Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.116679
Revised: December 20, 2025
Accepted: January 28, 2026
Published online: April 21, 2026
Processing time: 148 Days and 20.2 Hours
Chronic hepatitis B (CHB)-related liver fibrosis is a major driver of severe hepatic complications, with substantial interindividual heterogeneity in histological outcomes after antiviral therapy. Histopathological images contain rich biological information, but deep learning (DL) models for predicting on-treatment histo
To develop and independently validate a DL-based multimodal model to predict fibrosis reversal following antiviral therapy using histopathological images and clinical features.
This multicenter study included 238 patients from 14 institutions who received antiviral therapy and had both hematoxylin and eosin (HE) and Masson-stained liver biopsy slides available. The training, validation, and test cohorts comprised 114, 50, and 74 patients, respectively. Convolutional neural network models were independently developed using HE- and Masson-stained images, and subsequently combined with clinical features to construct a multimodal predictive model for evaluating fibrosis regression after antiviral treatment.
The HE model achieved areas under the receiver operating characteristic curves (AUCs) of 0.657 and 0.615 in the validation and test sets, respectively. The Masson model yielded AUCs of 0.727 and 0.676 for the corresponding sets. The clinical model exhibited AUCs of 0.658 in the validation set and 0.588 in the test set. The multimodal fusion model demonstrated enhanced discriminatory performance, reaching AUCs of 0.741 and 0.694 in the validation and test sets, respectively. Subgroup analysis showed robust predictive capacity in patients with progressive fibrosis (AUC = 0.779) and in those with hepatitis B e antigen-positive status (AUC = 0.755). Gradient-weighted class activation mapping revealed that the model focused primarily on key histological features associated with non-reversal, including hepatocyte degeneration, disorganized hepatic cords, and thick-bridging fibrous septa.
Digital pathology and clinical-based DL accurately predict fibrosis regression after antiviral therapy in CHB-related liver fibrosis, particularly in patients with progressive fibrosis and hepatitis B e antigen-positive status, supporting personalized treatment strategies.
Core Tip: This study presents a multimodal model that integrates pathological slide staining with clinical features to predict the probability of histological reversal following standard antiviral therapy in patients with advanced chronic hepatitis B-related fibrosis. The model demonstrates robust predictive accuracy in both internal validation and external test sets. This methodology supports patient risk stratification and informs the development of personalized, optimized treatment strategies.
- Citation: Han W, Cheng DY, He QW, Wang SH, Gong SJ, Chen Y, Yang YP. Deep learning-based multimodal model for predicting on-treatment histological outcomes in chronic hepatitis B-associated advanced liver fibrosis. World J Gastroenterol 2026; 32(15): 116679
- URL: https://www.wjgnet.com/1007-9327/full/v32/i15/116679.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i15.116679
Liver fibrosis can progress to cirrhosis and hepatocellular carcinoma (HCC), both of which significantly contribute to liver-related morbidity worldwide. Chronic hepatitis B (CHB) infection remains the predominant cause of liver fibrosis[1,2]. Extensive research has demonstrated that effective antiviral therapy can promote fibrosis regression[3,4]. However, treatment responses exhibit marked inter-individual heterogeneity, nearly half of patients do not achieve fibrosis regression and may even experience notable disease progression[5,6]. Therefore, it is essential to identify individuals with liver fibrosis who are at a heightened risk of disease progression and unlikely to benefit from antiviral therapy alone.
Despite advancements in noninvasive testing, liver biopsy with histopathological examination remains the gold standard for staging fibrosis and evaluating disease activity[7,8]. This preeminence is attributed to its ability to provide direct structural insights into liver architecture, including detailed visualization of fibrous septa morphology and distribution, as well as key pathological features, such as hepatocyte injury and inflammatory infiltration. Furthermore, several routinely employed scoring systems, including the Knodell histological activity index, Scheuer, Ishak, and Metavir systems[9-12], support simultaneous evaluation of inflammatory activity and fibrosis stage, thereby informing tailored clinical management. Nonetheless, conventional assessments are limited by interobserver variability[13] and by their reliance on semi-quantitative methodologies, which may overlook subtle continuous changes[14]. Importantly, beyond macrostructural changes detectable by manual assessment, complex textural and spatial patterns within pathological images remain underutilized.
Traditional pathology has relied on microscopic examination of tissue sections. However, with the rapid advancement of digital pathology, the field is undergoing a profound transformation. High-resolution whole-slide images (WSIs) facilitate improved data acquisition and storage, establishing a robust foundation for technological innovation[15]. The advancement of artificial intelligence (AI) has enabled the exploration of critical information embedded within WSIs using deep learning (DL). Convolutional neural networks (CNNs) can autonomously identify intricate and discriminative features from extensive digitized WSI datasets, revealing latent patterns associated with disease biology and progression that may elude human observation. In liver pathology, DL shows substantial promise for liver fibrosis staging, nodular lesion classification, and risk prediction for HCC development[16-20]. However, there is a notable lack of predictive algorithmic models for forecasting histological outcomes in longitudinal CHB-related liver fibrosis cohorts following targeted antiviral therapy.
We hypothesize that DL-extracted features from pretreatment liver biopsy images contain prognostic information regarding histological responses to antiviral therapy. Accordingly, this study aims to develop and validate a DL-based model utilizing pretreatment histopathological images and clinical features to predict histological outcomes in patients with CHB-related advanced liver fibrosis. The results seek to demonstrate that DL can transform static pathological snapshots into dynamic prognostic tools, thereby supporting the advancement of personalized and precise approaches to the management of liver fibrosis.
The patients were enrolled from a multicenter randomized controlled clinical trial (NCT01965418). The trial’s design, eligibility criteria, and outcomes have been reported elsewhere[5,6,21]. Briefly, this two-stage study recruited patients with CHB-related liver fibrosis who met the following criteria: Age of ≥ 18 years, treatment-naive CHB, and Ishak fibrosis score of ≥ 3 points. Participants were randomly divided into two groups: One received entecavir plus placebo, and the other received entecavir combined with Biejia-Ruangan tablets. Following a 72-week double-blind treatment period, liver fibrosis reversal was evaluated. An additional open-label extension lasting 168 weeks assessed the cumulative rates of HCC and decompensated cirrhosis between the groups.
Fourteen medical institutions participated in this trial: The Fifth Medical of Chinese PLA General Hospital, First Affiliated Hospital of Wenzhou Medical University, First Affiliated Hospital of Zhengzhou University, Fuzhou Infectious Diseases Hospital, Department of Clinical and Translational Medicine, Fuyang 2nd People’s Hospital, Third Military Medical University, 88th Hospital of PLA, Guangzhou 8th People’s Hospital, Shanghai Public Health Clinical Center, Affiliated Hospital of Chengdu University of Traditional Chinese Medicine, Affiliated Traditional Chinese Medicine Hospital of Southwest Medical University, Traditional Chinese Medicine Hospital of Chongqing, and Tianjin Second People’s Hospital. Patient distribution and histopathological image acquisition details across participating centers are present in Supplementary Table 1.
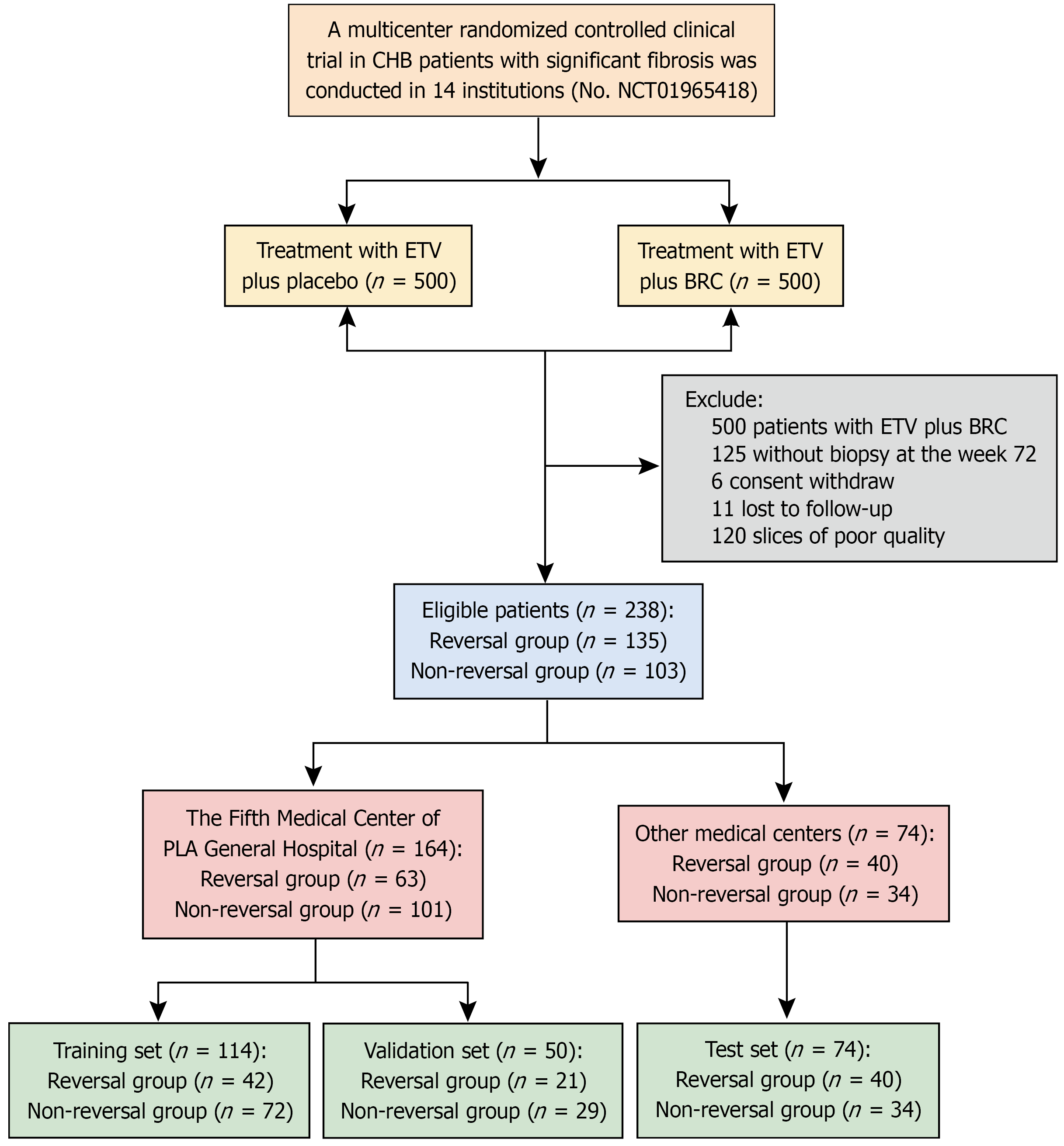
In total, 1000 patients were included in this study, and 762 patients were excluded for various reasons: 500 had received entecavir plus Biejia-Ruangan tablets, 125 did not undergo liver biopsy at week 72, 6 withdrew informed consent, 11 were lost to follow-up, and 120 had poor-quality pathological slides. As a result, 238 patients with baseline liver biopsy specimens were included in the final analysis (Figure 1).
Patients from the host institution (The Fifth Medical of Chinese PLA General Hospital, n = 164) were randomly split into training and internal validation sets at a 7:3 ratio using simple random sampling. Randomization was performed in R (version 4.3.1), with a fixed seed (seed = 42) to ensure reproducibility. Each patient was assigned a unique identification number, and a sampling function was employed to randomly select 114 patients for the training set, while the remaining 50 were allocated to the validation set.
All remaining patients from the other institutions were combined to form an external test cohort (n = 74), instead of evaluating each center separately. This strategy provided a more stable and representative assessment of model per
Ultrasound-guided liver biopsies were performed at baseline and after 72 weeks of treatment according to a standard protocol, using a 16-G quick-cut or Menghini needle (Allegiance Corporation, IL, United States). An adequate specimen was defined as one measuring at least 2.0 cm in length and containing 11 or more complete portal tracts[22]. Two experienced hepatopathologists independently conducted all histological evaluations, blinded to clinical information and experimental group allocation; any discrepancies were resolved by consensus.
The histological assessment focused on two key components[23]: (1) Fibrosis staging using the modified Ishak system (grades: 0 for no fibrosis, 1-2 for mild fibrosis, 3-4 for progressive fibrosis, 5-6 for cirrhosis); and (2) Grading inflammatory activity with the modified histologic activity index by Ishak (scores: 1-4 for mild inflammation, 5-8 for moderate inflammation, ≥ 9 for severe inflammation). Fibrosis reversal was defined as a decrease of at least one point in the Ishak fibrosis score at week 72 compared to baseline following treatment; all other patients were classified as non-reversers.
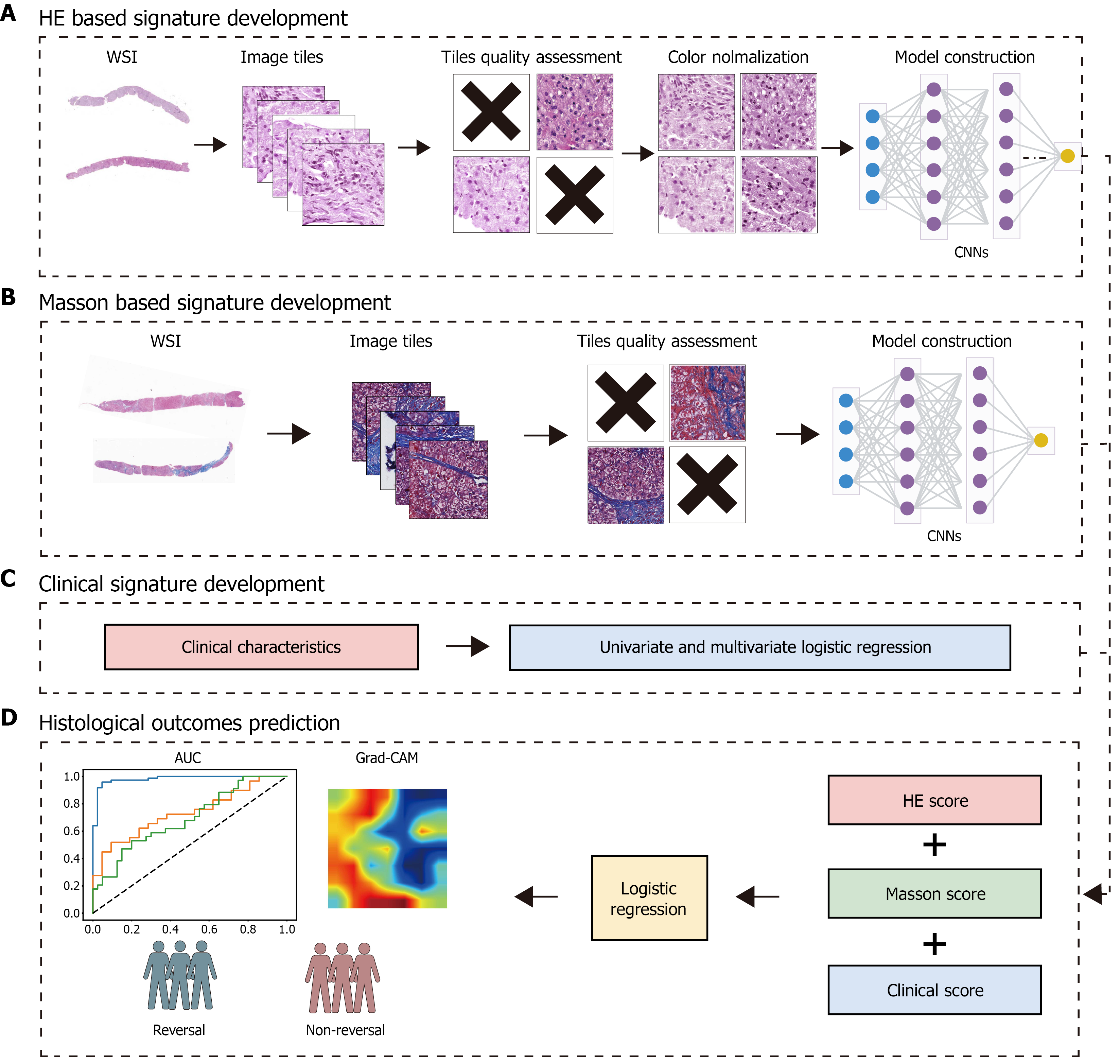
The hematoxylin and eosin (HE)-stained slides were scanned using a 3DHistech scanner (Hungary) with a 40 × objective lens (0.27 μm/pixel) in brightfield mode and stored as MRXS files. Masson’s trichrome-stained slides were digitized with a NanoZoomer scanner (Hamamatsu, Japan) at 20 × magnification (0.45 μm/pixel) and saved in NDPI format. All scanning parameters, including color calibration and focus, were set according to the manufacturer’s standard protocols. Representative images and the overall digitization workflow are presented in Figure 2.
WSIs of pathological liver biopsy specimens were segmented into non-overlapping image tiles of 512 × 512 pixels at 20 × magnification, which was selected as a unified working resolution to ensure consistency across staining modalities and to provide sufficient spatial detail for robust tissue-level analysis. Tissue regions were distinguished from the background using Otsu thresholding, and only tiles containing more than 60% tissue were retained. No predefined regions of interest were used; instead, tiles were sampled uniformly across the tissue to minimize selection bias. To enhance the model robustness and decrease overfitting, standard data augmentation techniques including random rotations and horizontal and vertical flips, were employed during training (Figure 2).
Differences in scanners and staining procedures can lead to substantial color variations in the WSIs, making stain normalization an essential preprocessing step. In this study, the Vahadane stain normalization method was employed. This technique relies on non-negative matrix factorization. Compared with commonly used normalization approaches, such as Macenko or Reinhard, the Vahadane method better preserves structural features by separating stain concentration from tissue morphology[24]. Hence, each image was split into matrices for stain color and density, and the stain elements from the source image were aligned with those of a selected reference image. The process generated normalized images with uniform coloring across all slides (Figure 2, Supplementary Figure 1).
The HE model was constructed using ResNet50 with pretrained weights in PyTorch (v2.8.0). Color-normalized image patches (512 × 512 pixels) were used as inputs. The model was trained using the Adam optimizer (batch size 256, initial learning rate 0.01), applying cosine annealing for learning rate decay and a dropout rate of 0.2. Multiple architectures (ResNet18, Inception_v3, ShuffleNet_v2, CrossFormer, and SimpleViT) were compared, and the optimal configuration was selected based on performance. The Masson model employed the ShuffleNet_v2 CNN architecture with pre-trained weights, also in PyTorch. Training followed identical settings, and the best-performing configuration among ResNet18, ResNet50, Inception_v3, CrossFormer, and SimpleViT was adopted. Model training was performed using an NVIDIA RTX 4090 graphics processing unit.
To build the clinical model, candidate variables were considered, including age, sex, smoking and drinking history, body mass index, platelet count (PLT), prothrombin time, alanine aminotransferase, aspartate aminotransferase, alkaline phosphatase (ALP), γ-glutamyltransferase (GGT), total bilirubin (TBiL), albumin, alpha-fetoprotein, hepatitis B virus DNA level, hepatitis B surface antigen level, liver stiffness measurement (LSM), aspartate aminotransferase to platelet index, and fibrosis index based on 4 factors. To prevent data leakage, all variable selection occurred solely within the training cohort. Univariate logistic regression was performed first, and variables with P < 0.05 were entered into a multivariate logistic regression model. Variables that remained significant were identified as independent predictors for the final clinical prediction model.
To combine the predictive information originating from the HE, Masson and clinical modalities, we implemented a probability-level multimodal fusion strategy. For each WSI, slide-level scores were generated by averaging prediction scores from all patches using the trained classifiers. Simultaneously, patients’ clinical scores were calculated via multivariate logistic regression based on independently identified clinical predictors. The HE, Masson, and clinical scores served as inputs to a logistic regression fusion model that determined optimal weights and combined the three modalities. This fusion strategy preserved each modality’s unique contribution while providing an interpretable, practical method for synthesizing heterogeneous predictive data into a single output. A cutoff value of 0.576 was set using the maximum Youden index; this threshold was consistently applied to evaluate performance in the validation and test cohorts.
To enhance model interpretability, Gradient-weighted Class Activation Mapping (Grad-CAM) was used to visualize the outputs in the corresponding images[25]. After completing model training and obtaining the optimal weights, they were applied to generate Grad-CAM visualizations from the model’s final convolutional layer.
The model’s performance on the validation and test datasets was assessed using multiple metrics: Area under the receiver operating characteristic curve (AUC), sensitivity, specificity, positive predictive value, and negative predictive value. The confidence interval (CI) for the AUC was calculated through 1000 bootstrap resampling iterations. Moreover, the model’s accuracy was analyzed using confusion matrices.
Continuous variables are recorded as mean ± SD or median (Q1-Q3) based on data distribution. Categorical variables are expressed as n (%). Skewed laboratory values for serum hepatitis B virus DNA and hepatitis B surface antigen were log-transformed. Univariate and multivariate logistic regression analyses were performed to screen clinical features. The Mann-Whitney U-test was used to compare continuous variables between two groups, whereas Spearman’s correlation was used to assess the association between two continuous variables. The χ2 test was applied to evaluate relationships between categorical variables. All statistical tests were two-sided, and P values < 0.05 were considered statistically significant. Data analyses were conducted using R (version 4.3.1; R Foundation for Statistical Computing) and Python (version 3.9.10).
A total of 238 patients were enrolled in our study, comprising 135 in the reversal group and 103 in the non-reversal group (Table 1). Clinical features and WSIs were collected from 114 cases comprising the training set for model development. An internal validation set of 50 cases was used for parameter optimization and internal validation, while an external test set of 74 cases was used for external validation (Figure 1). Baseline characteristics of the patients are summarized in Table 1. The mean ages of patients from the host institution and external set were 42.85 ± 10.06 years and 41.96 ± 10.21 years, respectively (Table 1). Median body mass index values were 23.53 for the host institution cohort and 23.03 for the external cohort (Table 1). Both cohorts exhibited a male predominance (Table 1).
| Characteristics | Training and validation sets | Test set | ||||||
| Overall | Reversal | Non-reversal | P value | Overall | Reversal | Non-reversal | P value | |
| Demography | ||||||||
| Age (years) | 42.85 ± 10.06 | 43.53 ± 11.50 | 42.44 ± 9.10 | 0.52 | 41.96 ± 10.21 | 37 (34.50-42) | 47 (39-50.75) | 0.001 |
| Gender | 0.49 | 0.09 | ||||||
| Male | 104 (63.80) | 37 (59.68) | 67 (66.34) | 55 (75.34) | 33 (84.62) | 22 (64.71) | ||
| Female | 59 (36.20) | 25 (40.32) | 34 (33.66) | 18 (24.66) | 6 (15.38) | 12 (35.29) | ||
| Drinks | 30 (18.40) | 10 (16.13) | 20 (19.80) | 0.70 | 5 (6.85) | 2 (5.13) | 3 (8.82) | 0.66 |
| Smoke | 33 (20.25) | 10 (16.13) | 23 (22.77) | 0.41 | 7 (9.59) | 4 (10.26) | 3 (8.82) | 1 |
| BMI (kg/m2) | 23.53 (21.15-25.41) | 23.47 (21.72-25.38) | 23.63 (20.96-25.43) | 0.85 | 23.03 (21.51-24.82) | 23.18 (21.14-24.56) | 23.03 (21.72-25) | 0.54 |
| Laboratory tests | ||||||||
| PLT (109/L) | 165.50 (120-198.50) | 177.50 (132-209.50) | 164 (115-187) | 0.02 | 153.73 ± 50.19 | 160.13 ± 49.54 | 144.68 ± 50.11 | 0.19 |
| ALB (g/L) | 42 (39.45-45) | 42 (39.78-44) | 42 (39-45) | 0.62 | 42.40 ± 4.60 | 42.50 ± 3.65 | 42.28 ± 5.55 | 0.84 |
| PT (seconds) | 12.70 (11.70-13.70) | 12.75 (11.70-13.57) | 12.6 (11.90-14) | 0.45 | 12.30 ± 1.60 | 12.60 ± 1.54 | 12.01 ± 1.63 | 0.11 |
| ALT (IU/L) | 51 (32.50-103) | 54 (27.25-107.25) | 50 (36-100) | 0.70 | 47 (30-90) | 55 (32-92) | 42 (29.25-66.60) | 0.48 |
| AST (IU/L) | 44 (29-73) | 41 (28.20-73.50) | 45 (30-69) | 0.50 | 34 (27-69) | 33 (27.50-74.50) | 34.50 (26-65.75) | 0.69 |
| ALP (U/L) | 81.50 (67-104) | 76 (65.75-90.25) | 89 (68-114.50) | 0.01 | 76 (60.75-90.25) | 72 (60-89.50) | 80.50 (67-90.25) | 0.67 |
| GGT (U/L) | 38 (22-70.75) | 32 (21-54) | 45 (22-75.50) | 0.08 | 32.5 (20-67) | 30 (20-48) | 34.5 (20-71.50) | 0.48 |
| TBiL (μmol/L) | 13 (10.15-17) | 12 (9.85-16.58) | 13.7 (11-17) | 0.22 | 15.40 (12.23-21.10) | 15.70 (13.40-20.70) | 16 (12.16-23.28) | 0.68 |
| AFP (ng/mL) | 5 (2.34-9) | 4.88 (2.30-9.20) | 5 (2.34-8.88) | 0.75 | 5.2 (2.90-11.80) | 4.51 (2.59-15.92) | 5.38 (3.14-9.51) | 0.47 |
| Fibrosis staging models | ||||||||
| LSM (kPa) | 10.6 (6.35-16.30) | 8.85 (5.80-12.80) | 12.30 (7.60-18.50) | 0.002 | 8.3 (6.20-15.30) | 7.6 (6.15-14) | 9.8 (6.40-16.15) | 0.270 |
| APRI | 0.73 (0.41-1.39) | 0.58 (0.39-1.18) | 0.76 (0.46-1.51) | 0.08 | 0.74 (0.40-1.20) | 0.77 (0.37-1.19) | 0.7 (0.47-1.29) | 0.70 |
| FIB4 | 1.63 (1.10-2.67) | 1.53 (0.97-2.39) | 1.69 (1.15-2.83) | 0.27 | 1.57 (1.08-2.54) | 1.43 (1.03-2.02) | 1.8 (1.17-3) | 0.07 |
| HBV markers | ||||||||
| HBsAg1 | 3.46 (3.09-4.01) | 3.59 (3.18-4.11) | 3.45 (2.98-3.85) | 0.21 | 3.5 (3.06-4.04) | 3.56 ± 0.91 | 3.53 ± 0.70 | 0.86 |
| HbeAg | 1.00 | 0.20 | ||||||
| Positive | 101 (61.96) | 38 (61.29) | 63 (62.38) | 37 (50.68) | 23 (58.97) | 14 (41.18) | ||
| Negative | 62 (38.04) | 24 (38.71) | 38 (37.62) | 36 (49.32) | 16 (41.03) | 20 (58.82) | ||
| HBV DNA1 | 5.98 (4.75-7.33) | 6.67 (4.89-7.78) | 5.71 (4.75-7.17) | 0.13 | 6.28 (5.04-7.75) | 6.91 (5.23-8.01) | 5.52 (4.95-6.63) | 0.07 |
| Histology | ||||||||
| HAI | 0.04 | 0.34 | ||||||
| 1-4 | 32 (19.63) | 8 (12.90) | 24 (23.76) | 18 (24.66) | 8 (20.51) | 10 (29.41) | ||
| 5-8 | 92 (56.44) | 42 (67.74) | 50 (49.50) | 38 (52.05) | 20 (51.28) | 18 (52.94) | ||
| 9-12 | 38 (23.31) | 11 (17.74) | 27 (26.73) | 16 (21.92) | 11 (28.21) | 5 (14.71) | ||
| 13-18 | 1 (0.61) | 1 (1.61) | 0 (0) | 1 (1.37) | 0 (0) | 1 (2.94) | ||
| Ishak score | 0.73 | 0.27 | ||||||
| 3 | 47 (28.83) | 21 (33.87) | 26 (25.74) | 24 (32.88) | 15 (38.46) | 9 (26.47) | ||
| 4 | 24 (14.72) | 8 (12.90) | 16 (15.84) | 12 (16.44) | 8 (20.51) | 4 (11.76) | ||
| 5 | 31 (19.02) | 11 (17.74) | 20 (19.80) | 17 (23.29) | 6 (15.38) | 11 (32.35) | ||
| 6 | 61 (37.42) | 22 (35.48) | 39 (38.61) | 20 (27.40) | 10 (25.64) | 10 (29.41) | ||
Within the primary center cohort, most baseline characteristics did not differ significantly between the reversal and non-reversal groups. However, notable differences were identified for PLT, ALP, LSM, and histologic activity index. Specifically, the non-reversal group demonstrated lower PLT levels (164 × 109/L vs 177.50 × 109/L), higher ALP levels (89 U/L vs 76 U/L), elevated LSM (12.30 kPa vs 8.85 kPa), and lower proportions of moderate inflammatory activity (49.50% vs 67.74%; Table 1). In the external validation cohort, patients in the non-reversal group were significantly older than those in the reversal group (47 years vs 37 years; Table 1).
Univariate logistic regression revealed significant associations between prothrombin time [odds ratio (OR) = 1.333, 95%CI: 1.005-1.766, P = 0.046], ALP (OR = 1.016, 95%CI: 1.000-1.031, P = 0.044), LSM (OR = 1.092, 95%CI: 1.025-1.163, P = 0.007) and the outcome (Table 2). Therefore, these variables were included in a multivariate logistic regression model, where only LSM remained a significant independent predictor (OR = 1.073, 95%CI: 1.003-1.149, P = 0.041) of outcome (Table 2). Accordingly, LSM was incorporated into the final clinical model, which achieved an AUC of 0.658 (95%CI: 0.502-0.812) in the validation set 0.588 (95%CI: 0.456-0.716) in the test cohort (Figure 3A and B, Tables 3 and 4).
| Characteristics | Univariate analysis | Multivariate analysis | ||
| OR (95%CI) | P value | OR (95%CI) | P value | |
| PT (seconds) | 1.333 (1.005-1.766) | 0.046 | 1.200 (0.896-1.609) | 0.222 |
| ALP (U/L) | 1.016 (1.000-1.031) | 0.044 | 1.006 (0.990-1.023) | 0.462 |
| LSM (kPa) | 1.092 (1.025-1.163) | 0.007 | 1.073 (1.003-1.149) | 0.041 |
| Age (years) | 1.111 (0.500-2.470) | 0.796 | ||
| Sex | 1.008 (0.464-2.190) | 0.983 | ||
| Drinking | 0.975 (0.371-2.562) | 0.959 | ||
| Smoking | 0.700 (0.262-1.872) | 0.477 | ||
| BMI (kg/m2) | 1.027 (0.931-1.133) | 0.591 | ||
| PLT (109/L) | 0.996 (0.989-1.002) | 0.169 | ||
| ALB (g/L) | 1.011 (0.924-1.106) | 0.809 | ||
| ALT (IU/L) | 1.000 (0.996-1.005) | 0.931 | ||
| AST (IU/L) | 0.999 (0.992-1.006) | 0.793 | ||
| GGT (U/L) | 0.999 (0.997-1.001) | 0.432 | ||
| TBil (μmol/L) | 1.053 (0.983-1.129) | 0.142 | ||
| AFP (ng/mL) | 1.006 (0.993-1.018) | 0.376 | ||
| APRI | 1.001 (0.716-1.397) | 0.997 | ||
| FIB4 | 1.058 (0.830-1.350) | 0.648 | ||
| HBsAg1 | 0.783 (0.449-1.366) | 0.389 | ||
| HBV DNA1 | 0.833 (0.651-1.066) | 0.146 | ||
| Metrics | Clinical | HE | Masson | HE + Masson | Multimodal model |
| AUC (mean) | 0.658 | 0.657 | 0.727 | 0.732 | 0.741 |
| AUC (95%CI) | 0.502-0.812 | 0.494-0.800 | 0.575-0.857 | 0.580-0.862 | 0.588-0.869 |
| Sensitivity | 0.586 | 0.828 | 0.724 | 0.724 | 0.655 |
| Specificity | 0.667 | 0.381 | 0.381 | 0.619 | 0.667 |
| PPV | 0.708 | 0.649 | 0.618 | 0.724 | 0.731 |
| NPV | 0.538 | 0.615 | 0.500 | 0.619 | 0.583 |
| Metrics | Clinical | HE | Masson | HE + Masson | Multimodal model |
| AUC (mean) | 0.588 | 0.615 | 0.676 | 0.690 | 0.694 |
| AUC (95%CI) | 0.456-0.716 | 0.484-0.741 | 0.547-0.799 | 0.564-0.812 | 0.570-0.815 |
| Sensitivity | 0.412 | 0.882 | 0.706 | 0.647 | 0.647 |
| Specificity | 0.750 | 0.250 | 0.450 | 0.525 | 0.525 |
| PPV | 0.583 | 0.500 | 0.522 | 0.537 | 0.537 |
| NPV | 0.600 | 0.714 | 0.643 | 0.636 | 0.636 |
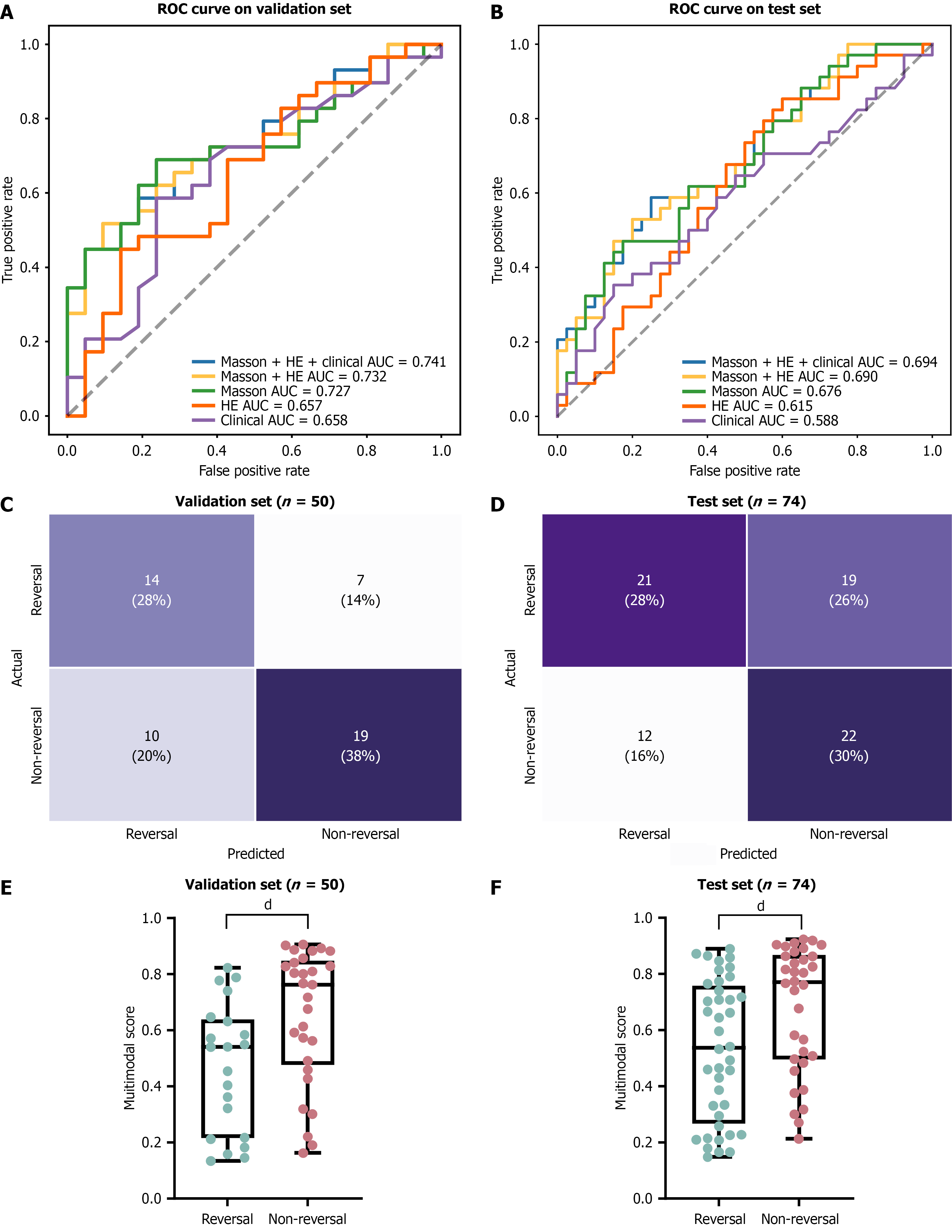
First, models were independently trained using HE- and Masson-stained WSIs, and a range of network architectures were systematically compared. Subsequently, a comprehensive assessment of the key performance metrics was performed, including AUC, sensitivity, specificity, negative predictive value, and positive predictive value, to identify the strongest model configuration (Tables 3 and 4). ResNet50 emerged as the optimal architecture for HE images, while ShuffleNet_v2 performed best for Masson-stained images. The final HE model achieved AUCs of 0.657 (95%CI: 0.494-0.800) and 0.615 (95%CI: 0.484-0.741) on the validation and test sets, respectively (Figure 3A and B, Tables 3 and 4). The Masson model yielded AUCs of 0.727 (95%CI: 0.575-0.857) and 0.676 (95%CI: 0.547-0.799) for the same sets (Figure 3A and B, Tables 3 and 4), slightly outperforming the HE model, which aligns with the established clinical value of Masson staining for liver fibrosis diagnosis and prognosis assessments[26].
Subsequently, the HE and Masson models were combined into a fusion model, yielding superior performance compared with either stain alone, achieving AUCs of 0.732 (95%CI: 0.580-0.862) and 0.690 (95%CI: 0.564-0.812) for the validation and test sets, respectively (Figure 3A and B, Tables 3 and 4). Given that predictions were generated at the slide level, an additional assessment was conducted to determine whether the number of slides per patient influenced model performance. The analysis showed that the slide count did not significantly impact the performance of the HE or Masson models across the training, validation, and test cohorts (Supplementary Figure 2).
To fully leverage patient-specific features, individual unimodal models were integrated at the probability level, and the resulting multimodal models were comprehensively evaluated in the validation and test cohorts. The multimodal model achieved an AUC of 0.741 (95%CI: 0.588-0.869) in the validation cohort and 0.694 (95%CI: 0.570-0.815) in the test cohort (Figure 3A and B, Tables 3 and 4), demonstrating favorable discriminative ability across datasets. In the validation set, the integrated model accurately identified 19 out of 29 non-reversal patients, while in the test set, it correctly classified 22 out of 34 non-reversal patients, with sensitivities of 0.655 and 0.647, and specificities of 0.667 and 0.525, respectively (Figure 3C and D; Tables 3 and 4).
The multimodal model generated a predictive probability for each patient ranging from 0 to 1, corresponding to the estimated likelihood of non-reversal to therapy. Utilizing the Youden index, a threshold of 0.576 was established; patients with scores ≥ 0.576 were considered to have a higher probability of non-reversal. Analysis of the continuous score distributions revealed that the non-reversal group consistently exhibited higher model scores than the reversal group across both validation and test sets (Figure 3E and F).
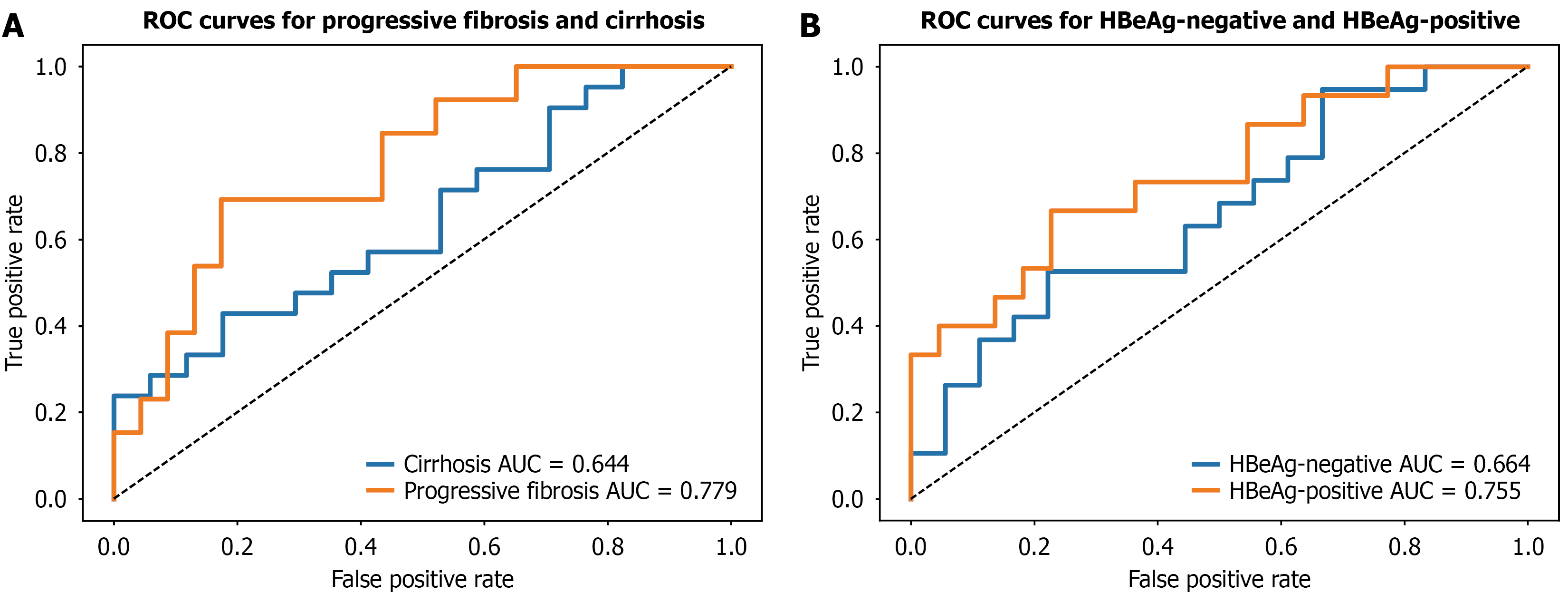
The test cohort was stratified into two subgroups based on baseline Ishak scores to facilitate comparative performance analysis of patients with advanced fibrosis (Ishak score 3-4) and those with cirrhosis (Ishak score 5-6). The model demonstrated strong predictive accuracy in the advanced fibrosis group, achieving an AUC of 0.779 (95%CI: 0.609-0.927; Figure 4A). Conversely, its performance in the cirrhosis subgroup was modest, with an AUC of 0.644 (95%CI: 0.461-0.816; Figure 4A). Additionally, the cohort was subdivided by hepatitis B e antigen (HBeAg) status to further evaluate the model’s predictive capacity. The combined model exhibited superior predictive ability in the HBeAg-positive subgroup (AUC: 0.755, 95%CI: 0.580-0.904; Figure 4B) compared to the HBeAg-negative subgroup (AUC: 0.664, 95%CI: 0.470-0.829; Figure 4B).
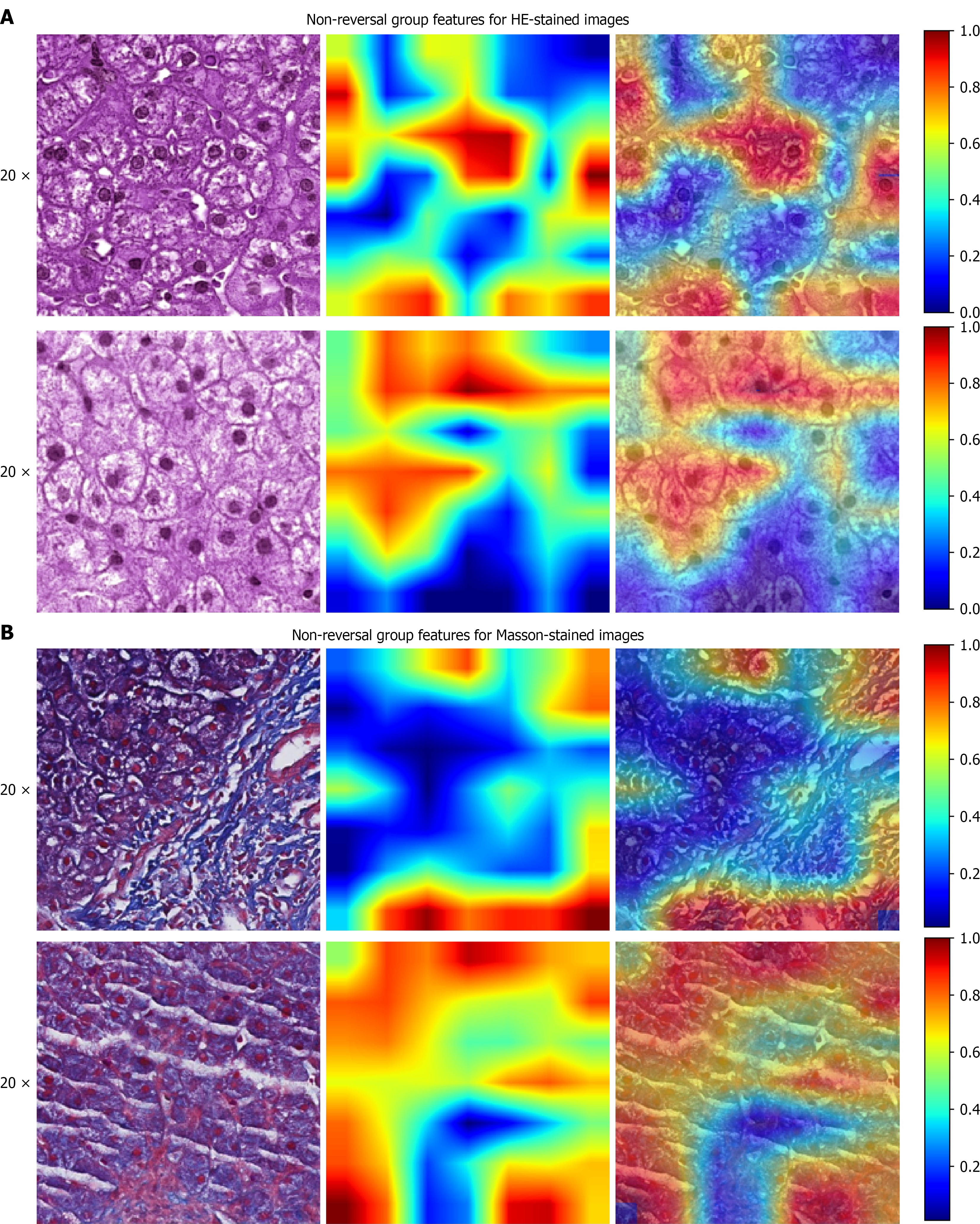
To enhance the model’s explainability, Grad-CAM was employed for visual analysis at the tile level to identify the main histopathological features associated with non-reversal of fibrosis. The model visualization outputs and representative pathological staining images from the non-reversal group are presented in Figure 5. Deeper red hues indicate areas with a higher probability of non-reversal characteristics. Key features seen in the red-highlighted regions from the HE stains included: (1) Disorganized hepatic cords: Hepatocytes lost their typical radial or plate-like arrangement, and instead appeared fragmented, scattered, or formed pseudo-glandular structures; (2) Ballooning degeneration and nuclear atypia of hepatocytes: Hepatocyte cytoplasm exhibited rarefaction and vacuolation, with some nuclei displaying hyper
In Masson staining images, high-risk regions predominantly corresponded to: (1) Thick, continuously bridging fibrous septa with parallel, layered, or laminar patterns that created coarse mesh-like networks; (2) Fine reticular fibers were found between the coarse septa and hepatocyte plates, indicating persistent perisinusoidal extracellular matrix deposition; and (3) Ductular reactions and inflammatory cell infiltration: Evident as bead-like arrangements of nuclei and duct-structured formation within or next to fibrous septa (Figure 5B).
Overall, models based on HE stains primarily capture hepatocellular states and parenchymal features, whereas those based on Masson stains tend to emphasize fibrous septa and alterations in portal tracts. The key histologic features, such as the condition of hepatocytes, architecture of fibrous septa, and the presence of intra-fibrotic inflammation and ductular reactions, may serve as potential prognostic markers for liver fibrosis.
This proof-of-concept study devised an AI driven model to predict histological outcomes following antiviral therapy in CHB patients with advanced liver fibrosis, using clinical features and digital pathology images. The integration of clinical and pathological DL models demonstrated superior performance compared with individual assessments in identifying patients unlikely to achieve fibrosis reversal with antiviral therapy alone. This multimodal model demonstrated robust predictive performance across internal and external test cohorts, further supporting its generalizability and suitability for practical applications.
Emerging evidence supports the view that liver fibrosis is a dynamic and potentially reversible process. As such, etiology-specific treatment forms the cornerstone of fibrosis reversal. Nonetheless, clinical observations indicate that, despite successful etiological control, a subset of patients fail to achieve fibrosis regression, with some experiencing continued progression toward liver-related endpoints[27]. This underscores the importance of identifying individuals at higher risk. Current noninvasive diagnostic modalities, including LSM and serum biomarker panels, are often influenced by inflammation and hemodynamic variability, leading to discrepancies with histologically confirmed regression, particularly during antiviral therapy[28,29]. Efforts to establish baseline predictors, such as angiopoietin-2 levels, have shown limited prognostic utility and fail to fully capture the complex tissue remodeling involved in fibrosis reversal[30].
In the current study, LSM emerged as the only clinical parameter associated with post-intervention fibrosis outcomes; however, it demonstrated modest predictive value, with AUCs of 0.658 and 0.588 in internal and external validation sets, respectively. Similarly, another study found that longitudinal multimodal assessments integrating LSM and biochemical markers provided only limited predictive accuracy of liver fibrosis regression after 78 weeks and 260 weeks of treatment, as validated by serial liver biopsies[31], highlighting a fundamental gap in existing approaches.
Advances in AI and radiomics have markedly improved the characterization of fibrosis severity and clinical risk using magnetic resonance imaging-derived features, automated image segmentation, and machine learning classifiers[32,33]. However, these efforts have largely focused on fibrosis staging or event prediction rather than on the more intricate biological process of fibrosis regression. Importantly, few studies have incorporated histological spatial information despite its direct relevance to architectural remodeling that accompanies fibrosis improvement. To address this important gap, the current study integrated clinical features with DL-based histopathological staining analyses to predict fibrosis regression. The combined model achieved an AUC of 0.694 on the test set, effectively identifying patients unlikely to experience fibrosis reversal despite etiological treatment. For these individuals, more proactive therapeutic strategies, such as the concomitant use of antifibrotic traditional medicines or closer clinical monitoring, may help maximize clinical benefits.
Our integrated model demonstrated superior predictive performance for Ishak stage 3-4 fibrosis, characterized by more dynamic and reversible histological features that the model could easily discern. Nonetheless, sustained efficacy in cirrhosis cases (Ishak 5-6) confirmed that significant regressive potential persisted within the cirrhotic microenvironment. The significant performance difference between the HBeAg-positive and HBeAg-negative cohorts further suggests divergent immunovirological mechanisms, necessitating distinct clinical management strategies.
CNNs, a major type of DL algorithm widely adopted in medical image analysis due to their capacity to capture local features and decipher complex image structures[34]. Models like ResNet and ShuffleNet_v2 have demonstrated robust stability and strong predictive performance in medical image classification tasks[35-37]. In the current comparative analysis, the CNNs (ResNet50, ResNet18, Inception_v3, and ShuffleNet_v2) consistently outperformed the transformer-based architectures (CrossFormer and SimpleViT). This is likely attributable to the intrinsic inductive bias of CNNs, which are particularly well-suited to capturing the localized texture and morphological patterns of histopathological images. In contrast, transformer models typically require substantially larger datasets to fully leverage their global attention mechanisms. Given the relatively small dataset and the strong dependence of liver histopathology on fine-grained local features, transformer-based models were likely limited by their unstable training dynamics and weaker generalization.
DL models are often regarded as “black boxes” due to their opaque, data-driven algorithms, making it difficult to interpret how they reach decisions[38]. To address this, visualization tools are commonly used to improve model explainability[25]. Accordingly, in this study, Grad-CAM was employed to summarize the image regions most critical to the probability scoring process[39,40]. By evaluating the tile regions highlighted by the model, the model was found to primarily focus on the state of parenchymal hepatocytes, the morphology of fibrous septa, and the characteristics of their local microenvironment.
This study presents several limitations that warrant consideration. First, although the multimodal model demonstrated a promising predictive performance, its accuracy diminished from the internal validation cohort to the external test cohort, suggesting suboptimal generalizability. This performance decline may be attributable to the relatively limited sample size, inter-center variations in staining protocols, and the inherent heterogeneity of multicenter digital pathology datasets. Second, the biological processes underlying image-derived DL features remain incompletely understood. Incorporating spatial transcriptomics or single-cell sequencing into future research could clarify the molecular substrates driving the distinct histologic patterns of fibrosis regression. Third, while our model leverages DL to quantify subtle histological features, it relies on retrospective biopsy data; therefore, prospective validation using larger, ethnically diverse cohorts is necessary before clinical application. Finally, although the current framework integrates clinical and pathological modalities, expanding the model to include longitudinal noninvasive biomarkers or radiologic signatures may further improve its robustness and clinical utility.
This multimodal model provides an effective tool for predicting posttreatment histological outcomes in patients with advanced CHB-related liver fibrosis. It has the potential to identify patients less likely to experience fibrosis reversal, enabling more proactive healthcare management, including closer monitoring, early adjunctive antifibrotic therapy, or enrollment in relevant clinical trials.
| 1. | GBD 2023 Causes of Death Collaborators. Global burden of 292 causes of death in 204 countries and territories and 660 subnational locations, 1990-2023: a systematic analysis for the Global Burden of Disease Study 2023. Lancet. 2025;406:1811-1872. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 151] [Cited by in RCA: 173] [Article Influence: 173.0] [Reference Citation Analysis (1)] |
| 2. | Moon AM, Singal AG, Tapper EB. Contemporary Epidemiology of Chronic Liver Disease and Cirrhosis. Clin Gastroenterol Hepatol. 2020;18:2650-2666. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 923] [Cited by in RCA: 836] [Article Influence: 139.3] [Reference Citation Analysis (6)] |
| 3. | Chang TT, Liaw YF, Wu SS, Schiff E, Han KH, Lai CL, Safadi R, Lee SS, Halota W, Goodman Z, Chi YC, Zhang H, Hindes R, Iloeje U, Beebe S, Kreter B. Long-term entecavir therapy results in the reversal of fibrosis/cirrhosis and continued histological improvement in patients with chronic hepatitis B. Hepatology. 2010;52:886-893. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 842] [Cited by in RCA: 790] [Article Influence: 49.4] [Reference Citation Analysis (3)] |
| 4. | Marcellin P, Gane E, Buti M, Afdhal N, Sievert W, Jacobson IM, Washington MK, Germanidis G, Flaherty JF, Aguilar Schall R, Bornstein JD, Kitrinos KM, Subramanian GM, McHutchison JG, Heathcote EJ. Regression of cirrhosis during treatment with tenofovir disoproxil fumarate for chronic hepatitis B: a 5-year open-label follow-up study. Lancet. 2013;381:468-475. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1532] [Cited by in RCA: 1421] [Article Influence: 109.3] [Reference Citation Analysis (8)] |
| 5. | Ji D, Chen Y, Bi J, Shang Q, Liu H, Wang JB, Tan L, Wang J, Chen Y, Li Q, Long Q, Song L, Jiang L, Xiao G, Yu Z, Chen L, Wang X, Chen D, Li Z, Dong Z, Yang Y. Entecavir plus Biejia-Ruangan compound reduces the risk of hepatocellular carcinoma in Chinese patients with chronic hepatitis B. J Hepatol. 2022;77:1515-1524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 58] [Article Influence: 14.5] [Reference Citation Analysis (1)] |
| 6. | Rong G, Chen Y, Yu Z, Li Q, Bi J, Tan L, Xiang D, Shang Q, Lei C, Chen L, Hu X, Wang J, Liu H, Lu W, Chen Y, Dong Z, Bai W, Yoshida EM, Mendez-Sanchez N, Hu KQ, Qi X, Yang Y. Synergistic Effect of Biejia-Ruangan on Fibrosis Regression in Patients With Chronic Hepatitis B Treated With Entecavir: A Multicenter, Randomized, Double-Blind, Placebo-Controlled Trial. J Infect Dis. 2022;225:1091-1099. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 41] [Article Influence: 10.3] [Reference Citation Analysis (1)] |
| 7. | Sun Y, Zhou J, Wang L, Wu X, Chen Y, Piao H, Lu L, Jiang W, Xu Y, Feng B, Nan Y, Xie W, Chen G, Zheng H, Li H, Ding H, Liu H, Lv F, Shao C, Wang T, Ou X, Wang B, Chen S, Wee A, Theise ND, You H, Jia J. New classification of liver biopsy assessment for fibrosis in chronic hepatitis B patients before and after treatment. Hepatology. 2017;65:1438-1450. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 131] [Article Influence: 14.6] [Reference Citation Analysis (1)] |
| 8. | Tong XF, Wang QY, Zhao XY, Sun YM, Wu XN, Yang LL, Lu ZZ, Ou XJ, Jia JD, You H. Histological assessment based on liver biopsy: the value and challenges in NASH drug development. Acta Pharmacol Sin. 2022;43:1200-1209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 30] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 9. | Knodell RG, Ishak KG, Black WC, Chen TS, Craig R, Kaplowitz N, Kiernan TW, Wollman J. Formulation and application of a numerical scoring system for assessing histological activity in asymptomatic chronic active hepatitis. Hepatology. 1981;1:431-435. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2758] [Cited by in RCA: 2491] [Article Influence: 55.4] [Reference Citation Analysis (0)] |
| 10. | Scheuer PJ. Classification of chronic viral hepatitis: a need for reassessment. J Hepatol. 1991;13:372-374. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1293] [Cited by in RCA: 1222] [Article Influence: 34.9] [Reference Citation Analysis (4)] |
| 11. | Bedossa P, Poynard T. An algorithm for the grading of activity in chronic hepatitis C. The METAVIR Cooperative Study Group. Hepatology. 1996;24:289-293. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3413] [Cited by in RCA: 3115] [Article Influence: 103.8] [Reference Citation Analysis (1)] |
| 12. | Ishak K, Baptista A, Bianchi L, Callea F, De Groote J, Gudat F, Denk H, Desmet V, Korb G, MacSween RN. Histological grading and staging of chronic hepatitis. J Hepatol. 1995;22:696-699. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4060] [Cited by in RCA: 3796] [Article Influence: 122.5] [Reference Citation Analysis (4)] |
| 13. | Bedossa P, Dargère D, Paradis V. Sampling variability of liver fibrosis in chronic hepatitis C. Hepatology. 2003;38:1449-1457. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1571] [Cited by in RCA: 1403] [Article Influence: 61.0] [Reference Citation Analysis (5)] |
| 14. | Davison BA, Harrison SA, Cotter G, Alkhouri N, Sanyal A, Edwards C, Colca JR, Iwashita J, Koch GG, Dittrich HC. Suboptimal reliability of liver biopsy evaluation has implications for randomized clinical trials. J Hepatol. 2020;73:1322-1332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 384] [Cited by in RCA: 345] [Article Influence: 57.5] [Reference Citation Analysis (2)] |
| 15. | Aggarwal A, Bharadwaj S, Corredor G, Pathak T, Badve S, Madabhushi A. Artificial intelligence in digital pathology - time for a reality check. Nat Rev Clin Oncol. 2025;22:283-291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 30.0] [Reference Citation Analysis (1)] |
| 16. | Abdurrachim D, Lek S, Ong CZL, Wong CK, Zhou Y, Wee A, Soon G, Kendall TJ, Idowu MO, Hendra C, Saigal A, Krishnan R, Chng E, Tai D, Ho G, Forest T, Raji A, Talukdar S, Chin CL, Baumgartner R, Engel SS, Ali AAB, Kleiner DE, Sanyal AJ. Utility of AI digital pathology as an aid for pathologists scoring fibrosis in MASH. J Hepatol. 2025;82:898-908. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 14] [Article Influence: 14.0] [Reference Citation Analysis (1)] |
| 17. | Yu H, Sharifai N, Jiang K, Wang F, Teodoro G, Farris AB, Kong J. Artificial intelligence based liver portal tract region identification and quantification with transplant biopsy whole-slide images. Comput Biol Med. 2022;150:106089. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 14] [Article Influence: 3.5] [Reference Citation Analysis (2)] |
| 18. | Cheng N, Ren Y, Zhou J, Zhang Y, Wang D, Zhang X, Chen B, Liu F, Lv J, Cao Q, Chen S, Du H, Hui D, Weng Z, Liang Q, Su B, Tang L, Han L, Chen J, Shao C. Deep Learning-Based Classification of Hepatocellular Nodular Lesions on Whole-Slide Histopathologic Images. Gastroenterology. 2022;162:1948-1961.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 110] [Cited by in RCA: 82] [Article Influence: 20.5] [Reference Citation Analysis (1)] |
| 19. | Nakatsuka T, Tateishi R, Sato M, Hashizume N, Kamada A, Nakano H, Kabeya Y, Yonezawa S, Irie R, Tsujikawa H, Sumida Y, Yoneda M, Akuta N, Kawaguchi T, Takahashi H, Eguchi Y, Seko Y, Itoh Y, Murakami E, Chayama K, Taniai M, Tokushige K, Okanoue T, Sakamoto M, Fujishiro M, Koike K. Deep learning and digital pathology powers prediction of HCC development in steatotic liver disease. Hepatology. 2025;81:976-989. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 23] [Article Influence: 23.0] [Reference Citation Analysis (1)] |
| 20. | Yu Y, Wang J, Ng CW, Ma Y, Mo S, Fong ELS, Xing J, Song Z, Xie Y, Si K, Wee A, Welsch RE, So PTC, Yu H. Deep learning enables automated scoring of liver fibrosis stages. Sci Rep. 2018;8:16016. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 91] [Cited by in RCA: 70] [Article Influence: 8.8] [Reference Citation Analysis (5)] |
| 21. | Qu J, Yu Z, Li Q, Chen Y, Xiang D, Tan L, Lei C, Bai W, Li H, Shang Q, Chen L, Hu X, Lu W, Li Z, Chen D, Wang X, Zhang C, Xiao G, Qi X, Chen J, Zhou L, Chen G, Li Y, Zeng Z, Rong G, Dong Z, Chen Y, Lou M, Wang C, Lu Y, Zhang C, Yang Y. Blocking and reversing hepatic fibrosis in patients with chronic hepatitis B treated by traditional Chinese medicine (tablets of biejia ruangan or RGT): study protocol for a randomized controlled trial. Trials. 2014;15:438. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 32] [Article Influence: 2.7] [Reference Citation Analysis (2)] |
| 22. | Rockey DC, Caldwell SH, Goodman ZD, Nelson RC, Smith AD; American Association for the Study of Liver Diseases. Liver biopsy. Hepatology. 2009;49:1017-1044. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1851] [Cited by in RCA: 1626] [Article Influence: 95.6] [Reference Citation Analysis (6)] |
| 23. | Goodman ZD. Grading and staging systems for inflammation and fibrosis in chronic liver diseases. J Hepatol. 2007;47:598-607. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 743] [Cited by in RCA: 678] [Article Influence: 35.7] [Reference Citation Analysis (1)] |
| 24. | Vahadane A, Peng T, Sethi A, Albarqouni S, Wang L, Baust M, Steiger K, Schlitter AM, Esposito I, Navab N. Structure-Preserving Color Normalization and Sparse Stain Separation for Histological Images. IEEE Trans Med Imaging. 2016;35:1962-1971. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 714] [Cited by in RCA: 369] [Article Influence: 36.9] [Reference Citation Analysis (5)] |
| 25. | Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. 2017 IEEE International Conference on Computer Vision (ICCV); 2017 Oct 22-29; Venice, Italy. NJ, United States: IEEE, 2017: 618-626. [DOI] [Full Text] |
| 26. | Arjmand A, Tsipouras MG, Tzallas AT, Forlano R, Manousou P, Giannakeas N. Quantification of Liver Fibrosis—A Comparative Study. Appl Sci. 2020;10:447. [RCA] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 37] [Article Influence: 6.2] [Reference Citation Analysis (1)] |
| 27. | Watson AG, Mulay AS, Gill US. Chronic hepatitis B in 2025: diagnosis, treatment and future directions. Clin Med (Lond). 2025;25:100527. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 28. | Lai JC, Liang LY, Wong GL. Noninvasive tests for liver fibrosis in 2024: are there different scales for different diseases? Gastroenterol Rep (Oxf). 2024;12:goae024. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 31] [Article Influence: 15.5] [Reference Citation Analysis (3)] |
| 29. | Xu W, Hu Q, Chen C, Li W, Li Q, Chen L. Non-invasive Assessment of Liver Fibrosis Regression in Patients with Chronic Hepatitis B: A Retrospective Cohort Study. Infect Dis Ther. 2023;12:487-498. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 30. | Kawagishi N, Suda G, Kimura M, Maehara O, Yamada R, Tokuchi Y, Kubo A, Kitagataya T, Shigesawa T, Suzuki K, Ohara M, Nakai M, Sho T, Natsuizaka M, Morikawa K, Ogawa K, Kudo Y, Nishida M, Sakamoto N. Baseline elevated serum angiopoietin-2 predicts long-term non-regression of liver fibrosis after direct-acting antiviral therapy for hepatitis C. Sci Rep. 2021;11:9207. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 12] [Article Influence: 2.4] [Reference Citation Analysis (1)] |
| 31. | Zhang J, Chen S, Zhou J, Wang B, Wu X, Xu X, Zhao X, Kong Y, Ou X, Sun Y, You H. Serial Liver Stiffness Measurement and Serum Biomarkers Are Not Strong Predictors of the Regression of Fibrosis among Chronic Hepatitis B Patients Receiving Antiviral Therapy Based on Triple Liver Biopsies. Gut Liver. 2025;19:889-899. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 32. | Luo Y, Luo Q, Wu Y, Zhang S, Ren H, Wang X, Liu X, Yang Q, Xu W, Wu Q, Li Y. MRI-based machine-learning radiomics of the liver to predict liver-related events in hepatitis B virus-associated fibrosis. Eur Radiol Exp. 2025;9:81. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 33. | Li C, Wang Y, Bai R, Zhao Z, Li W, Zhang Q, Zhang C, Yang W, Liu Q, Su N, Lu Y, Yin X, Wang F, Gu C, Yang A, Luo B, Zhou M, Shen L, Pan C, Wang Z, Wu Q, Yin J, Hou Y, Shi Y. Development of fully automated models for staging liver fibrosis using non-contrast MRI and artificial intelligence: a retrospective multicenter study. EClinicalMedicine. 2024;77:102881. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 14] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 34. | LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436-444. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70666] [Cited by in RCA: 21014] [Article Influence: 1910.4] [Reference Citation Analysis (10)] |
| 35. | Xie Y, Xia Y, Zhang J, Song Y, Feng D, Fulham M, Cai W. Knowledge-based Collaborative Deep Learning for Benign-Malignant Lung Nodule Classification on Chest CT. IEEE Trans Med Imaging. 2019;38:991-1004. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 211] [Article Influence: 30.1] [Reference Citation Analysis (1)] |
| 36. | Qi S, Shan H, Fu Y, Chen Y, Zhang Q. A clinically oriented and interpretable AI framework for classifying dentin caries severity on CBCT images. J Prosthet Dent. 2025;S0022-3913(25)00831. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (1)] |
| 37. | Jiang X, Sun Q, Wang C, Li W, Chen W, Xu J, Yu L. CT-based radiomics and deep learning to predict EGFR mutation status in lung adenocarcinoma. Front Oncol. 2025;15:1597548. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 38. | Rakaee M, Tafavvoghi M, Ricciuti B, Alessi JV, Cortellini A, Citarella F, Nibid L, Perrone G, Adib E, Fulgenzi CAM, Hidalgo Filho CM, Di Federico A, Jabar F, Hashemi S, Houda I, Richardsen E, Rasmussen Busund LT, Donnem T, Bahce I, Pinato DJ, Helland Å, Sholl LM, Awad MM, Kwiatkowski DJ. Deep Learning Model for Predicting Immunotherapy Response in Advanced Non-Small Cell Lung Cancer. JAMA Oncol. 2025;11:109-118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 56] [Article Influence: 56.0] [Reference Citation Analysis (1)] |
| 39. | Lu MY, Williamson DFK, Chen TY, Chen RJ, Barbieri M, Mahmood F. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat Biomed Eng. 2021;5:555-570. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1755] [Cited by in RCA: 956] [Article Influence: 191.2] [Reference Citation Analysis (1)] |
| 40. | Foersch S, Eckstein M, Wagner DC, Gach F, Woerl AC, Geiger J, Glasner C, Schelbert S, Schulz S, Porubsky S, Kreft A, Hartmann A, Agaimy A, Roth W. Deep learning for diagnosis and survival prediction in soft tissue sarcoma. Ann Oncol. 2021;32:1178-1187. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 81] [Article Influence: 16.2] [Reference Citation Analysis (4)] |