 2).
Correlation analysis between host age and microbial abundance revealed that Alistipes and Intestinimonas increased with age, whereas Bifidobacterium breve and Erysipelatoclostridium ramosum exhibited negative correlations with age (Figure 2F). Interestingly, the abundances of Catenulispora and Luteipulveratus also significantly increased in an age-dependent manner (Figure 2G).
To further characterize age-related changes in the gut microbiome composition in healthy rats (hereafter referred to as the HR cohort), fecal samples were collected at 2, 10, 20, and 48 weeks (named HR2, HR10, HR20, and HR48, respectively) and subjected to 16S rRNA gene sequencing analysis.
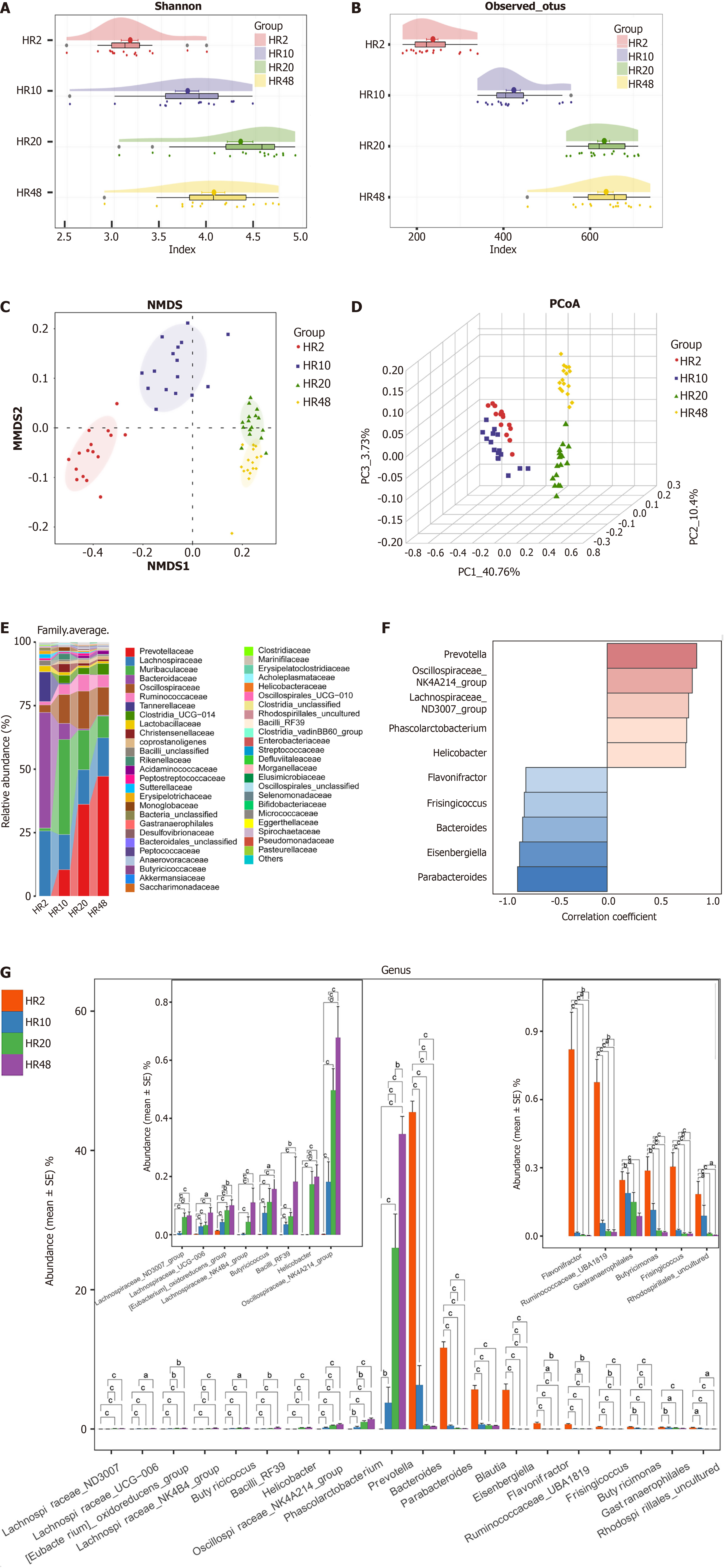
The Venn diagram indicated that 389 of the 1886 OTUs were shared among all four groups (Supplementary Figure 3A), whereas 154 were unique to the HR48 group. The Simpson index (Supplementary Figure 3B) and Shannon index (Figure 3A) revealed that taxonomic diversity increased from the HR2 group to the HR10 group and further to the HR20 group but then gradually decreased from HR20 to HR48. According to the observed OTU diversity index (Figure 3B), the Chao index (Supplementary Figure 3C) and the ACE index (Supplementary Figure 3D), gut microbial diversity increased progressively from the HR2 (n = 16) to the HR10 (n = 16) group to the HR20 (n = 16) but decreased significantly in the HR48 group compared with the HR20 group (all P < 0.05). Both nonmetric multidimensional scaling (Figure 3C) and PCoA (Figure 3D) analyses revealed clear separation of the gut microbiomes of the HR2, HR10, HR20, and HR48 groups into four distinct clusters. The average composition and relative abundance of the bacterial community at the genus level across groups are presented in Figure 3E.
Notably, correlation analysis revealed age-related trends in microbial composition: The abundance of Prevotella was positively correlated with age, while that of Parabacteroides was negatively correlated (Figure 3F). The longitudinal abundance dynamics of these aging-associated microbes are shown in Figure 3G. LDA at the genus level indicated that 32 genera, including Bacteroides, Parabacteroides, and Lachnoclostridium, which were significantly enriched in HR2 group, whereas 15 genera, such as Muribaculaceae, Alistipes, and Monoglobus, which were significantly enriched in HR10 group compared with the other three groups (P < 0.01, LDA score > 3) (Supplementary Figure 3E).
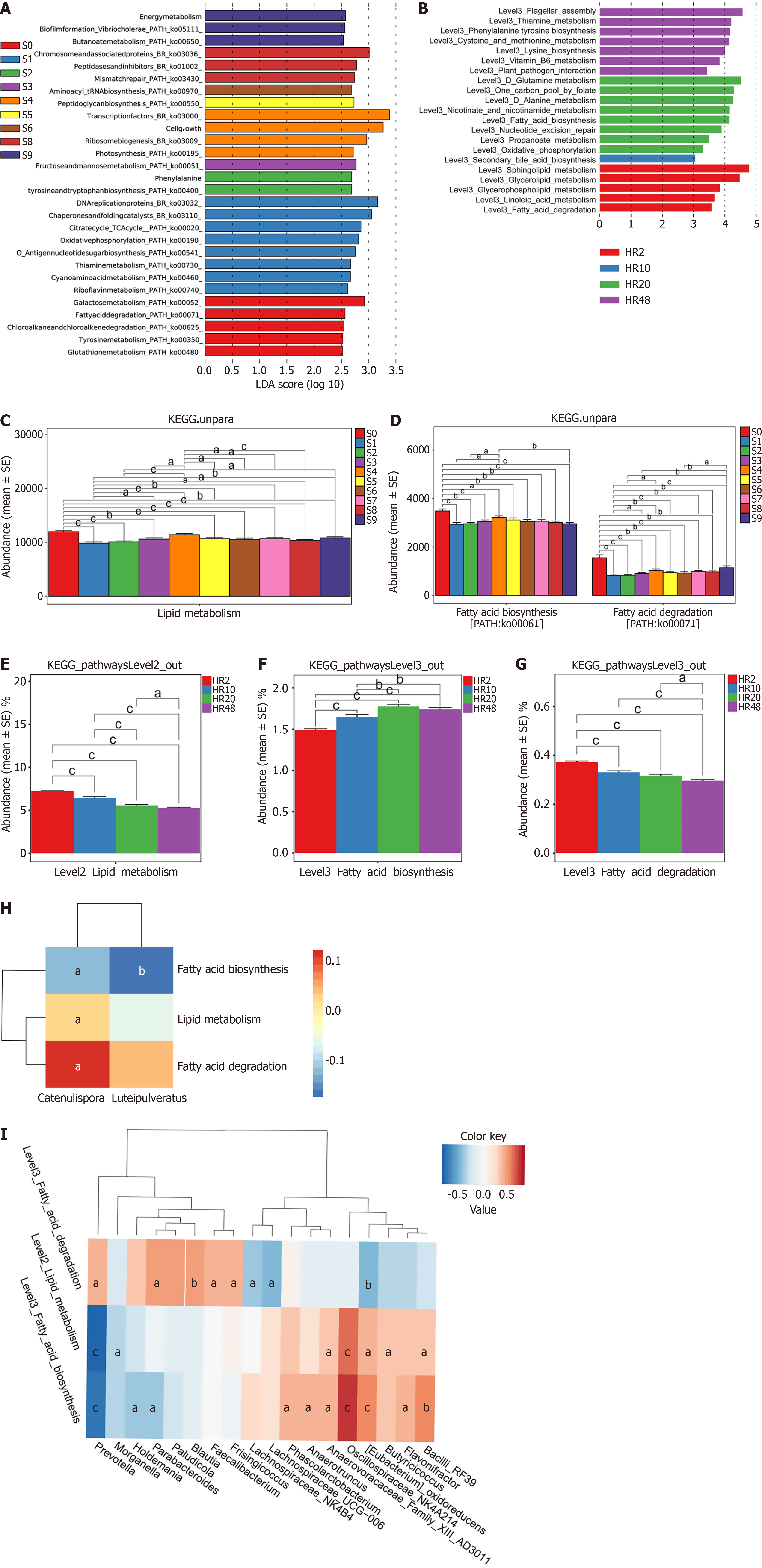
The functional potential of the microbiota was inferred from 16S rRNA and metagenomic data through parallel annotation against the KEGG, CAZy, and COG databases. KEGG pathway analysis revealed that stage-specific metabolic characteristics were associated with age. In the HP group, KEGG pathway analysis revealed that specific metabolic characteristics occurred with age at specific age stages. The functional predictions highlighted the significant role of lipid metabolism in healthy aging in both the human and the rat models (Figure 4A and B). Microbial abundance bar plots revealed progressive enrichment of lipid metabolism-associated microbiota from S0 to S9, accompanied by a decrease in the number of taxa linked to fatty acid biosynthesis and an increase in the number of those involved in fatty acid degradation (Figure 4C and D). Notably, the HR cohort exhibited similar trends to those of the HP group, indicating reproducible microbial patterns (Figure 4E-G).
Analysis of the lipid-metabolizing microbiota in the HP group revealed significant negative correlations between Catenulispora and Luteipulveratus and fatty acid biosynthesis pathways (P < 0.05), whereas Catenulispora was positively associated with β-oxidation enzymes (Figure 4H). Concurrently, in the HR cohort, Bacteroides, Flavonifractor, and Parabacteroides were positively correlated with lipid metabolism (all ρ = 0.79, P < 0.05), whereas Phascolarctobacterium (ρ = -0.73, P < 0.05), Butyricicoccus (ρ = -0.52, P < 0.05), and Faecalibacterium (ρ = -0.43, P < 0.05) were negatively correlated (Figure 4I).
COG enrichment indicated protein functions significantly represented across groups (P < 0.05, LDA > 2.5) (Supplementary Figure 4A). CAZy analysis revealed age-enriched metabolic pathways such as glycosyl transferases (GT) (GT51, GT9, GT19, GT5), which were significantly enriched in S8 and S9, suggesting elevated glycoside metabolism in older groups (P < 0.05, LDA > 3.0) (Supplementary Figure 4B). KEGG level 2 analysis revealed enrichment of infectious diseases (parasitic), excretory system, and amino acid metabolism in S0, whereas genetic information processing and replication and repair pathways were enriched in S8 (P < 0.05, LDA > 2.5) (Supplementary Figure 4C). KEGG level 3 analysis further revealed characteristic metabolic shifts during aging (Supplementary Figure 4D).
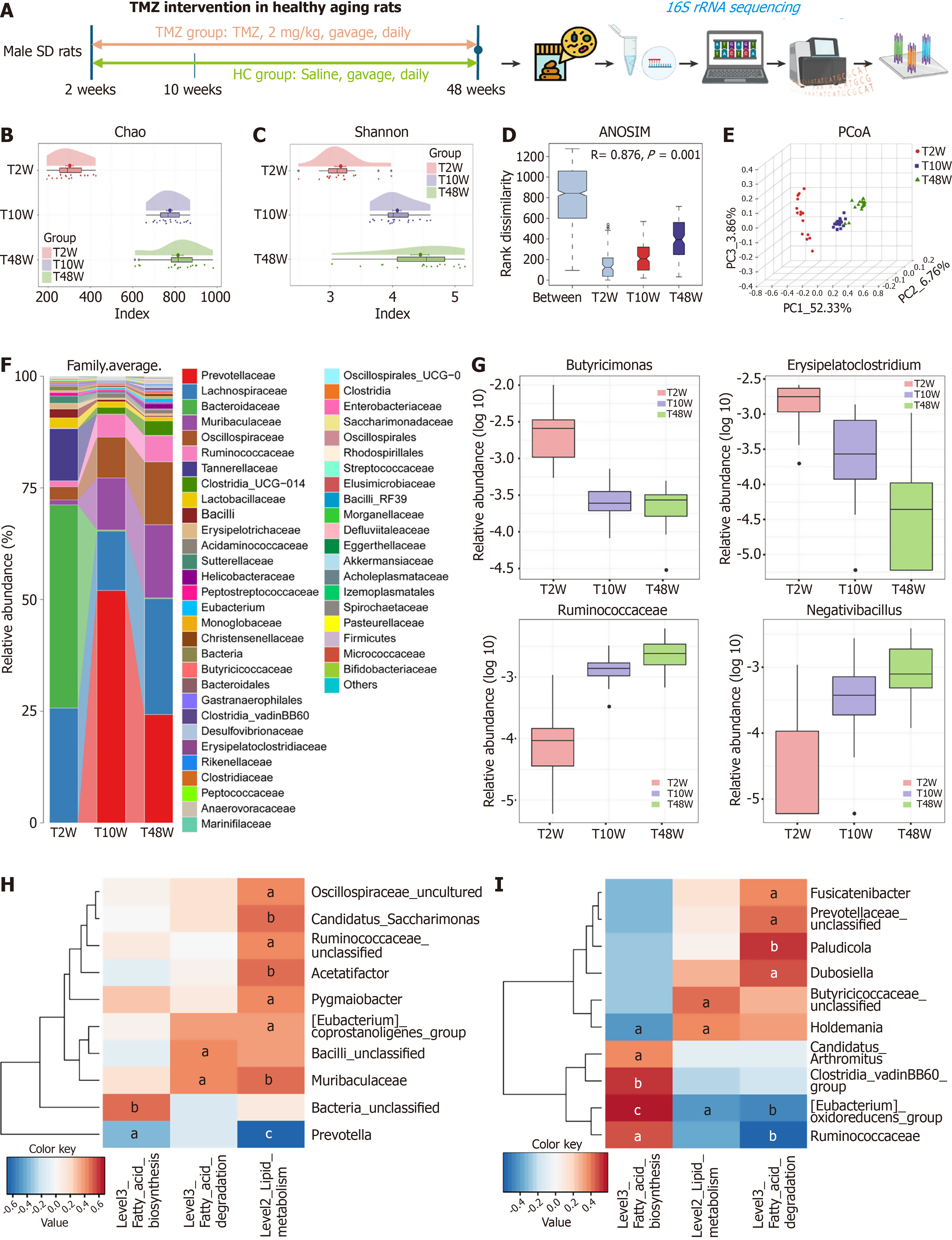
To clarify the crucial role of FAO in aging, the FAO inhibitor TMZ was subsequently used to treat natural aging rats (hereafter referred to as the TMZ cohort). The TMZ cohort was randomly divided into a HC group (which was treated with normal saline; n = 12) and a TMZ intervention (T) group (n = 18). Samples were collected at 2, 10, 20, and 48 weeks (named H2W, H10W, H20W, and H48W and T10W, T20W, and T48W, respectively). The experimental design for TMZ intervention is outlined in Figure 5A. The Chao index (Figure 5B) and Shannon index (Figure 5C) indicated a gradual increase in gut microbial diversity from T2W to T10W and T48W. ANOSIM (Figure 5D) and PCoA (Figure 5E) revealed clear separation of the gut microbiome into three distinct clusters corresponding to T2W, T10W, and T48W. The average relative abundance of bacterial communities at the genus level across the three TMZ-treated groups is shown in Figure 5F. Characteristic changes in gut microbial abundance during aging under TMZ intervention included a progressive decrease in Butyricimonas and Erysipelatoclostridium and a gradual increase in Ruminococcaceae and Negativibacillus (Figure 5G). Additionally, correlation analysis between lipid metabolism pathways and the microbiota in the T10W and H10W groups revealed that Acetatifactor and Candidatus Saccharimonas were positively correlated with lipid metabolism (P < 0.05), whereas Prevotella was negatively correlated (P < 0.05) (Figure 5H and I).
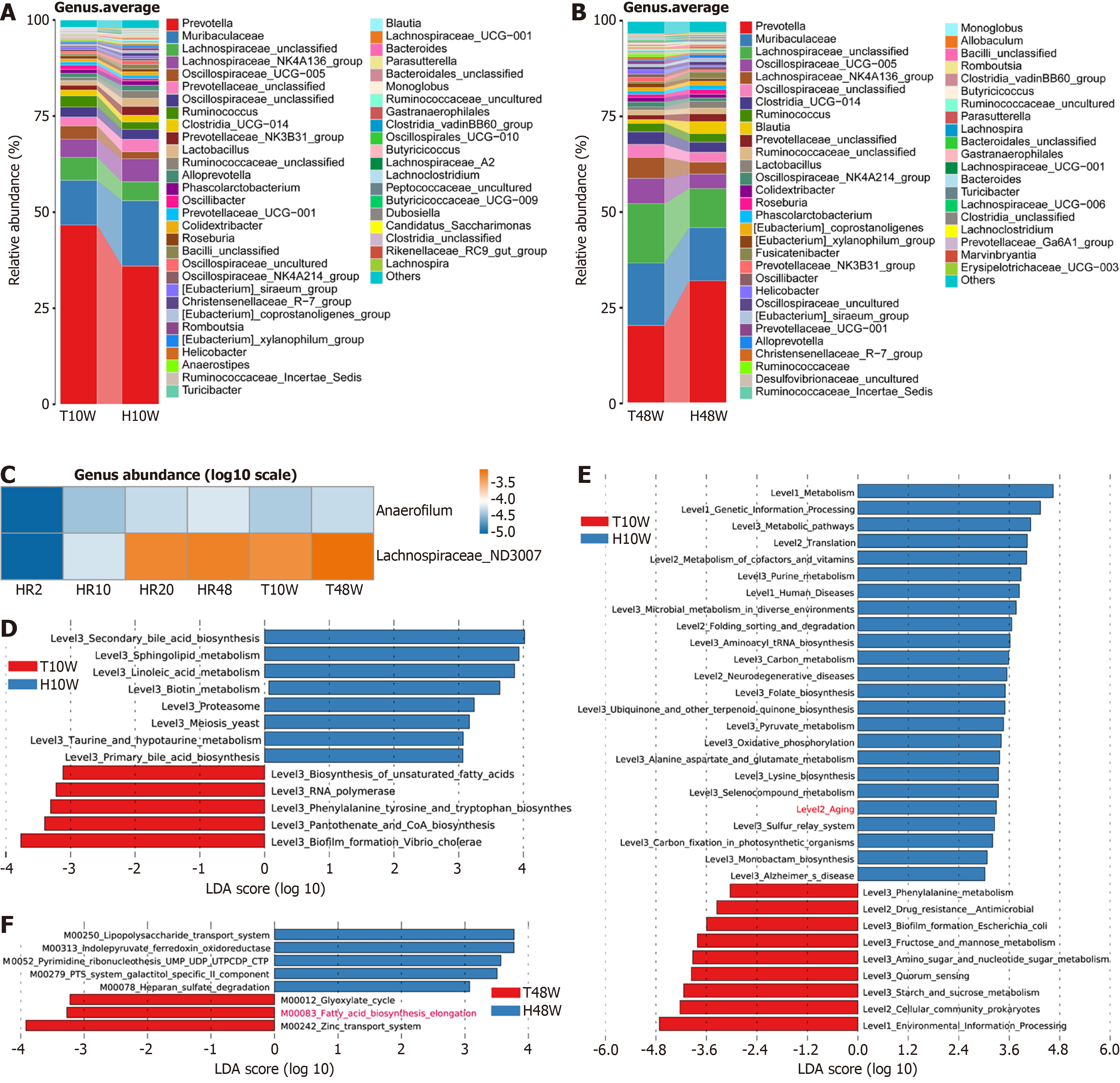
To assess the effect of TMZ on the intestinal microbiota during the growth process, we analyzed the gut microbiota of rats treated with TMZ. At the genus level, compared with the H10W group, the T10W group presented decreased average relative abundance of Muribaculaceae but increased relative abundance of Prevotella (Figure 6A). In the T48W group, the abundances of Prevotella and Blautia were lower than those in the H48W group were (Figure 6B). Correlation analysis between microbial abundance and age revealed significant associations for Anaerofilum and Lachnospiraceae (Figure 6C).
To assess functional and metabolic alterations in the gut microbiota, we performed functional prediction. LEfSe analysis of the KEGG modules indicated that M00122 (cobalamin biosynthesis) was significantly enriched in the T10W group compared with the H10W group (LDA > 3.0, P < 0.05) (Supplementary Figure 5). KEGG level 3 analysis further revealed enrichment of fatty acid biosynthesis in the T10W group compared with that in the H10W group (LDA > 3.0, P < 0.05) (Figure 6D). In contrast, compared with the T10W group, the H10W group showed greater enrichment in aging-related metabolic pathways (Figure 6E). Consistent with the 10-week results, the microbial enrichment in fatty acid elongation was significantly greater in the T48W group (KEGG map 00083) than in the H48W group (LDA > 3.0, P < 0.05) (Figure 6F). These findings indicated that TMZ might modulate the gut microbial community structure and species richness in rats, thus regulating lipid metabolism.
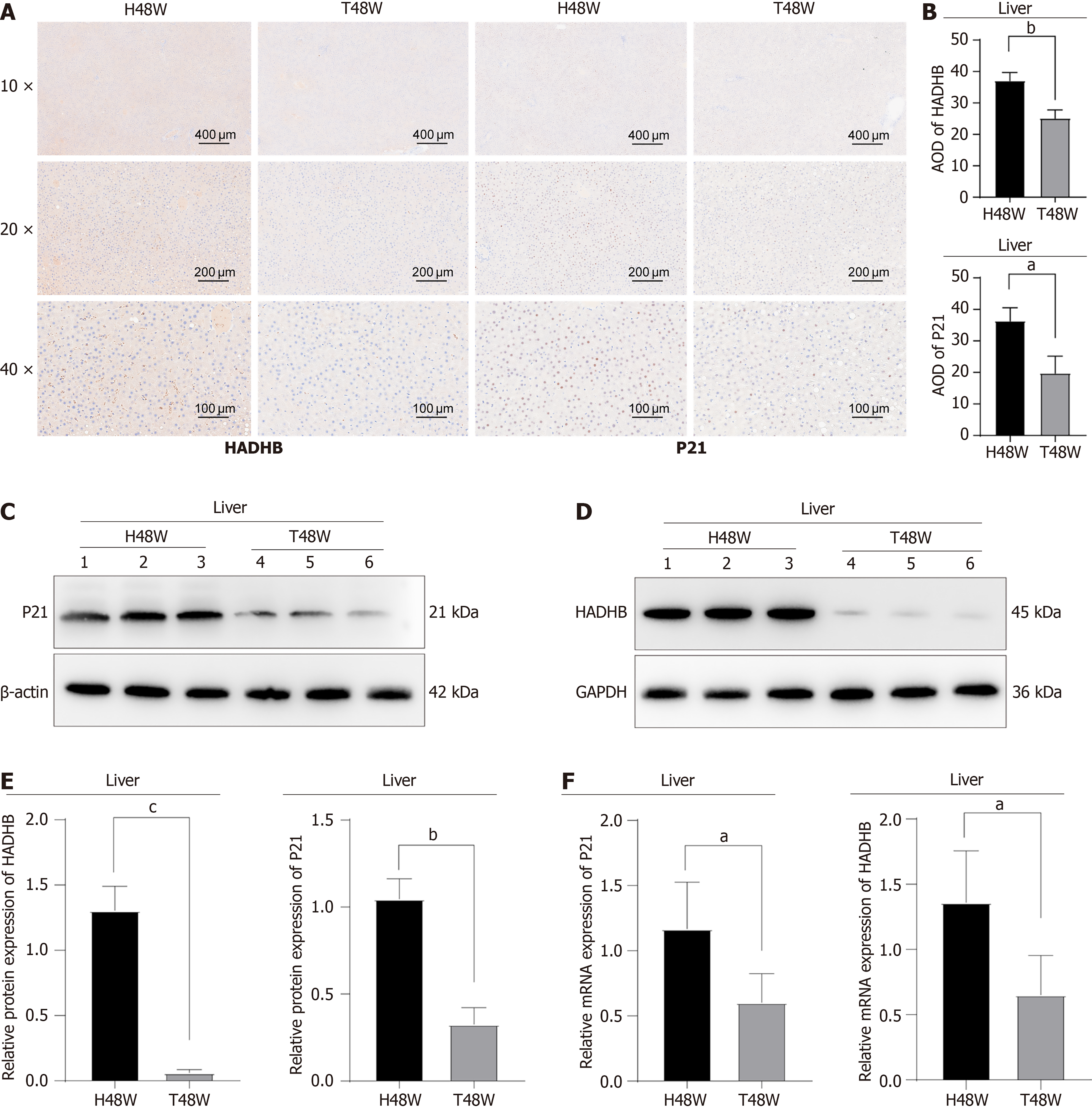
The liver serves as a central hub for fatty acid metabolism and plays a crucial role in regulating whole-body lipid homeostasis and energy metabolism during aging. As the primary organ responsible for fatty acid β-oxidation and lipid metabolism, the liver represents a critical target for understanding age-related metabolic alterations. To investigate the effects of TMZ on hepatic FAO and senescence markers, we performed immunohistochemical analysis, western blotting, and quantitative PCR on liver tissues from rats treated with TMZ for 48 weeks and demonstrated that TMZ significantly modulates FAO pathways (HADHB, a critical enzyme in the mitochondrial trifunctional protein complex involved in FAO) and cellular senescence markers during the normal aging process. Immunohistochemical analysis revealed significantly lower expression of P21 and HADHB in the T48W group than in the H48W group (P < 0.05) (Figure 7A and B). Consistent with these findings, both the protein and gene expression levels of P21 and HADHB were markedly reduced in the T48W group (P < 0.05) (Figure 7C-F).
The gut microbiota is increasingly implicated in the aging process. The interaction between gut microorganisms and host metabolism is essential for maintaining overall physiological homeostasis. Although the gut microbiota has been implicated in the aging process, the specific characteristics and functional roles of aging-associated microbial communities remain insufficiently explored[24]. In this study, we employed metagenomic sequencing to analyze fecal samples from a cohort of 300 healthy Chinese individuals spanning 0-100 years of age, delineating microbial dynamics across the human lifespan and their influence on host metabolic pathways. Parallel investigations in a rodent model of healthy aging further revealed conserved shifts in microbial diversity and composition, particularly involving taxa associated with lipid metabolism. On the basis of these observational findings, we further conducted an in vivo interventional study in which the fatty acid β-oxidation inhibitor TMZ was used to evaluate the effects of TMZ on the intestinal microbiota and FAO during the growth process of rats.
Consistent with previous studies[25,26], our multicohort analysis revealed significant alterations in the gut microbiome throughout the aging process. Notably, we found that age-associated remodeling of the gut microbiome followed a nonlinear trajectory. Microbial diversity initially increased gradually with age but began to decline after 70 years of age. Our findings are consistent with those of a multicohort study by Wilmanski et al[27], which demonstrated that the α diversity of the gut microbiota increases with age in young adults (20-45 years) and stabilizes at approximately 40 years of age. However, our study extends this temporal pattern by encompassing the complete lifespan from 0 year to 100 years, revealing that α diversity decreases continuously after 70 years of age, thus demonstrating a biphasic relationship between aging and microbial diversity that may be modulated by multiple factors, including sex, geography, and ethnicity. Specifically, the abundance of bacterial species, including Faecalibacterium, prauznitzii, Phocaeicola vulgatus and Bifidobacterium longum, decreased in the aged population. At the species level, Proteobacteria are the key microbes for the age range from 90 years to 100 years. Proteobacteria are present in the gut microbes of healthy adults and maintain low abundance through host-microbe signaling, which constitutes the predominant component of pathogenic bacteria and plays a key role in driving aging processes[28]. Specifically, the bacterial species were found decreased in the aged population, including Faecalibacterium, prauznitzii, Phocaeicola vulgatus and Bifidobacterium longum. At the species level, Proteobacteria are the key microbes for the age stage from 90 years to 100 years. Proteobacteria presents in the gut microbes of healthy adults and maintains low abundance through host-microbe signaling, which constitute the predominant composition of pathogenic bacteria and play a key role in driving aging processes[29]. Interestingly, we also found that healthy bacterial species associated with metabolism, including Proteobacteria, Actinobacteria and Verrucomicrobia, Christensenellaceae and Lactobacillus, were more abundant in older adults than in younger adults, indicating a close possible connection between metabolism and aging. Additionally, similar phenomena were observed in the gut microbiota of mice of different ages.
Our findings reveal that there was a significant increase in the abundance of microbes linked to lipid metabolism as age progressed. Metabolic intermediates produced during FAO, including reactive oxygen species, accumulate in cells, resulting in damage to mitochondria and other organelles and thus inducing oxidative stress. Notably, oxidative stress is recognized as among the important triggers of aging. Li et al[30] reported that after adjusting for age and sex, healthy centenarians exhibited increased metabolic processes such as fatty acid β-oxidation. Additionally, cellular fatty acid storage, FAO, pyruvate metabolism, branchial amino acid metabolism, the tricarboxylic acid cycle and oxidative phosphorylation were also elevated in centenarians. Hence, a reciprocal relationship exists between microbial dysbiosis, aging and overall health. The mechanism through which gut microbiota dysbiosis affects the aging process may involve significant changes in the human gut microbiota, resulting in reduced microecological diversity. Disrupted gut microbes interact with host metabolic signaling pathways through their metabolites to promote inflammation and tissue dysfunction, thus contributing to systemic metabolic dysfunction and various age-related pathologies[31].
To further investigate the role of lipid metabolism in aging, we treated healthy rats with the FAO inhibitor TMZ. The gut microbiota directly influences liver metabolism via portal circulation, while bile acids secreted by the liver in turn regulate the gut microbiota, forming a gut-liver axis feedback loop. Additionally, as the liver is the primary site for fatty acid β-oxidation and lipid metabolism, it represents the most relevant organ for assessing the consequences of alterations in the gut microbiota and enhanced FAO. The liver, which is characterized by accelerated aging, is recognized as an ideal experimental model for investigating senescence mechanisms[32]. Therefore, this study focused on phenotypic validation in liver tissue. Our results revealed that TMZ modified the composition of the gut microbiota and significantly reduced the expression of liver aging markers (P21 and P16) and the FAO enzyme HADHB (P < 0.05). These results indicate, to some extent, a mechanistic link between changes in the gut microbiota and liver metabolic function during aging. Elmansi et al[33] proposed that TMZ could inhibit lysine acetyltransferase, activate adenosine 5’-monophosphate-activated protein kinase, and subsequently inhibit endoplasmic reticulum stress-induced apoptosis and the pro-inflammatory effects of nuclear factor kappa-B, serving as a common target in aging, which might be the possible mechanism by which TMZ exerts its anti-aging effects. The mechanisms linking lipid metabolism and aging are complex. Lipids play crucial roles in regulating aging and longevity. In the past few decades, a series of genetic pathways have been discovered to regulate lifespan in model organisms. Interestingly, many of these regulatory pathways are linked to lipid metabolism and lipid signaling. Lipid metabolic enzymes undergo significant changes during aging and are regulated by different longevity pathways[34]. One study used the genome scale precision metabolic modeling approach to examine the metabolic network in an elderly Chinese population, identifying elevated FAO as the most prominent metabolic feature in these older individuals[30]. This suggests that correcting aging associated reprogramming of lipid metabolism, particularly FAO dysregulation may be a potential mechanism underlying its anti-aging effects. By alleviating lipotoxicity, limiting excessive FAO flux, and reducing mitochondrial burden and oxidative stress, aging may be delayed[35].
Furthermore, TMZ intervention resulted in significant alterations in the gut microbiota composition. Notably, microbial diversity in TMZ-treated rats continued to increase with age but did not decrease in the HR and HP groups, suggesting that TMZ might increase gut microbiota diversity and maintain a more youthful microbial state. Our analysis revealed that following TMZ intervention, the relative abundance of lipid metabolism-related bacteria, including Muribaculaceae, Prevotellaceae and Lachnospiraceae, significantly increased in the altered gut microbial communities. These findings are highly consistent with existing research, confirming the critical relationship between microbiota diversity maintenance and healthy aging[36]. With advancing age, the gut microbiome undergoes significant compositional changes that often lead to dysbiosis, which is associated with increased inflammation, compromised immunity, and metabolic dysfunction[37]. TMZ intervention, by reversing the age-related decline in microbial diversity, may represent an effective antiaging therapeutic strategy. Muribaculaceae bacteria participate in short-chain fatty acid synthesis and influence host energy homeostasis and lipid metabolism[38], while Lachnospiraceae exhibit anti-inflammatory properties and are enriched in long-lived populations[39]. These bacterial families play crucial roles in lipid metabolism, including peroxisome proliferator-activated receptor signaling pathways and unsaturated fatty acid biosynthesis[40]. Therefore, TMZ may exert antiaging effects by modulating the gut microbiota composition, particularly by increasing the abundance of these beneficial bacteria, via mechanisms including maintaining a “youthful” state of microbial diversity, increasing short-chain fatty acid production, and optimizing lipid metabolic pathways.
Limitations of this study: First, all the samples in this study were collected from Henan Province, China, which may introduce sampling bias. The findings of this study represent age-related associations rather than absolute causal relationships, and these changes may be influenced by other age-related confounding factors within the cohorts. Additionally, while this study demonstrated that TMZ could alter the gut microbiota in rats and inhibit hepatic FAO while reducing the expression of liver aging markers, whether changes in the microbiota are the direct cause of FAO inhibition requires further experimental validation. Finally, this study used metagenomic data for functional prediction but did not incorporate metabolomics analysis. Future research integrating metabolomics will more directly elucidate the role of microbiota-host co-metabolism in aging.
This study delineated the evolutionary landscape of the gut microbiota during the aging process in both humans and rats. Functional enrichment analysis revealed a possible association between the gut microbiota and lipid metabolism. Finally, the FAO inhibitor TMZ may delay the aging process in rats by targeting the functional gut microbiota associated with lipid metabolism.
We thank all the generous volunteer subjects who enrolled in the study. We thank Dr. Ren HY and Liu C (Shanghai Mobio Biomedical Technology Co., Ltd., Shanghai, China) for their help on data analysis.
| 1. | Dogra S, Dunstan DW, Sugiyama T, Stathi A, Gardiner PA, Owen N. Active Aging and Public Health: Evidence, Implications, and Opportunities. Annu Rev Public Health. 2022;43:439-459. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 182] [Article Influence: 36.4] [Reference Citation Analysis (0)] |
| 2. | Fu J, Qiu W, Zheng H, Qi C, Hu S, Wu W, Wang H, Wu G, Cao P, Ma Z, Zheng C, Ma WJ, Zhou HW, He Y. Ageing trajectory of the gut microbiota is associated with metabolic diseases in a chronological age-dependent manner. Gut. 2023;72:1431-1433. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 25] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 3. | Xu K, Guo Y, Wang Y, Ren Y, Low V, Cho S, Ping L, Peng K, Li X, Qiu Y, Liu Q, Li Z, Wang Z. Decreased Enterobacteriaceae translocation due to gut microbiota remodeling mediates the alleviation of premature aging by a high-fat diet. Aging Cell. 2023;22:e13760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 23] [Reference Citation Analysis (0)] |
| 4. | Ijaz MU, Vaziri F, Wan YY. Effects of Bacillus Calmette-Guérin on immunometabolism, microbiome and liver diseases(⋆). Liver Res. 2023;7:116-123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 5. | Leite G, Pimentel M, Barlow GM, Chang C, Hosseini A, Wang J, Parodi G, Sedighi R, Rezaie A, Mathur R. Age and the aging process significantly alter the small bowel microbiome. Cell Rep. 2021;36:109765. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 112] [Article Influence: 22.4] [Reference Citation Analysis (3)] |
| 6. | Lin SW, Tsai YS, Chen YL, Wang MF, Chen CC, Lin WH, Fang TJ. Lactobacillus plantarum GKM3 Promotes Longevity, Memory Retention, and Reduces Brain Oxidation Stress in SAMP8 Mice. Nutrients. 2021;13:2860. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 40] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 7. | Maynard C, Weinkove D. The Gut Microbiota and Ageing. Subcell Biochem. 2018;90:351-371. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 87] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 8. | Thevaranjan N, Puchta A, Schulz C, Naidoo A, Szamosi JC, Verschoor CP, Loukov D, Schenck LP, Jury J, Foley KP, Schertzer JD, Larché MJ, Davidson DJ, Verdú EF, Surette MG, Bowdish DME. Age-Associated Microbial Dysbiosis Promotes Intestinal Permeability, Systemic Inflammation, and Macrophage Dysfunction. Cell Host Microbe. 2017;21:455-466.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1066] [Cited by in RCA: 943] [Article Influence: 104.8] [Reference Citation Analysis (4)] |
| 9. | Das UN. "Cell Membrane Theory of Senescence" and the Role of Bioactive Lipids in Aging, and Aging Associated Diseases and Their Therapeutic Implications. Biomolecules. 2021;11:241. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 94] [Article Influence: 18.8] [Reference Citation Analysis (0)] |
| 10. | Collino S, Montoliu I, Martin FP, Scherer M, Mari D, Salvioli S, Bucci L, Ostan R, Monti D, Biagi E, Brigidi P, Franceschi C, Rezzi S. Metabolic signatures of extreme longevity in northern Italian centenarians reveal a complex remodeling of lipids, amino acids, and gut microbiota metabolism. PLoS One. 2013;8:e56564. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 143] [Cited by in RCA: 191] [Article Influence: 14.7] [Reference Citation Analysis (0)] |
| 11. | Yamauchi S, Sugiura Y, Yamaguchi J, Zhou X, Takenaka S, Odawara T, Fukaya S, Fujisawa T, Naguro I, Uchiyama Y, Takahashi A, Ichijo H. Mitochondrial fatty acid oxidation drives senescence. Sci Adv. 2024;10:eado5887. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 33] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 12. | Tsugawa H, Ishihara T, Ogasa K, Iwanami S, Hori A, Takahashi M, Yamada Y, Satoh-Takayama N, Ohno H, Minoda A, Arita M. A lipidome landscape of aging in mice. Nat Aging. 2024;4:709-726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 42] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 13. | Ogłuszka M, Chen CY, Poławska E, Starzyński RR, Liput K, Siekierko U, Pareek CS, Pierzchała M, Kang JX. Elevated tissue status of omega-3 fatty acids protects against age-related telomere attrition in fat-1 transgenic mice. Clin Nutr. 2024;43:1488-1494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 14. | Mei Z, Yilamu K, Ni W, Shen P, Pan N, Chen H, Su Y, Guo L, Sun Q, Li Z, Huang D, Fang X, Fan S, Zhang H, Shen S. Chondrocyte fatty acid oxidation drives osteoarthritis via SOX9 degradation and epigenetic regulation. Nat Commun. 2025;16:4892. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 24] [Reference Citation Analysis (0)] |
| 15. | Gao M, Wang H, Luo H, Sun Y, Wang L, Ding S, Ren H, Gang J, Rao B, Liu S, Wang X, Gao X, Li M, Zou Y, Liu C, Yuan C, Sun J, Cui G, Ren Z. Characterization of the Human Oropharyngeal Microbiomes in SARS-CoV-2 Infection and Recovery Patients. Adv Sci (Weinh). 2021;8:e2102785. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 35] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 16. | Ren Z, Liu S, Wang Q, Rao B, Zeng Z, Xu Y, Wang H, Luo H, Gou J, Yu Z. Alterations of the oral and gut mycobiome and cytokines during long-term follow-up of COVID-19 convalescents. Signal Transduct Target Ther. 2023;8:166. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 17. | Cui GY, Rao BC, Zeng ZH, Wang XM, Ren T, Wang HY, Luo H, Ren HY, Liu C, Ding SY, Tan JJ, Liu ZG, Zou YW, Ren ZG, Yu ZJ. Characterization of oral and gut microbiome and plasma metabolomics in COVID-19 patients after 1-year follow-up. Mil Med Res. 2022;9:32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 30] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 18. | Zou Y, Sun Y, Chen X, Hong L, Dong G, Bai X, Wang H, Rao B, Ren Z, Yu Z. Nanosecond pulse effectively ablated hepatocellular carcinoma with alterations in the gut microbiome and serum metabolites. Front Pharmacol. 2023;14:1163628. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 19. | Zhang Y, Gu Y, Ren H, Wang S, Zhong H, Zhao X, Ma J, Gu X, Xue Y, Huang S, Yang J, Chen L, Chen G, Qu S, Liang J, Qin L, Huang Q, Peng Y, Li Q, Wang X, Kong P, Hou G, Gao M, Shi Z, Li X, Qiu Y, Zou Y, Yang H, Wang J, Xu G, Lai S, Li J, Ning G, Wang W. Gut microbiome-related effects of berberine and probiotics on type 2 diabetes (the PREMOTE study). Nat Commun. 2020;11:5015. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 299] [Cited by in RCA: 245] [Article Influence: 40.8] [Reference Citation Analysis (4)] |
| 20. | Vatanen T, Jabbar KS, Ruohtula T, Honkanen J, Avila-Pacheco J, Siljander H, Stražar M, Oikarinen S, Hyöty H, Ilonen J, Mitchell CM, Yassour M, Virtanen SM, Clish CB, Plichta DR, Vlamakis H, Knip M, Xavier RJ. Mobile genetic elements from the maternal microbiome shape infant gut microbial assembly and metabolism. Cell. 2022;185:4921-4936.e15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 135] [Article Influence: 33.8] [Reference Citation Analysis (1)] |
| 21. | Chen X, Shen J, Jiang X, Pan M, Chang S, Li J, Wang L, Miao M, Feng X, Zhang L, Shu G, Liu W, Xu F, Zhang W, Ding Z, Zong H, Liu W, Li D, Chen B, Shao M, Fei G, Zha X, Fan X. Characterization of dipyridamole as a novel ferroptosis inhibitor and its therapeutic potential in acute respiratory distress syndrome management. Theranostics. 2024;14:6947-6968. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 22] [Reference Citation Analysis (0)] |
| 22. | Parker A, Romano S, Ansorge R, Aboelnour A, Le Gall G, Savva GM, Pontifex MG, Telatin A, Baker D, Jones E, Vauzour D, Rudder S, Blackshaw LA, Jeffery G, Carding SR. Fecal microbiota transfer between young and aged mice reverses hallmarks of the aging gut, eye, and brain. Microbiome. 2022;10:68. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 295] [Cited by in RCA: 261] [Article Influence: 65.3] [Reference Citation Analysis (4)] |
| 23. | Sato Y, Atarashi K, Plichta DR, Arai Y, Sasajima S, Kearney SM, Suda W, Takeshita K, Sasaki T, Okamoto S, Skelly AN, Okamura Y, Vlamakis H, Li Y, Tanoue T, Takei H, Nittono H, Narushima S, Irie J, Itoh H, Moriya K, Sugiura Y, Suematsu M, Moritoki N, Shibata S, Littman DR, Fischbach MA, Uwamino Y, Inoue T, Honda A, Hattori M, Murai T, Xavier RJ, Hirose N, Honda K. Novel bile acid biosynthetic pathways are enriched in the microbiome of centenarians. Nature. 2021;599:458-464. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 449] [Article Influence: 89.8] [Reference Citation Analysis (1)] |
| 24. | Boyajian JL, Ghebretatios M, Schaly S, Islam P, Prakash S. Microbiome and Human Aging: Probiotic and Prebiotic Potentials in Longevity, Skin Health and Cellular Senescence. Nutrients. 2021;13:4550. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 75] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 25. | Yang H, Wang T, Qian C, Wang H, Yu D, Shi M, Fu M, Liu X, Pan M, Rong X, Xiao Z, Chen X, Yeerken A, Wu Y, Zheng Y, Yang H, Zhang M, Liu T, Qiao P, Qu Y, Lin Y, Huang Y, Jin J, Liu N, Wen Y, Sun N, Zhao C. Gut microbial-derived phenylacetylglutamine accelerates host cellular senescence. Nat Aging. 2025;5:401-418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 45] [Reference Citation Analysis (0)] |
| 26. | Zumerle S, Sarill M, Saponaro M, Colucci M, Contu L, Lazzarini E, Sartori R, Pezzini C, Rinaldi A, Scanu A, Sgrignani J, Locatelli P, Sabbadin M, Valdata A, Brina D, Giacomini I, Rizzo B, Pierantoni A, Sharifi S, Bressan S, Altomare C, Goshovska Y, Giraudo C, Luisetto R, Iaccarino L, Torcasio C, Mosole S, Pasquini E, Rinaldi A, Pellegrini L, Peron G, Fassan M, Masiero S, Giori AM, Dall'Acqua S, Auwerx J, Cippà P, Cavalli A, Bolis M, Sandri M, Barile L, Montopoli M, Alimonti A. Targeting senescence induced by age or chemotherapy with a polyphenol-rich natural extract improves longevity and healthspan in mice. Nat Aging. 2024;4:1231-1248. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 64] [Article Influence: 32.0] [Reference Citation Analysis (0)] |
| 27. | Wilmanski T, Diener C, Rappaport N, Patwardhan S, Wiedrick J, Lapidus J, Earls JC, Zimmer A, Glusman G, Robinson M, Yurkovich JT, Kado DM, Cauley JA, Zmuda J, Lane NE, Magis AT, Lovejoy JC, Hood L, Gibbons SM, Orwoll ES, Price ND. Gut microbiome pattern reflects healthy ageing and predicts survival in humans. Nat Metab. 2021;3:274-286. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 575] [Cited by in RCA: 480] [Article Influence: 96.0] [Reference Citation Analysis (4)] |
| 28. | de la Cuesta-Zuluaga J, Kelley ST, Chen Y, Escobar JS, Mueller NT, Ley RE, McDonald D, Huang S, Swafford AD, Knight R, Thackray VG. Age- and Sex-Dependent Patterns of Gut Microbial Diversity in Human Adults. mSystems. 2019;4:e00261-e00219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 211] [Cited by in RCA: 283] [Article Influence: 40.4] [Reference Citation Analysis (0)] |
| 29. | Pang S, Chen X, Lu Z, Meng L, Huang Y, Yu X, Huang L, Ye P, Chen X, Liang J, Peng T, Luo W, Wang S. Longevity of centenarians is reflected by the gut microbiome with youth-associated signatures. Nat Aging. 2023;3:436-449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 116] [Article Influence: 38.7] [Reference Citation Analysis (0)] |
| 30. | Li GH, Han F, Xiao FH, Gu KS, Shen Q, Xu W, Li WX, Wang YL, Liang B, Huang JF, Xiao W, Kong QP. System-level metabolic modeling facilitates unveiling metabolic signature in exceptional longevity. Aging Cell. 2022;21:e13595. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 23] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 31. | Gu P, Wei R, Liu R, Yang Q, He Y, Guan J, He W, Li J, Zhao Y, Xie L, He J, Guo Q, Hu J, Bao J, Wang W, Guo J, Zeng Z, Chen Z, Jiang Y, Liu Z, Chen P. Aging-induced Alternation in the Gut Microbiota Impairs Host Antibacterial Defense. Adv Sci (Weinh). 2025;12:e2411008. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 32. | Rossetti GG, Dommann N, Karamichali A, Dionellis VS, Asensio Aldave A, Yarahmadov T, Rodriguez-Carballo E, Keogh A, Candinas D, Stroka D, Halazonetis TD. In vivo DNA replication dynamics unveil aging-dependent replication stress. Cell. 2024;187:6220-6234.e13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 14] [Reference Citation Analysis (0)] |
| 33. | Elmansi AM, Kassem A, Castilla RM, Miller RA. Downregulation of the NF-κB protein p65 is a shared phenotype among most anti-aging interventions. Geroscience. 2025;47:3077-3094. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 34. | Mutlu AS, Duffy J, Wang MC. Lipid metabolism and lipid signals in aging and longevity. Dev Cell. 2021;56:1394-1407. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 229] [Article Influence: 45.8] [Reference Citation Analysis (0)] |
| 35. | Papsdorf K, Brunet A. Linking Lipid Metabolism to Chromatin Regulation in Aging. Trends Cell Biol. 2019;29:97-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 116] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 36. | Zhao Z, Hu Z, Li L. Cardiac energy metabolic disorder and gut microbiota imbalance: a study on the therapeutic potential of Shenfu Injection in rats with heart failure. Front Microbiol. 2025;16:1509548. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 37. | Boehme M, Guzzetta KE, Bastiaanssen TFS, van de Wouw M, Moloney GM, Gual-Grau A, Spichak S, Olavarría-Ramírez L, Fitzgerald P, Morillas E, Ritz NL, Jaggar M, Cowan CSM, Crispie F, Donoso F, Halitzki E, Neto MC, Sichetti M, Golubeva AV, Fitzgerald RS, Claesson MJ, Cotter PD, O'Leary OF, Dinan TG, Cryan JF. Microbiota from young mice counteracts selective age-associated behavioral deficits. Nat Aging. 2021;1:666-676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 206] [Article Influence: 41.2] [Reference Citation Analysis (0)] |
| 38. | Xia M, Xu Y, Li H, Huang J, Zhou H, Gao C, Han J. Structural and functional alteration of the gut microbiota in elderly patients with hyperlipidemia. Front Cell Infect Microbiol. 2024;14:1333145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 11] [Reference Citation Analysis (0)] |
| 39. | DeJong EN, Surette MG, Bowdish DME. The Gut Microbiota and Unhealthy Aging: Disentangling Cause from Consequence. Cell Host Microbe. 2020;28:180-189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 332] [Cited by in RCA: 272] [Article Influence: 45.3] [Reference Citation Analysis (0)] |
| 40. | Šardzíková S, Gajewska M, Gałka N, Štefánek M, Baláž A, Garaiová M, Holič R, Świderek W, Šoltys K. Can longer lifespan be associated with gut microbiota involvement in lipid metabolism? FEMS Microbiol Ecol. 2024;100:fiae135. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |