Published online Mar 28, 2026. doi: 10.3748/wjg.v32.i12.112725
Revised: October 21, 2025
Accepted: January 16, 2026
Published online: March 28, 2026
Processing time: 196 Days and 23.3 Hours
Pancreatic cancer (PC) is a prevalent and highly malignant tumor. Reports indi
To investigate RHEB’s role in PC metastasis and the underlying mechanisms.
RHEB functions were validated using in vitro, cell counting kit-8 colony forma
RHEB promoted PC proliferation and metastasis in vitro. Higher RHEB expression was negatively correlated with longer survival times, with the gene also interacting directly with colony stimulating factor 1 receptor (CSF1R) to inhibit autophagy. The RHEB-CSF1R complex further advanced phosphorylation levels of phosphatidylinositol 3-kinase (PI3K), AKT serine/threonine kinase 1, and the mammalian target of rapamycin, as well as autophagy markers. The use of autophagy inhibitors confirmed that the RHEB-CSF1R complex could promote epithelial-mesenchymal transition markers expression levels. Finally, Transwell assays with CSF1R-overexpression/si
This study found that RHEB lowered PC patients’ survival. RHEB expression promoted PC proliferation, migration, and invasive ability, but it also inhibited autophagy by upregulating and interacting with CSF1R, leading to PI3K, AKT serine/threonine kinase 1, and mammalian target of rapamycin phosphorylation. This, in turn, promoted PC metastasis by promoting epithelial-mesenchymal transition marker expression. The above results indicated that RHEB can be a promising biomarker for predicting prognosis and developing new treatment strategies in PC.
Core Tip: Ras homolog enriched in brain (RHEB), small GTPases, play a crucial role in regulating neuronal activity and have gained attention for their implications in cancer development, particularly in pancreatic cancer (PC). Clarifying the precise mechanism through which the autophagy-related gene RHEB promotes PC metastasis holds the potential for improving prognosis as well as for developing new prevention and treatment strategies. This study aims to elucidate the role of the autophagy-related gene RHEB in PC metastasis.
- Citation: Deng QX, Yang K, He J, Li JF, Li XQ, Zou L, Li YM, Xu SM, Jiang Z, Wu LJ. Autophagy related RHEB-CSF1R complex promotes tumor metastasis via advancing phosphorylation levels of PI3K, AKT, mTOR in pancreatic cancer. World J Gastroenterol 2026; 32(12): 112725
- URL: https://www.wjgnet.com/1007-9327/full/v32/i12/112725.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i12.112725
Pancreatic cancer (PC), a prevalent and highly malignant tumor, is primarily responsible for cancer-related deaths worldwide[1], with ductal adenocarcinoma being the most common type[2]. According to GLOBOCAN 2022 data, of all malignancies, PC ranked 12th in terms of morbidity but 6th in mortality, involving about 530000 new cases and 531000 deaths[3]. Patients with this condition typically have a short median survival time as well as an alarmingly low five-year survival rate of only 4%[4], thereby imposing a significant burden on global healthcare systems. Such a high mortality rate can largely be attributed to challenges faced for early diagnosis, coupled with the tumor’s propensity for invasion and metastasis, as well as its poor responsiveness to most conventional chemotherapeutic interventions[4]. Therefore, there is an urgent need to enhance the current understanding of the underlying biological mechanisms governing PC’s onset and progression given the improving prognosis.
Autophagy, a core cytoprotective mechanism that is widely present in eukaryotes, is essential for preserving cellular and organismal homeostasis, thereby maintaining a stable internal environment[5]. Therefore, it is not surprising that altered levels of gene expression in autophagy-related processes are associated with various diseases, including malignancies[6]. In particular, as far as PC is concerned, autophagy has been reported to play a dual role, potentially promoting or inhibiting tumors at different stages[7-10]. In addition, autophagy can also influence the immunogenicity of chemotherapeutic-induced tumor cell death[11], thus indicating a close correlation between autophagy and the tumor immune microenvironment. However, since its role in PC metastasis remains poorly studied, further investigation is warranted.
Ras homolog enriched in brain (RHEB), a member of the Ras superfamily within the Ras/Rap/Ral subfamily, is one of the most prominent GTP-binding proteins as it exerts crucial regulatory roles in various fundamental cellular activities such as cell proliferation, differentiation, adhesion, and migration. In addition, it is tightly implicated in diverse physiological and pathological processes at the cellular and organismal levels[12]. Interestingly, RHEB is also known to be an essential gene that regulates cellular autophagy via two pathways: The direct modulation of AMP-activated protein kinase and indirect control of mammalian target of rapamycin (mTOR) and other autophagy-related genes[13]. Recent studies have suggested a close relationship between RHEB and cancer, especially regarding the former’s role in hepatocellular carcinoma[14], lung cancer[15], and prostate cancer[16] development. However, investigations into RHEB’s involvement in PC remain limited, particularly concerning its contribution to the metastatic processes leading to malignancy. Therefore, clarifying the precise mechanism through which the autophagy-related gene RHEB promotes PC metastasis holds the potential for improving prognosis as well as for developing new prevention and treatment strategies.
The present research aimed to elucidate the role of the autophagy-related gene RHEB in PC metastasis, along with the underlying regulatory mechanisms. The results revealed elevated RHEB expression in PC compared with normal tissues, with even higher expression observed in the case of metastatic samples. Furthermore, increased RHEB expression was correlated with reduced survival times for PC patients, with RHEB silencing attenuating the ability of cell proliferation and metastasis. Moreover, this study unveiled, for the first time, interactions between RHEB and colony stimulating factor 1 receptor (CSF1R). Mechanistic investigations suggested that RHEB promoted the phosphorylation of pho
PC patients, admitted to the Third Hospital of Mianyang between 2011 and 2021, were enrolled in this study to validate RHEB expression’s correlation with overall survival (OS), with the OS defined as the duration from diagnosis to all-cause deaths or the final follow-up in May 2021. The participant inclusion standards were: (1) Pathologically confirmed PC; (2) Tissue samples had > 70% of tumor cells as confirmed by pathological and histological examinations; and (3) Complete clinicopathological data were available. On the other hand, patients who disagreed with the project were excluded. Overall, 44 PC patients were retained for the study, and their tumor-node-metastasis staging was performed in accordance with the eighth edition American Joint Committee on Cancer staging system[17], while tumor grading was based on the degree of tumor differentiation. The study protocol got approval from the Ethics Committee of the Third Hospital of Mianyang, and followed the guidelines of the Declaration of Helsinki. The First Affiliated Hospital of Chongqing Medical University provided tissue samples, and the local Ethics Committee ensured that all procedures followed the guidelines of the Declaration of Helsinki for human research. All participants or their guardians signed the informed consent.
For reverse transcription-quantitative polymerase chain reaction (RT-qPCR), tissue samples were provided by the First Affiliated Hospital of Chongqing Medical University and consisted of 5 primary tumor tissue samples and 5 paired normal tissue samples (surgical margin tissues). Tissue microarrays were also acquired from Outdo Biotech Co. Ltd. (HPanA060CD02, Shanghai, China) and included 15 non-paired normal pancreatic tissues, 20 primary PC, as well as 24 distant metastatic paraffin-embedded tissues (18 liver metastasis and 6 lymph node metastasis).
Capan-1, CFPAC-1, PANC-1, and MIA-PaCa2 (human PC cell lines) were sourced from American Type Culture Collection, VA, United States, while Colo357 (a human PC cell line) was donated by Xiang TX from Chongqing University. Additionally, HPDE6-C7 (a human normal pancreatic ductal epithelial cell line, Cat number: 338285) was commercially available from the BeNa Culture Collection (Henan, China). Cell cultures were established in Dulbecco’s modified eagle’s medium (Gibco, 12491015, CA, United States) medium containing high glucose levels as well as 12.5% fetal bovine serum (VivaCell, C2630-0100, Telaviv Yafo, Israel) prior to incubation at 37 °C in a humidified environment and under 5% CO2. The Dulbecco’s modified eagle’s medium cell culture medium was prepared at a ratio of basal medium/serum/penicillin-streptomycin double antibody (Thermo Fisher, 10378016, CA, United States) = 8.5:1.5:0.1. The cell strain to be resuscitated was taken out from the liquid nitrogen tank and placed into a pre-warmed constant temperature water bath (Precision CIR 19, Thermo Fisher, CA, United States) at 37 °C. Then, it was quickly shaken to thaw the cells. Subsequently, the thawed cell suspension was transferred into a sterile centrifuge tube filled with 2 mL of cell culture medium in advance by using a pipette gun (Eppendorf, 3123000144, Hamburg, Germany) and mixed evenly by pipetting. It was then placed into a centrifuge and centrifuged at 800 rpm (75002545, Thermo Fisher, CA, United States) for 5 minutes. After that, the supernatant was discarded. 1 mL of cell culture medium was added again to the centrifuge tube to resuspend the cells. The cell suspension was transferred into a culture flask filled with 2 mL of cell culture medium in advance. After being fully shaken, it was placed in a cell incubator (371GPCN, Thermo Fisher, CA, United States) for cultivation and observed every other day. Furthermore, small interfering (si) RNA/short hairpin (sh) RNA targeting human RHEB and shRNA targeting human CSF1R were obtained from Tsingke (Tsingke Biotechnology, China), while RHEB-overexpressing and CSF1R-overexpressing plasmids were supplied by GeneCreate (GENECREATE Biotechnology, China). Capan-1 and CFPAC-1 cell lines were selected for silencing, with stable transfected shRHEB and sh-negative control (NC) subsequently generated over a two-week period through a selection process that involved exposure to 1 μg/mL puromycin (Biosharp, BL528A, Shanghai, China). The same cell lines were then used for transient transfection with siRHEB and siNC for 48 hours, after which pcDNA3.1 and RHEB-harboring plasmid (4 μg) were introduced into Colo357. This was followed by a two-week screening process using G418 (Amresco, E859, OH, United States) to yield stable transfected RHEB and control cell lines. In subsequent rescue experiments, pcDNA3.1 and plasmid carrying CSF1R (4 μg) were introduced into Capan-1 and CFPAC-1 cell lines stably transfected with shRHEB and shNC for silencing. The Colo357 cell line stably transfected with pcDNA3.1 and carrying the RHEB plasmid was selected for CSF1R silencing; and assessment of stability was performed as previously described. All experiments were performed according to the user manual, and Lipofectamine 2000 (Invitrogen, 11668030, CA, United States) was employed for all transfection processes. Cells were eventually harvested after transfection for 48 hours for RT-qPCR and western blot (WB) assays in order to assess the efficiency of gene silencing and overexpression in cells (Table 1).
| Sequences | |
| Negative control | UUCUCCGAACGUGUCACGU(dT)(dT) |
| ACGUGACACGUUCGGAGAA(dT)(dT) | |
| RHEB/siRNA | RHEB-siRNA1: GTGTATTCTGTTACATCAA |
| RHEB-siRNA2: CACAGTAAATGGACAAGAA | |
| RHEB-siRNA3: GAAAGACCTGCATATGGAA | |
| RHEB/shRNA | RHEB-shRNA1: CCGGCCTCAGACATACTCCATAGATCTCGAGATCTATGGAGTATGTCTGAGGTTTTTT |
| RHEB-shRNA2: CCGGTTATGTTGGTTGGGAATAAGACTCGAGTCTTATTCCCAACCAACATAATTTTTT | |
| RHEB-shRNA3: CCGGCCTACGATCCAACCATAGAAACTCGAGTTTCTATGGTTGGATCGTAGGTTTTTT | |
| RHEB-shRNA4: CCGGTATCATCTTCAACTTGTAGACCTCGAGGTCTACAAGTTGAAGATGATATTTTTT | |
| CSF1R/shRNA | CSF1R-shRNA1: CTGCTGACTGTTGAGACCTTA |
| CSF1R-shRNA2: CGACTATAAGAACATCCACCT | |
| CSF1R-shRNA3: CCAACAACGCTACCTTCCAAA | |
| CSF1R-shRNA4: ACAGGAGAGAGCGGGACTATA |
The experiments for cell and tissue RNA isolation were performed following the manufacturer’s instructions for ES Science® (YiShan Biotech, ES-RN001, Shanghai, China). Briefly, aliquots containing 1 μg total RNA were subjected to RT-PCR to generate 20 μL of complementary DNA, followed by amplification under the following conditions as per the instructions for Go-Taq (Promega, m7421, WI, USA): 2 minutes of initial denaturation (95 °C), followed by 32 amplification cycles and a final 3-minute extension (72 °C). GAPDH was amplified as a control, and 23 PCR cycles were conducted using the relevant primer sequences (species origin of primers: Human) (Table 2). SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biology, AG11718, Hunan, China) was applied for all PCR amplifications, and all experiments were performed in triplicate on the 7500 RealTime PCR System (Thermo Fisher, CA, United States). Relative quantification of RNA expression levels was performed using the 2-ΔΔCt method. Three independent replicate experiments were done to verify the experimental results.
| Primer | Sequence (5′-3′) | Molecular weight (kDa) | Annealing temperature (°C) |
| CSF1R-F | ATCTTCTCACTTGGGCTGAATCC | 6965.46 | 60 |
| CSF1R-R | AGATTGGTATAGTCCCGCTCTCT | 7005.46 | |
| FGF22-F | GTGGTCATCAAAGCAGTGTCCT | 6750.27 | 60 |
| FGF22-R | GTAGGTGTTGTGGCCGTTCT | 6185.83 | |
| AMTN-F | AGCAGGAGGAGCAGGTGTAAAT | 6897.3 | 60 |
| AMTN-R | ACCAAATTCGAGGCAGCTTAGTT | 7047.51 | |
| TNFSF12-TNFSF13-F | ATAGCCAGGTCCTGTTTCAAGAC | 7023.48 | 60 |
| TNFSF12-TNFSF13-R | AAATGGAAGACACCTGCGCTATA | 7065.63 | |
| UQCRHL-F | GGAGGAAGAGGAAGAGGAGGAAT | 7324.5 | 60 |
| UQCRHL-R | CGTGCAACCTCTTCTGTATGTG | 6996.45 | |
| DACT1-F | AGTGACCTGAGACTGGATGTAGA | 7152.49 | 60 |
| DACT1-R | GGTGCTGGAATGACAACATGGATA | 7152.49 | |
| ARRDC4-F | ACTGCCATTAGTGATCGGTACAA | 7047.51 | 60 |
| ARRDC-R | CACACTTCTCCCTCACAGTTAGG | 6919.45 | |
| GAPDH-F | CACTCCTCCACCTTTGACGC | 5948.9 | 60 |
| GAPDH-R | CTGTTGCTGTAGCCAAATTCGT | 6716.4 |
WB analysis was carried out as previously described[18]. Rapamycin-induced proximity assay (Beyotime Technology, P0013K, Shanghai, China) buffer with phenylmethylsulfonyl fluoride (1 mmol/L, ST505, Beyotime Technology, Shanghai, China) and NaF (1 mmol/L; AG5355, Beyotime Technology, Shanghai, China) was employed to obtain whole-cell lysates. After centrifugation (5 minutes, 13000 × g), the supernatants were collected. A bicinchoninic acid Protein Assay Kit (P0010S, Beyotime Technology, Shanghai, China) was used for total protein concentration quantification. The extracted protein was added to 5 × loading buffer (Mengbio, MBW141, Chongqing, China) and denatured by boiling for 10 minutes at 100 °C. Aliquots of 40 μg of protein lysates were then electrophoresed on 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (Epizyme, LK303, Shanghai, China) prior to the transfer of the resulting separated proteins to polyvinylidene fluoride membranes (Invitrogen, 88520, CA, United States). The latter were blocked with blocking buffer (P0222, Beyotime Technology, Shanghai, China), after which they underwent incubation (overnight, 4 °C) with primary antibodies against RHEB (1:3000, Abcam, ab92313, Cambridge, United Kingdom), CSF1R (1:550, Santa Cruz, sc-46662, CA, United States), GAPDH (1:3000, Huabio, 12D6, Hangzhou, China), PI3K (1:500, Wanleibio, WL03380, Shenyang, China), corresponding to the P110 catalytic subunit, it is the P110β isoform, p-PI3K (1:1000, PA5-85820, Thermo Fisher, CA, United States), corresponding to the Tyr524 phosphorylation site of PI3K. AKT (1:750, WL0003b, Wanleibio, Shenyang, China), p-AKT (1:750, Proteintech, 80455-1-RR, Wuhan, China), corresponding to the Ser473 phosphorylation site of AKT. mTOR (1:500, WL02477, Wanleibio, Shenyang, China), p-mTOR (1:500, s2448, Abcam, Cambridge, United Kingdom), Beclin 1 (1:500, WL02237, Wanleibio, Shenyang, China), autophagy related 5 (ATG5; 1:750, WL02411, Wanleibio, Shenyang, China), microtubule-associated protein 1 light chain 3 (LC3α/β; 1:750, WL01506, Wanleibio, Shenyang, China), E-cadherin (1:400, 20874-1-AP, Proteintech, Wuhan, China), N-cadherin (1:750, 22018-1-AP, Proteintech, Wuhan, China), vimentin (1:500, 10366-1-AP, Proteintech, Wuhan, China), interleukin 4 (1:1500, WL05916, Wanleibio, Shenyang, China) and interleukin 10 (1:1500, WL03088, Wanleibio, Shenyang, China). Next, horseradish peroxidase-conjugated secondary antibodies (mouse: 1:5000, Servicebio, G1214, Wuhan, China; rabbit: 1:5000, Servicebio, G1213, Wuhan, China) were added for 2-hour incubation at 4 °C. The horseradish peroxidase Substrate kit (Merckmillipore Corporation, WBKLS0100, Darmstadt, Germany) was eventually used for visualizing the blots. Rescue experiments were used with the autophagy inhibitor chloroquine (MedChemExpress, HY-17589, NJ, United States), the PI3K activator 740Y-P (HY-P0175, MedChemExpress, NJ, United States), and the PI3K inhibitor LY294002 (HY-10108, MedChemExpress, NJ, United States). Three independent experiments were performed to ensure the accuracy of the results.
Capan-1 and CFPAC-1 cells in which RHEB was stably silenced and Colo357 cells in which RHEB was stably overexpressed were applied. Briefly, after cell transfection with shRHEB plasmid, total protein was extracted for overnight incubation on the Ferris wheel (60 yards) at 4 °C. Next, antibodies containing anti-Flag M2 (Sigma-Aldrich, F3165, MO, United States), RHEB (1:750, ab92313, Abcam, Cambridge, United Kingdom), and CSF1R (1:2000, Santa Cruz, sc-46662, CA, United States) were added for incubation. The Millipore SigmaTm Pure ProteomeTm Protein A/G Mix Magnetic Bead System (LSKAGAG10, Merck Millipore Corporation, Darmstadt, Germany) was eventually used for purification of the resulting co-immunoprecipitation (Co-IP) complex. Three independent experiments were performed in each case.
Immunohistochemistry (IHC) was performed on paraffin-embedded tissues and tissue microarray chips. This involved overnight incubation with the primary antibodies against RHEB (1:500, Abcam, ab92313, Cambridge, United Kingdom), Ki67 (1:2000, 28074-1-AP, Proteintech, Wuhan, China), LC3 (1:300, WL01506, Wanleibio, Shenyang, China), p-AKT (1:400, 80455-1-RR, Proteintech, Wuhan, China), and p-mTOR (1:500, ab109268, Abcam, Cambridge, United Kingdom) at 4 °C. Take out the humid box on the next day. After the tissue microarray returns to room temperature, rinse it in phosphate-buffered saline (PBS) Tween-20 (PBST, 500 mL PBS, G4202, Servicebio, Wuhan, China; 2 mL Tween-20, GC204002, Servicebio, Wuhan, China) solution three times, with each rinse lasting for 5 minutes. Use filter paper again to absorb the excessive PBST solution around the tissue, then drop an appropriate amount of anti-rabbit secondary antibody (mouse: 1:200, G1214, Servicebio, Wuhan, China; rabbit: 1:200, G1213, Servicebio, Wuhan, China) onto the tissue to cover it and incubate it in a 37 °C incubator for 30 minutes. After taking out the tissue microarray, rinse it with PBST solution three times again, with each rinse lasting for 5 minutes. Then, use filter paper to absorb the excessive PBST solution around the tissue, drop an appropriate amount of streptavidin-peroxidase onto the tissue to cover it, and incubate it in a 37 °C incubator for 30 minutes. Subsequently, rinse it with PBST solution three times, with each rinse lasting for 5 minutes. After re-probing with secondary antibodies, standard DAB (Beijing ZhongShanJinQiao Biotechnology Co., Ltd., PV-6000D, Beijing, China) was employed for visualizing the labeled antigens, and this was followed by counter-staining with hematoxylin (G1004, Servicebio, Wuhan, China). Take out the tissue microarray and then put it into 70% ethanol, 85% ethanol, 90% ethanol, 95% ethanol, and 100% ethanol in sequence, soaking it in each for 2 minutes. Then, soak it in xylene A and xylene B, respectively 10 minutes each. After that, place it in a fume hood to dry. The staining intensity (0-3 points: Low-medium-high), as well as positive cell percentage (0-4 points: < 1%, 1%-10%, 11%-50%, 51%-80%, > 80%), were then scored before calculating the immunoreactive score as follows: Staining intensity score × positive cell percentage score. These immunoreactive scores were computed independently by two scorers, and the average was taken as the result.
Immunofluorescence (IF) staining was carried out using sections that had been incubated with primary antibodies against RHEB (1:1000, ab92313, Abcam, Cambridge, United Kingdom) and LC3 (1:400, WL01506, Wanleibio, Shenyang, China) prior to incubation with the Cy3-labeled goat anti-rabbit IgG (H + L) secondary antibodies (1:500, P0183, Beyotime, Shanghai, China). This was followed by counterstaining using 2 μg 4’,6-diamidino-2-phenylindole (C1002, Beyotime, Shanghai, China), after which images were captured on a fluorescence microscope (Leica Microsystems GmbH, DM6000, Wetzlar, Germany).
After fixing using 2.5% glutaraldehyde (Servicebio, G1102, Wuhan, China), stable transfected cells embedded in Epon 812 resin (SPI, SPI-02660, CA, United States) were dehydrated in acetone of different concentrations (Sinopharm Chemical Reagent Co., Ltd, CSDS67-64-1, Shanghai, China). Sections (1-μm thick) were then dyed with sodium acetate and lead citrate (Ted Pella Inc., 19314, CA, United States) prior to visualization by transmission electron microscope (TEM, Hitachi, HT7800, Tokyo, Japan).
Cells that had been cultured in 96-well plates (2000 cells/well) were kept at 37 °C and under 5% CO2 for 0, 24 hours, 48 hours, and 72 hours. Cells were then incubated (2 hours) with cell counting kit-8 (CCK-8) reagent (BS350A, Biosharp, Shanghai, China), after which absorbance readings were taken with a microplate reader (Tecan, Infinite 2000-PRO, Switzerland) at 450 nm. Each experiment was performed with three replicates.
Cells that had been added to six-well plates (1000 cells/well, 2000 cells/well, and 3000 cells/well) were cultured for 10-14 days. After fixation in 4% paraformaldehyde (G1109, Servicebio, Wuhan, China), a scanner (CanoScan, 8800F MOEL-85, Japan) was utilized for scanning and counting the number of surviving colonies prior to visualization by Gentian violet (G1014, Servicebio, Wuhan, China). For each experiment, there were three independent replicates.
Cell migration and invasion were assessed using Transwell chambers (Corning Inc., NY, United States; 6.5-mm diameter inserts, 8-mm pore size). A Matrigel glue (Becton, Dickinson and Company, 3422, NJ, United States) was added to the Transwell membrane for the cell invasion assay. After the incubation is completed, take out the Transwell chambers and gently wipe off the cells that have not migrated or invaded the surface of the upper chamber membrane with cotton swabs. Then put the chamber membrane into paraformaldehyde to fix it for a period of time. Subsequently, stain the cells that have migrated or invaded onto the surface of the lower chamber membrane with 1 mL crystal violet staining solution. The staining time is usually more than 30 minutes. After staining, rinse the chamber membrane with clean water to remove the excess staining solution. Forty-eight hours post inoculation, a phase contrast microscope (DVM6, Leica, Wetzlar, Germany) was employed to photograph and count cells on the lower chamber surface. Each experiment was independently performed three times.
Female BALB/c nude mice (4 weeks, National Laboratory Animal Center, Shanghai, China) were housed in standard pathogen-free environments. The animals were then subjected to subcutaneous administration of stable RHEB-silenced Capan-1 cells and shNC cells (2 × 106 cells resuspended in 150 μL PBS; MB1980, Mengbio, Chongqing, China). Seven days later, the tumor size was measured, with subsequent measurements taken every three days. After nineteen days, mice were euthanized, and tumor tissues were excised for experimentation.
To establish a metastasis model in nude mice, a suspension of shRHEB and shNC stably-transfected Capan-1 cells (resuspended in 100 μL PBS) was prepared at a concentration of 5 × 107/mL. The mice were intraperitoneally administered with 1% sodium pentobarbital (50 mg/kg, TCI, 144-02-05, Hashimoto, Japan), and then fixed in the right decubitus position. The spleen was subsequently exposed with a longitudinal incision on the left side of the abdomen post-disinfection, and its lower pole was slowly injected with tumor cell suspension. After the injection, the abdomen was closed entirely. The animals were routinely fed for 4 weeks post-administration, and mice were euthanized through cervical dislocation, and metastatic tumors were observed.
The UCSC Xena website provided the phenotypic and clinicopathological information as well as the survival data of pancreatic adenocarcinoma (PAAD) patients of The Cancer Genome Atlas (TCGA), along with data regarding RNA-seq batch effects for a group of normalized gene expression. From the Gene Expression Omnibus dataset GSE19279, four primary pancreatic ductal adenocarcinoma samples and five liver metastasis samples were then selected into the primary and metastasis groups, respectively. The HADb database (https://www.autophagy.lu/v1/) provided autophagy-associated gene data, while the UALCAN database (https://ualcan.path.uab.edu/analysis.html) was employed for detecting the pan-cancer expression of RHEB. Eventually, interactions between known and predicted proteins were identified and visualized by the STRING database (https://cn.string-db.org/) and the Cytoscape software (3.9.0 Beijing Bio-Tech Pack Technology Co. Ltd., Beijing, China). All datasets are publicly available.
SPSS Statistics 26.0 (IBM, NY, United States) or GraphPad Prism 9 (Dotmatics, United Kingdom) was employed for the statistical data processing. Data with or without normal distribution were compared using Student’s t-tests and Mann-Whitney U-tests, separately, while Fisher’s exact or χ2 tests were employed for categorical variable analysis. In addition, for continuous variables, multiple group comparisons were performed using ANOVA with the Tukey multiple comparison test correction. Pearson or Spearman correlation coefficient analysis also undertaken to assess the relationship between two variables. OS and disease-free survival were then assessed using Kaplan-Meier survival curves along with the log-rank test, while uni- and multi-variable Cox regression methods were employed for survival analysis. Finally, the R software (version 4.2.1, https://www.r-project.org) and the “clusterProfiler” package were used for Gene Set Enrichment Analysis (GSEA) as well as Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses. The significance threshold was determined by multiple hypothesis testing, with a set value of 0.05. Results were considered statistically significant at P value < 0.05. Multiple comparisons were adjusted to maintain statistical rigor in the experimental design. Three times repeated for each assay in the statistical analysis and figure legends. However, there may be potential biases or limitations due to sample size limitations, variability in cell line characteristics, or assay sensitivity.
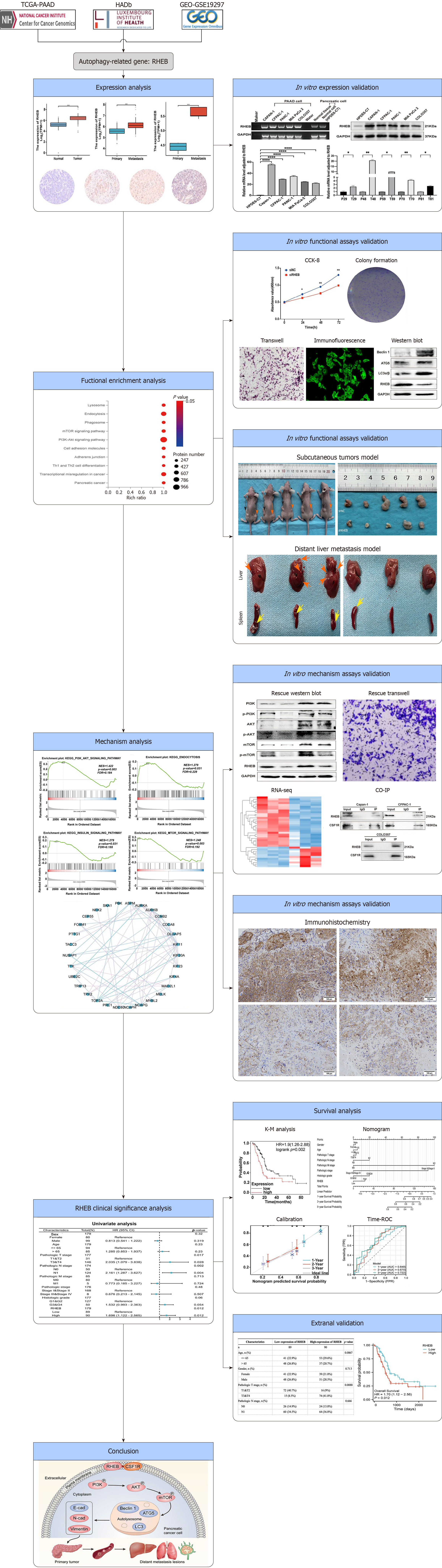
The study methodology is presented in Figure 1. Overall, the results of the TCGA tumor tissues and their corresponding normal ones showed that RHEB was highly expressed in most tumors (Supplementary Figure 1A). In particular, a significant upregulation of RHEB was observed in PC tumor tissues within the TCGA-PAAD database (Supplementary Figure 1B), while a more pronounced upregulation was validated in metastatic tissues compared with primary tissues in the TCGA-PAAD (Supplementary Figure 1C) and GSE17297 datasets (Supplementary Figure 1D).
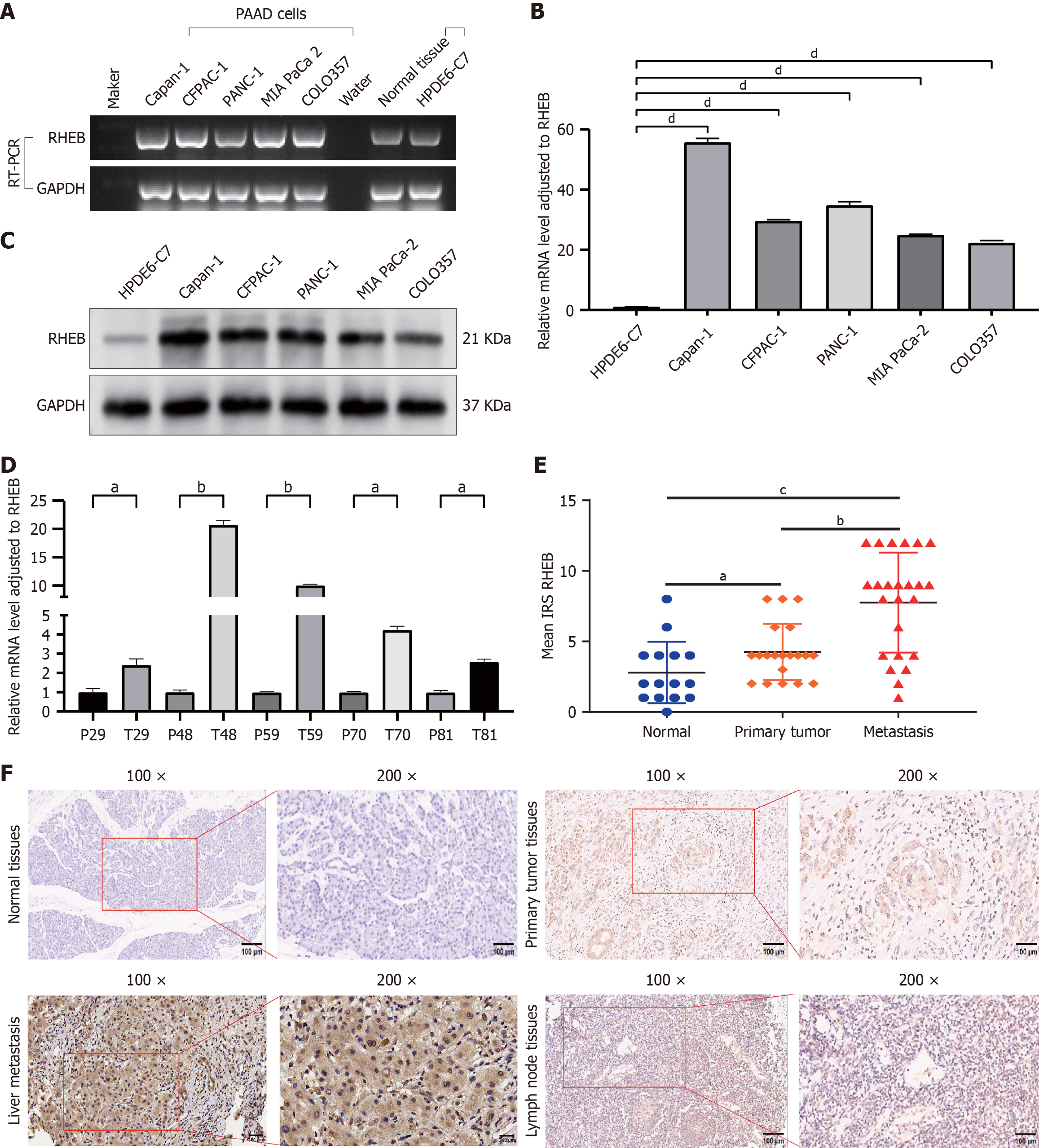
Additionally, RHEB expression in PC was experimentally assessed. The findings indicated a remarkable increase in RHEB mRNA level in Capan-1, CFPAC-1, PANC-1, MIA PaCa2, and Colo357 PC cell lines (Figure 2A and B). On the other hand, although the results were generally reflected at the protein level, only a slight increase in RHEB expression was observed in Colo357 (Figure 2C). Moreover, 5 PC tissues also showed elevated mRNA expression levels relative to paired non-cancerous pancreatic ones (Figure 2D). Subsequent experiments were aimed at detecting RHEB protein expression in 15 non-paired normal pancreas, 20 primary PC, and 24 distant metastatic tissues (18 liver metastasis and 6 lymph node metastasis) using IHC. In this case, it was found that RHEB exhibited higher expression in the primary PC tissues than in normal pancreatic ones. Similarly, higher RHEB expression was observed in metastatic tissues as opposed to primary PC ones (Figure 2E and F). Bioinformatics analysis results were consistent with the above findings, especially in confirming the increased RHEB expression in PC.
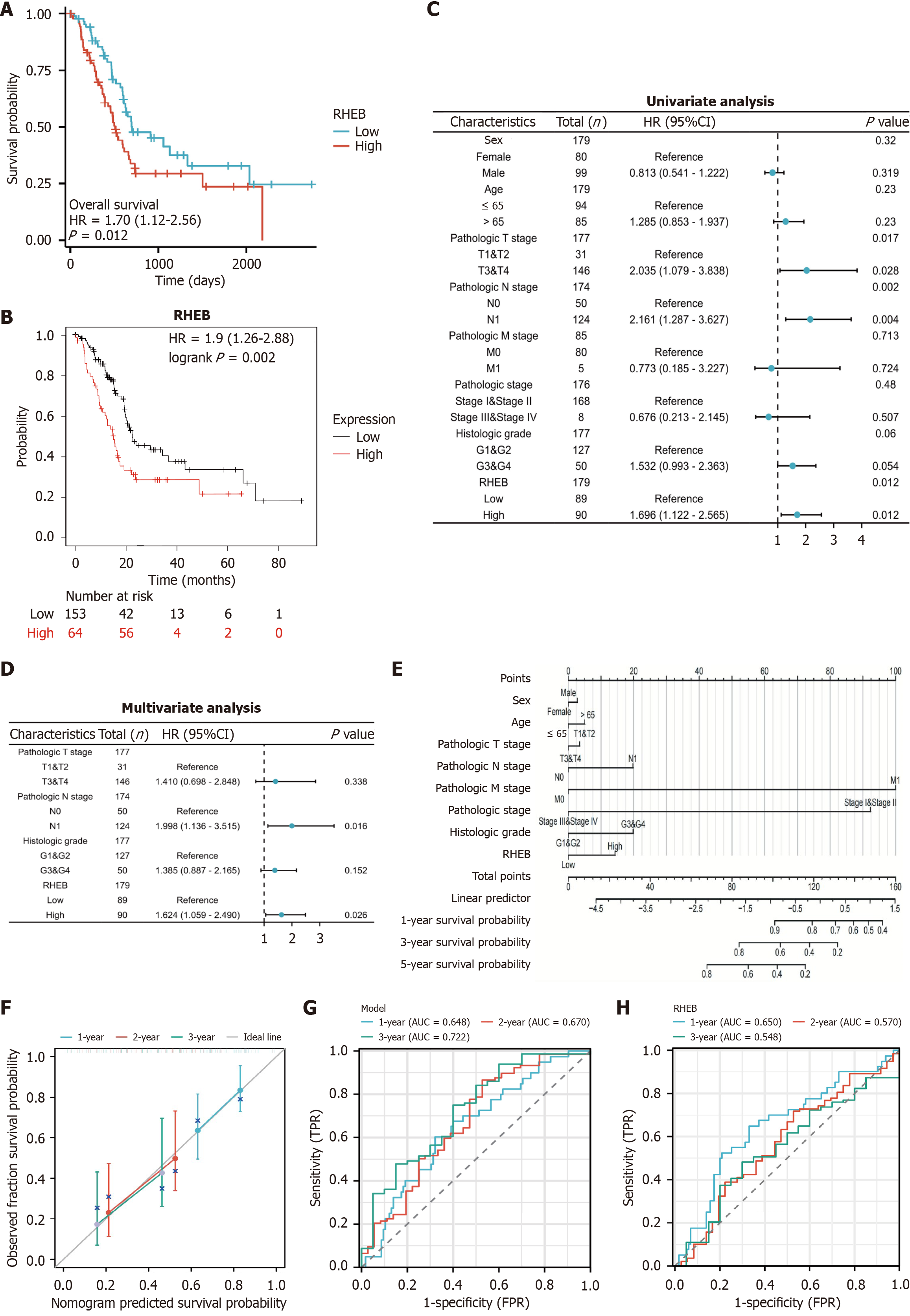
To validate the clinical significance of RHEB in PC, the correlation of its expression with disease prognosis was determined for 44 PC patients admitted to the authors’ hospital. The results reflected those obtained for the TCGA database (Figure 3A). Furthermore, based on the TCGA-PAAD database (n = 177), higher RHEB expression was correlated to poorer survival rates (Figure 3B), as determined in the previous results.
Differential clinicopathological characteristics between the subgroups (high- and low-RHEB expression) were then studied. Overall, the T and N stages, histologic grades, and number of deaths were markedly higher in PC patients within the high-RHEB subgroup (Figure 3C, Supplementary Table 1). Furthermore, through multivariate Cox analysis of those three characteristics, RHEB expression and the N stage were identified as independent prognostic factors of the condition (Figure 3D). An RHEB-related nomogram prognostic model was then constructed and visualized (Figure 3E) before using calibration curves and time-receiver operating characteristic to evaluate the predictive potential and sensitivity of the model. In this case, the model’s predicted outcomes for years 1, 2, and 3 were close to the standard curves (Figure 3F). Specifically, the area under the curves for 1-, 3- and 5-year OS for the model (0.648, 0.670 and 0.722; Figure 3G) was higher than that for RHEB single-gene 1-, 3- and 5-year OS (0.650, 0.570, and 0.548; Figure 3H), highlighting the prognostic potential and sensitivity of the above models. Therefore, the results suggest that RHEB could potentially represent a predictive biomarker of PC, although this remains to be further studied.
To determine whether RHEB could be related to PC proliferation and metastasis, the correlation between its expression in primary and metastatic PC and poor prognosis was determined. To verify the hypothesis, paired samples were subjected to KEGG and GSEA analysis (Supplementary Figure 2), with the results revealing RHEB upregulation to be significantly related to 10 pathways (P value < 0.05) within the TCGA-PAAD and GSE19279 datasets (Supplementary Figure 2A and
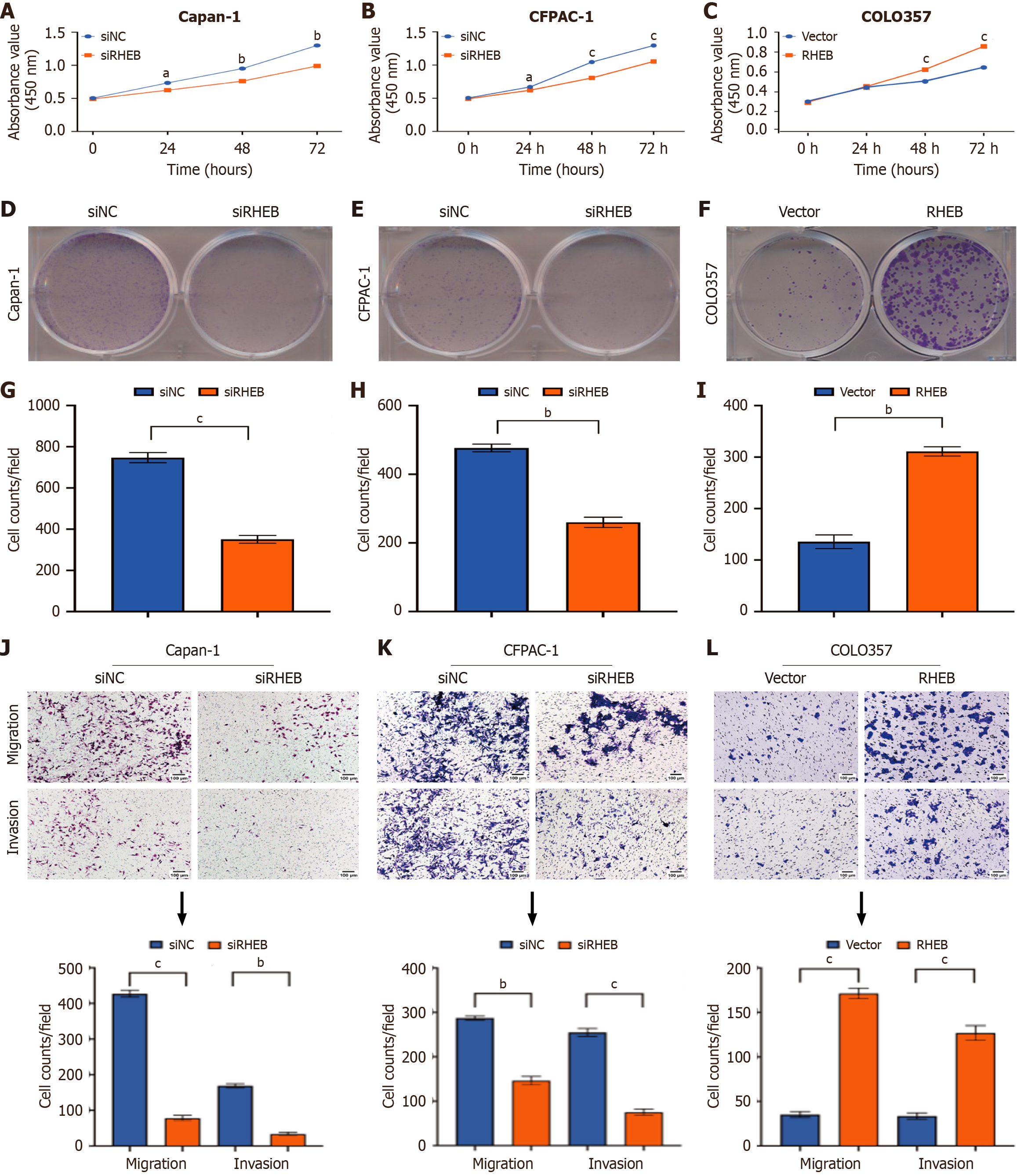
Taking into account RHEB expression patterns in PC cell lines, RHEB silencing models were established in Capan-1 and CFPAC-1 cell lines, while an overexpression model was generated using the Colo357 cell line. The efficiency of RHEB silencing via siRNA and shRNA in the two cell lines was then validated by RT-qPCR and WB before selecting the RHEB/siRNA2 and RHEB/shRNA4 constructs exhibiting optimal silencing effects for subsequent experiments (Supplementary Figure 3). Similarly, the successful construction of the RHEB overexpression cell line in Colo357 was confirmed (Supplementary Figure 3B and F). As revealed by CCK-8 and colony formation assay results (Figure 4), RHEB silencing notably inhibited PC cell proliferation (Figure 4A, B, D, E, G, and H), while its overexpression promoted cell proliferation (Figure 4C, F, and I). Furthermore, the potential effects of RHEB on metastasis were investigated. In this case, fewer RHEB-silenced cells passed through the Transwell chamber in the Capan-1 and CFPAC-1 cell lines. In contrast, significantly more RHEB-overexpressing Colo357 cells passed through the chamber. Finally, RHEB-deficient PC cells exhibited reduced invasion through the Matrigel barrier compared with the control, with the opposite result noted for RHEB-overexpressing PC cell lines (Figure 4J-L).
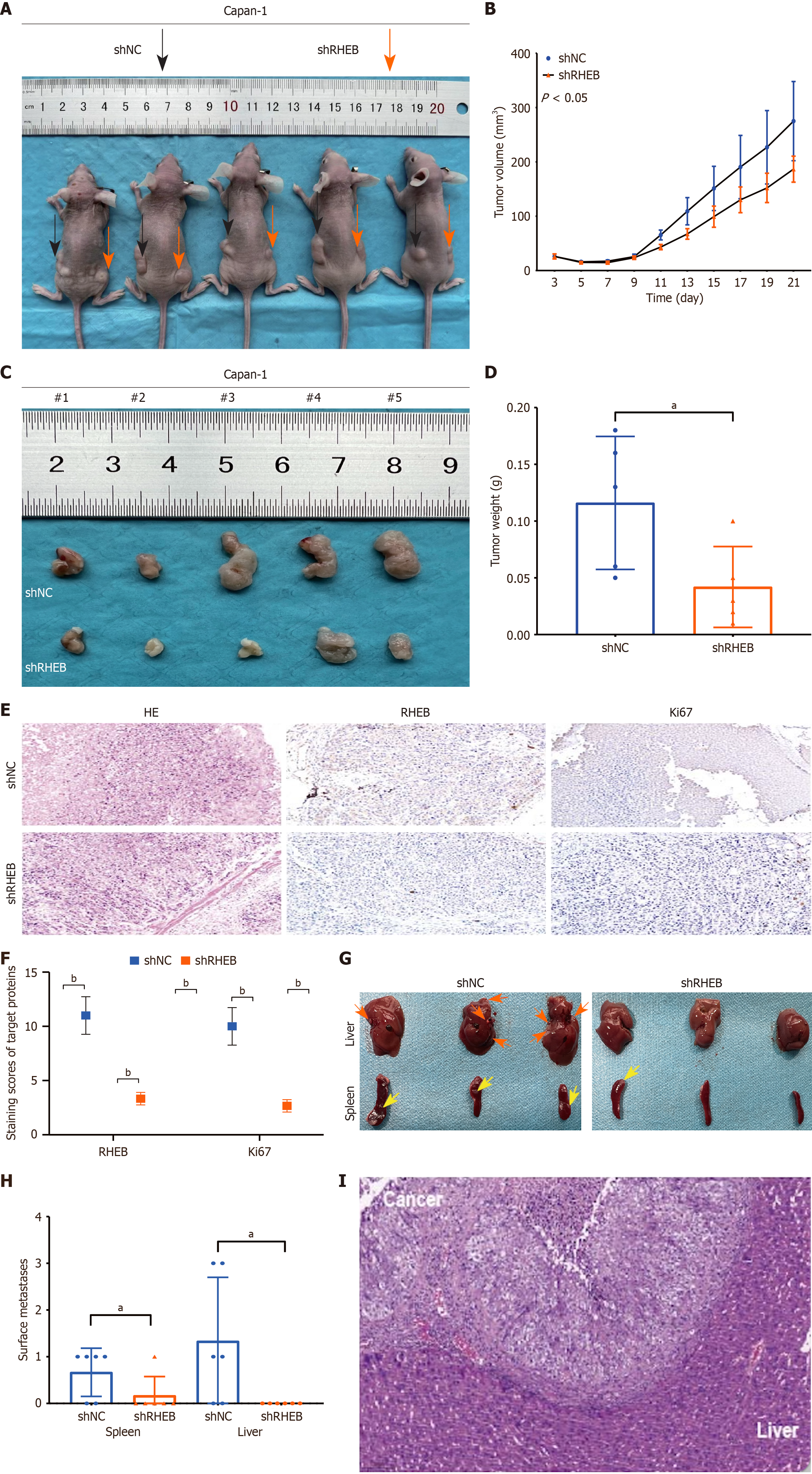
To examine RHEB’s anticancer effects in vivo, nude mice were injected with Capan-1 cells to form a subcutaneous xenograft tumor as well as a liver metastasis model. Since the efficiency of shRNA silencing of RHEB was already verified in the Capan-1 cell line in vitro (Supplementary Figure 3D), a similar RHEB/shRNA4 construct was selected for the in vivo experiments. It was found that the tumor volume in the shRHEB group was significantly smaller than that in the control group as time went by, and the weight of the tumor was also significantly lighter than that of the control group. Figure 5A-D with primary tumors diagnosed by hematoxylin-eosin staining. In addition, IHC verified the downregulation of RHEB and Ki67 levels compared to the shNC group (Figure 5E and F). In the distant liver metastasis model, the shNC group showed a notably much higher average number of primary tumors in the spleen and liver metastases relative to the shRHEB group (Figure 5G and H). These distant liver metastases were successfully identified by hematoxylin-eosin staining (Figure 5I). The above experiments implied that PC cell proliferation and metastasis were weakened after silencing of RHEB in vivo.
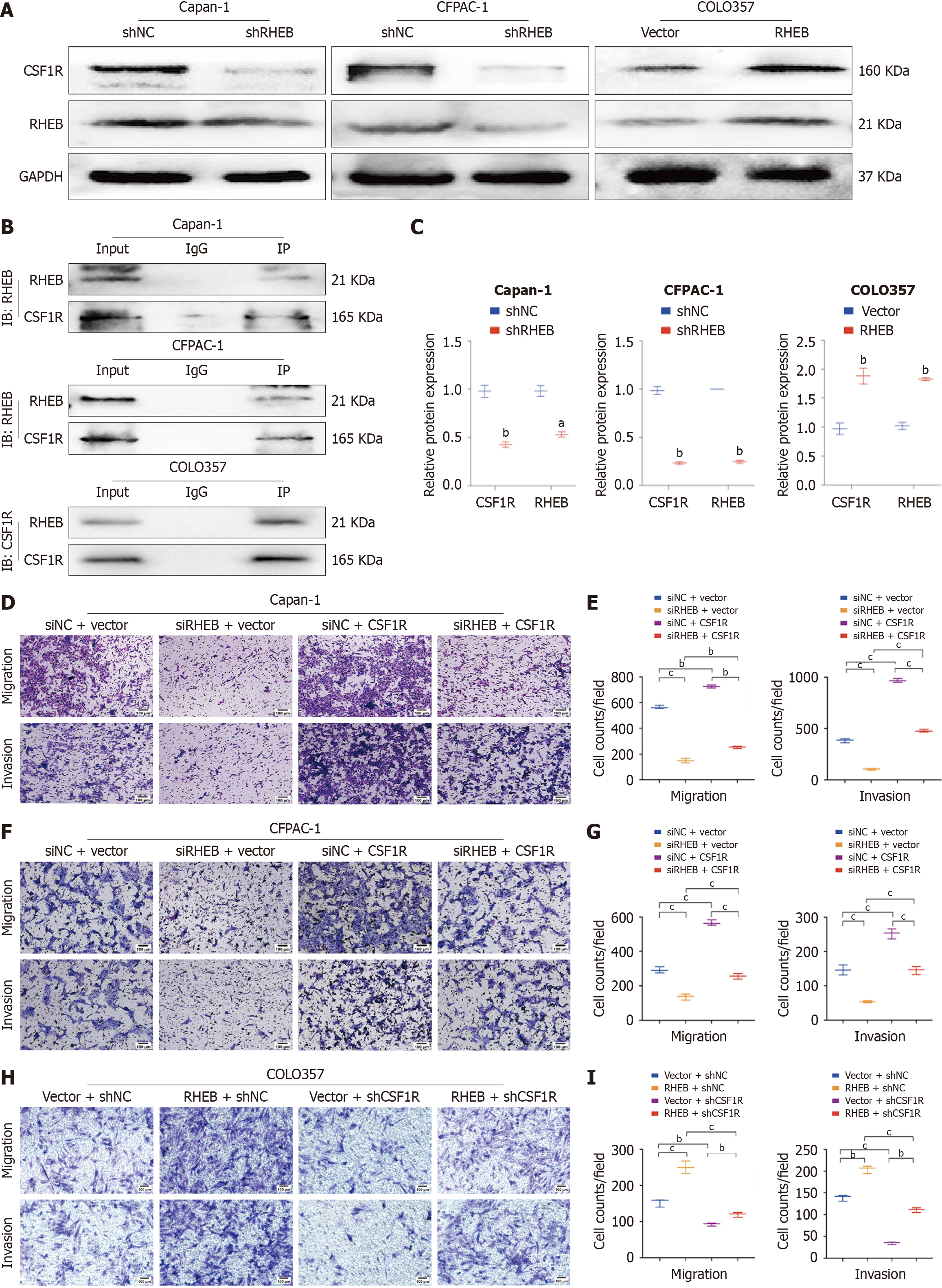
Early metastasis contributes to unfavorable outcomes in PC patients. Therefore, to further investigate the underlying mechanisms regulating metastasis phenotypes, the transcriptome (RNA-Seq) of Capan-1 cell lines, stably transfected with shRHEB and shNC, was examined by high-throughput sequencing, and differential genes between the two groups were identified (Supplementary Figure 4A). The criteria for screening differentially expressed genes were set to a |log2fold change| value of > 1 and a P value of < 0.05, with this process yielding 39 eligible genes (Supplementary Figure 4B). Their correlation with RHEB was analyzed separately, and the results showed that CSF1R, FGF22, AMTN, TNFSF12-TNFSF13, DACT1, ARRDC4, and UQCRHL were strongly correlated with RHEB (Supplementary Figure 4C). Subsequent validation of this finding by RT-qPCR indicated that the mRNA expression of only CSF1R in Capan-1, CFPAC-1, and Colo357 cell lines was consistent with the sequencing trends (Supplementary Figure 4D). Previous research has documented elevated CSF1R levels in different types of tumors[19-21], with WB confirming the downregulation of CSF1R in Capan-1 and CFPAC-1 cells that had been transfected with shRHEB as well as its upregulation in Colo357 cells that had been transfected with RHEB-overexpressing constructs (Figure 6A-C). Co-IP, validated through the binding of CSF1R and RHEB in Capan-1, CFPAC-1, and Colo357 (Figure 6B), indicated that the two proteins could form a complex. It was therefore hypothesized that RHEB could interact and form a complex with CSF1R.
CSF1R was subsequently analyzed by GO and KEGG enrichment analyses, and it was found that the CSF1R function was enriched in cell adhesion and migration (Supplementary Figure 5A), with these results corroborated by those obtained during GSEA (Supplementary Figure 5B-D). By integrating these findings with previous studies on RHEB gene function, it was hypothesized that RHEB could complex with CSF1R to influence the metastatic phenotype of PC.
To investigate whether RHEB can regulate PC metastasis through CSF1R, replenishment Transwell experiments were performed based on CSF1R-overexpression in Capan-1/siNC and Capan-1/siRHEB or CFPAC-1/siNC and CFPAC-1/siRHEB cell lines or CSF1R-silencing in Colo357/vector and Colo357/RHEB. Firstly, the successful construction of CSF1R overexpression in Capan-1 and CFPAC-1 cell lines was confirmed by RT-qPCR and WB, while the efficiency of shRNA silencing was verified in the Colo357 cell line. The CSF1R/shRNA3 construct was then selected for subsequent experiments based on RT-qPCR and WB results (Supplementary Figure 5E and F). Quite expectedly, the CSF1R-overexpression group showed enhanced migration and invasion relative to the vector one, but these abilities were largely diminished in the CSF1R-silencing group. Moreover, the migration and invasion of Capan-1/siRHEB, CFPAC-1/siRHEB, and Colo357/vector cells were significantly enhanced compared with those of Capan-1/siNC, CFPAC-1/siNC, and Colo357/RHEB cells (Figure 6D-I). These results further confirm the tumor metastasis-promoting role of RHEB, possibly by upregulating CSF1R in PC.
The downstream mechanisms through which RHEB mediates PC metastasis were then studied. GSEA and KEGG analyses of differential genes in the shNC and shRHEB groups revealed significant enrichment in the mTOR, PI3K-AKT, and insulin pathways (Supplementary Figure 6A-E). Protein-protein interaction networks further demonstrated a close association between RHEB and proteins such as PI3K, AKT, and autophagy-related protein 13 (Supplementary Figure 6F). Taken together, RHEB might promote PC metastasis via the PI3K/AKT/mTOR pathway.
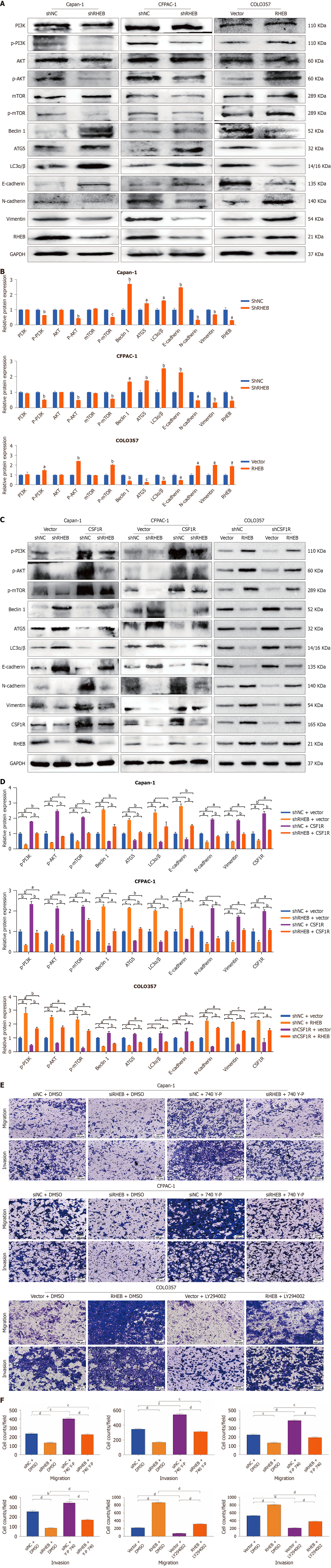
Total protein levels of markers (PI3K, AKT, and mTOR) showed no significant differences between RHEB-silenced and RHEB-overexpressed groups compared with the control (Figure 7A and B). However, a notable decrease in PI3K (p-PI3K), AKT (p-AKT), and mTOR (p-mTOR) protein phosphorylation extents was noted within RHEB-silenced groups, while an increase was observed within the RHEB-overexpressed group (Figure 7A and B). In replenishment experiments involving CSF1R overexpression and silencing, it was found that CSF1R overexpression elevated p-PI3K, p-AKT, and p-mTOR expressions, while its silencing caused the opposite trends. However, in these experiments, the magnitude of increases and decreases was comparatively smaller than that observed after RHEB silencing and overexpression within control groups (Figure 7C and D). These results suggested that RHEB might participate in PC metastasis by promoting PI3K, AKT, and mTOR phosphorylation levels.
To further verify the above hypothesis, a rescue experiment was performed. p-PI3K, being an upstream molecular segment of this axis, its activator (740 Y-P) and inhibitor (LY294002) were used for respectively silencing and overexpressing RHEB cell lines. In the Transwell assay, 740 Y-P enhanced Capan-1 and CFPAC-1 cell invasion and migration, but this phenomenon was less pronounced in the shRHEB groups than in the shNC groups. However, overexpressing RHEB promoted cell invasion and migration with or without LY294002, with the results being more significant than those in the vector group (Figure 7E and F). The verification results are consistent with the hypothesis.
Being an autophagy-related gene, it was speculated that RHEB could regulate PC metastasis through autophagy-related mechanisms. At the same time, given that previous studies established that p-PI3K, p-AKT, and p-mTOR are crucial in autophagy[13,22], subsequent investigations were focused on whether RHEB affects autophagy through these genes.
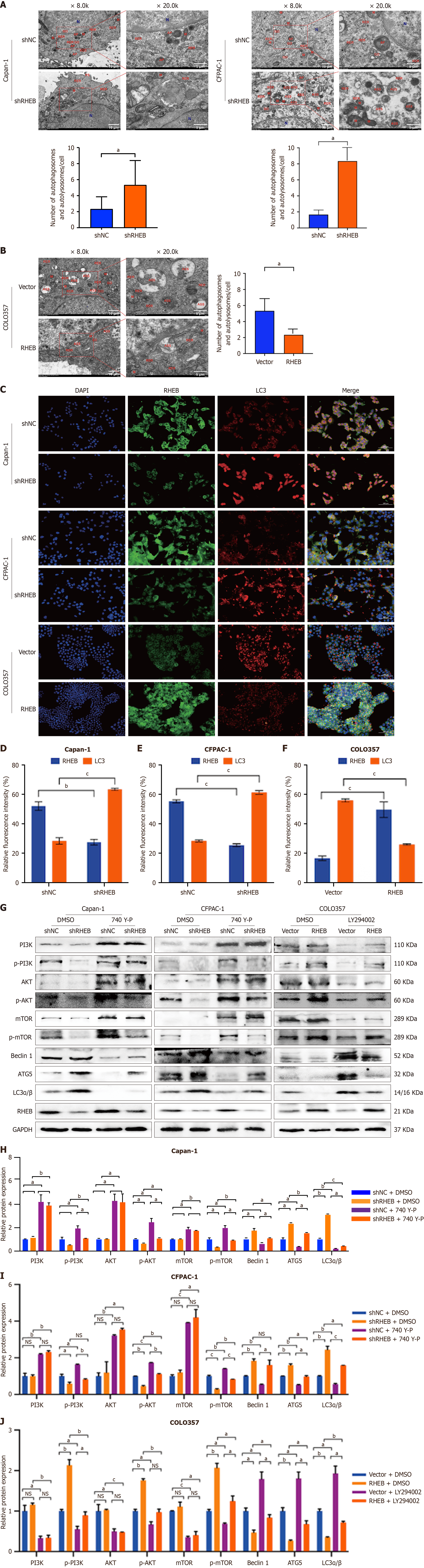
In this study, ATG5, Beclin 1, and LC3 were selected as autophagy markers before applying TEM and IF analyses for visualizing RHEB’s relationship with autophagy. According to the results, RHEB-silenced Capan-1 and CFPAC-1 cell lines had a higher total number of autophagosomes and autolysosomes than the control, while in the RHEB-overexpressing Colo357 cell line, these features were less in number compared with the control (Figure 8A and B). Indirect IF analysis further confirmed that silencing RHEB increased the expression of the autophagy process marker LC3 in Capan-1 and CFPAC-1 PC cell lines, while overexpressing the gene lowered LC3 expression in Colo357 cell lines (Figure 8C-F). Subsequent WB analyses confirmed that silencing RHEB enhanced Beclin 1, ATG5, and LC3 protein levels, with the opposite results also holding true when RHEB is overexpressed (Figure 7A, Supplementary Figure 7A). Based on the above, it was inferred that RHEB may inhibit the autophagy process.
To further confirm whether RHEB influences PC metastasis by mediating autophagy through the CSF1R/p-PI3K/p-AKT/p-mTOR axis, WB analyses were performed to clarify changes in autophagy marker levels in the rescue experiments. In replenishment experiments involving CSF1R overexpression and silencing, it was found that the former led to a decrease in Beclin1, ATG5, and LC3 expression, while silencing it resulted in opposite changes to those expressions. Meanwhile, the magnitude of variations in these expressions was relatively smaller than those observed after RHEB silencing/overexpression in the control group (Figure 7B, Supplementary Figure 7B). Likewise, despite insignificant changes in PI3K, AKT, and mTOR levels between the RHEB-silencing and control groups, the PI3K activator induced an increase in p-PI3K, p-AKT, and p-mTOR expression. At the same time, the decrease in Beclin 1, ATG5, and LC3 levels was significantly reversed by RHEB silencing. Moreover, in Colo357 cell lines where RHEB is stably overexpressed, p-PI3K, p-AKT, and p-mTOR upregulation coupled with Beclin 1, ATG5, and LC3 downregulation were remarkably diminished relative to the control group (Figure 8G-J). In this case, it was suggested that the RHEB-CSF1R complex might inhibit autophagy by phosphorylating PI3K, AKT, and mTOR, thereby further affecting the biological process of PC.
EMT represents a hallmark event of tumor metastasis[23], and existing evidence suggests that cells undergoing EMT need to be regulated by autophagy to survive the migration and dissemination process[24]. In Transwell assays, the RHEB-overexpressing cell lines exhibited an EMT-like cell morphology as they lost their epithelial characteristics, leading to a spindle-shape. However, when RHEB was silenced, it caused some morphological alterations from the scattered growth structures of tightly arranged colonies (Figure 4J-L). In addition, previous WB analyses showed that silencing RHEB increased E-cadherin levels (epithelial marker) while suppressing N-cadherin and vimentin (mesenchymal markers) levels. Opposite results were also observed when RHEB is overexpressed (Figure 7A, Supplementary Figure 7A). On the other hand, experiments involving CSF1R overexpression indicated that CSF1R overexpression could lead to elevated N-cadherin and vimentin levels but reduced E-cadherin expression. However, the magnitude of these changes in expression level was comparatively smaller than that achieved following RHEB silencing/overexpression within control groups (Figure 7B, Supplementary Figure 7B). Hence, it was speculated that RHEB possibly affected EMT markers through changes in the expression of autophagy markers, which regulate the process of PC metastasis.
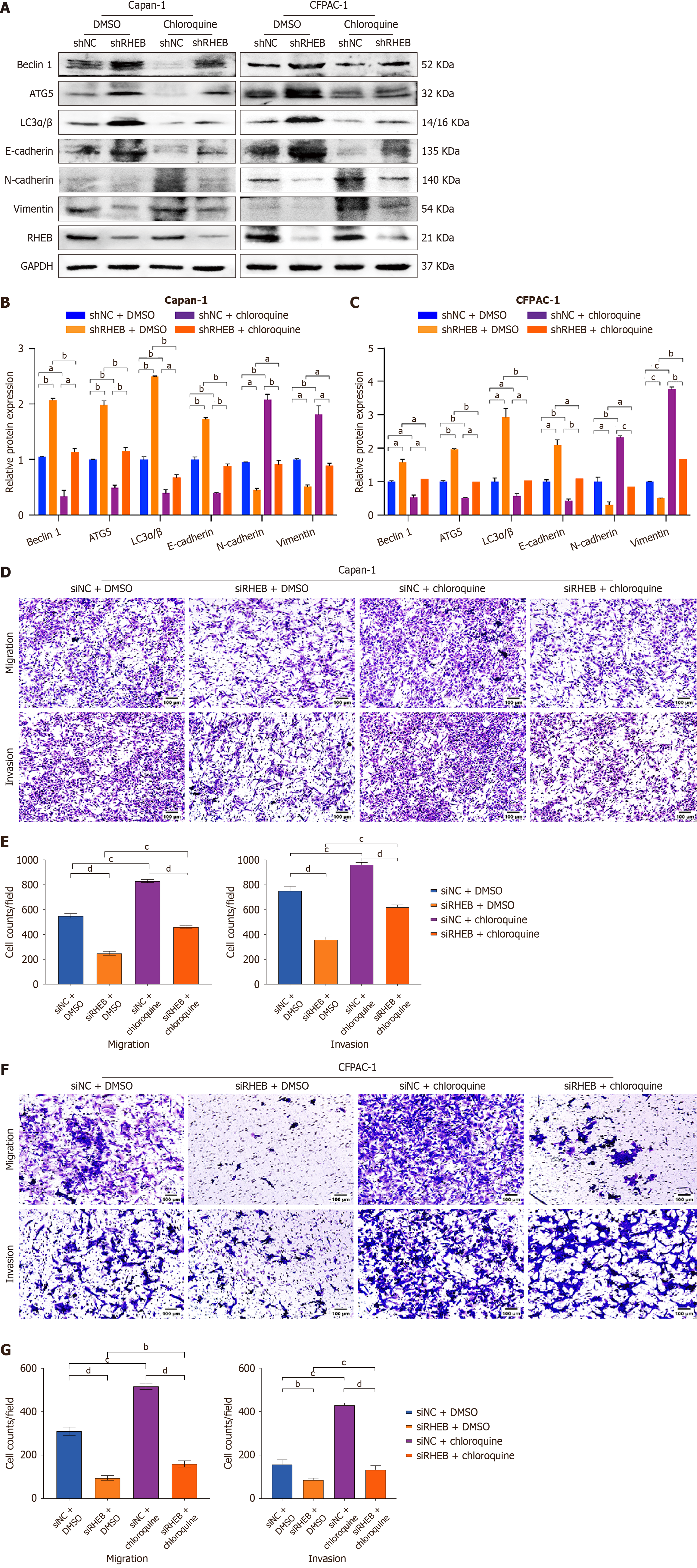
A rescue experiment was also undertaken to validate the above hypothesis. This was achieved using chloroquine, an autophagy inhibitor, for silencing RHEB in cell lines and their controls. After treatment with chloroquine, E-cadherin was downregulated relative to the control group. However, the shRHEB group maintained a higher E-cadherin expression than the shNC group. In contrast, N-cadherin and vimentin showed elevated expression with chloroquine treatment, and lower expression was observed in the shRHEB group than the shNC group (Figure 9A-C). Furthermore, chloroquine even potentiated the invading and migrating capacity of Capan-1 and CFPAC-1 cells, although these chloroquine-induced effects were compromised by RHEB silencing in vitro. In particular, the disparity within the siRHEB groups following chloroquine intervention was less significant as compared to the siNC groups (Figure 9D-G). The above results revealed that the RHEB-CSF1R complex might promote PC metastasis by upregulating the EMT markers N-cadherin and vimentin and downregulating E-cadherin via autophagy inhibition.
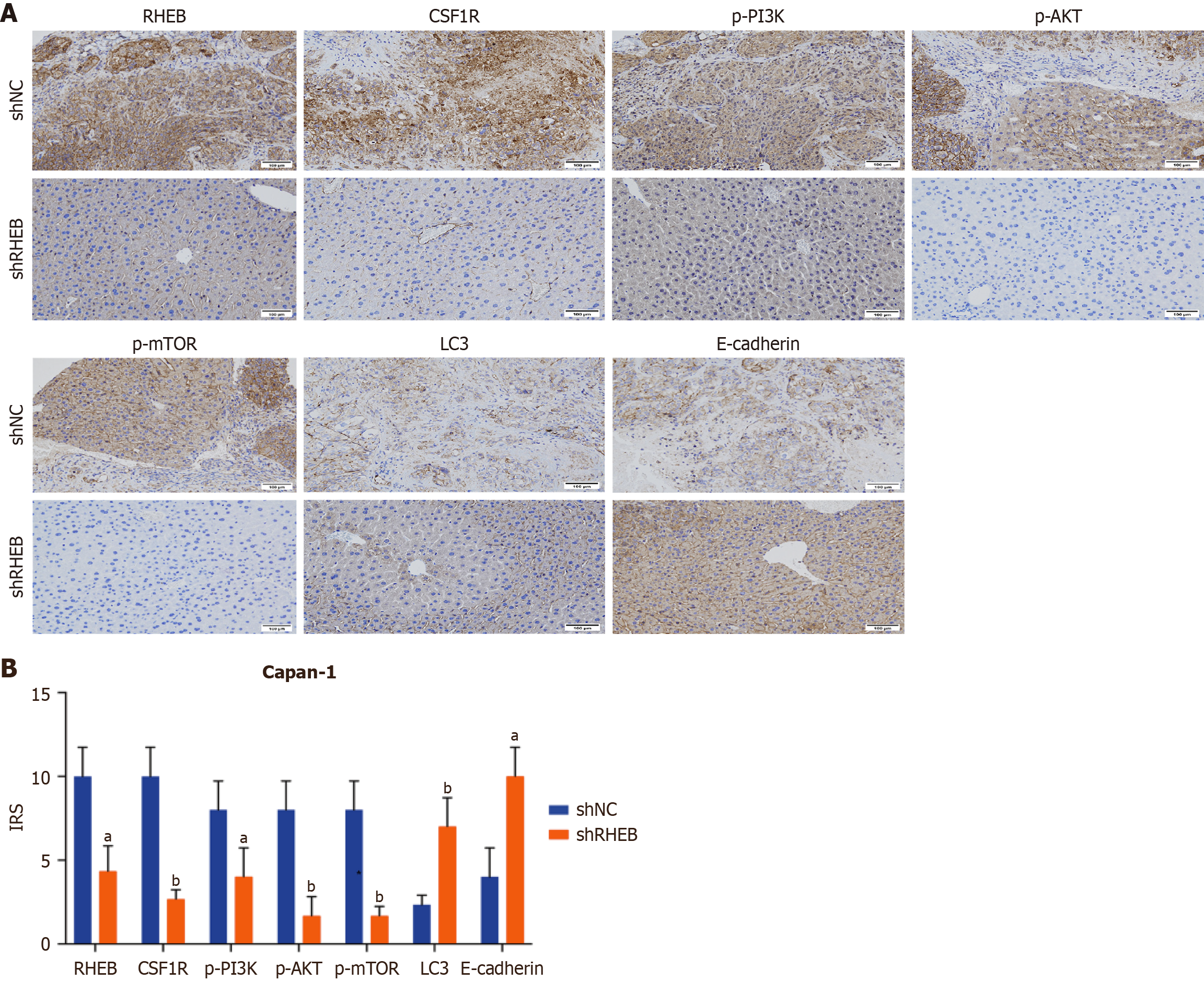
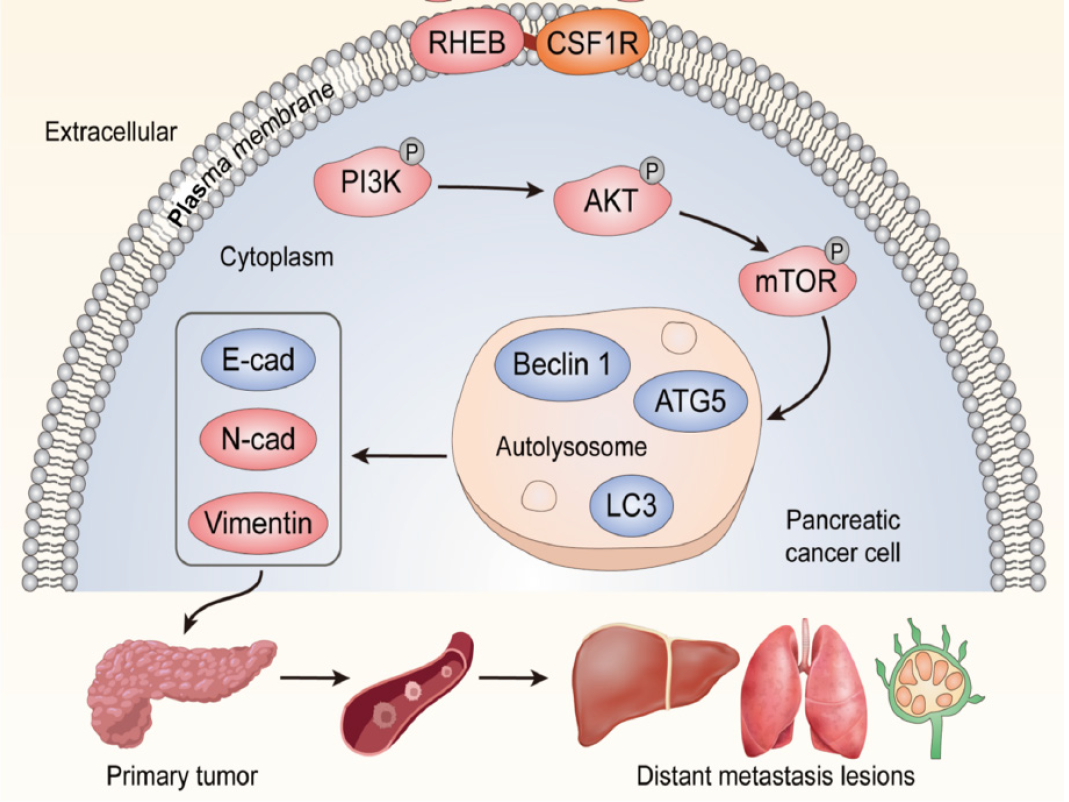
The mechanism of RHEB affecting PC metastasis was further validated. Immunohistochemical analysis was performed in a mouse distant liver metastasis model. The upregulation of RHEB, CSF1R, p-PI3K, p-AKT, and p-mTOR expression was confirmed using IHC. However, the downregulation of LC3 and E-cadherin was also observed in the shNC groups in comparison with those in the shRHEB groups (Figure 10). Altogether, these in vivo results suggest that silencing RHEB may inhibit PC metastasis, likely through changes in CSF1R expression and in PI3K, AKT, and mTOR phosphorylation. However, these mechanisms still require validation through further experiments. The above mechanistic study has been summarized and illustrated in Figure 11.
Over the last few decades, PC has dramatically increased the burden on global healthcare systems, and may represent a major cause of cancer-related mortality, especially due to susceptibility to metastasis[1]. Current studies suggest a close relationship between autophagy and tumor metastasis[5,6], and hence, identifying autophagy-relevant genes that are closely linked to PC metastasis and clarifying their mechanisms of action is crucial to improving survival rates and developing innovative forms of treatment. A previous study by the current authors proposed a prognostic model of autophagy-related genes to predict PC metastasis, with RHEB being one of the hub genes[25]. In addition, existing studies have indicated that RHEB is relevant to both autophagy and cancer[12,13]. Therefore, the role of this gene was further explored to understand its role in PC in view of improving prognosis and providing better treatments.
RHEB, one of the small GTPases related to Ras, Rap, and Ral[12], is an oncogene for tumor growth[26-28]. In various cancers, such as colorectal cancer[26], non-small cell lung carcinoma[27], and hepatocellular cancer[28], increased RHEB expression has been reported and linked to cancer progression. Similarly, in the case of PC, over-expression of RHEB enhanced cell proliferation, migration, and invasion[29], with elevated RHEB transcription and translation in PC confirmed by in vitro and in vivo experiments. Functional assays, including CCK-8, colony formation, and Transwell, revealed that RHEB can promote proliferation and metastasis in vitro, while its silencing suppressed distant liver metastasis in mice. Overall, the different experiments yielded consistent results. Moreover, high mRNA levels of RHEB were linked to PC poor prognosis[29]. In contrast, our results revealed that downregulating RHEB was implicated in superior prognosis in terms of both OS and recurrence-free survival in PC. In addition, the prognostic role of RHEB was significant regardless of sex age, tumor-node-metastasis stage, pathologic stage, and histologic grade. Taken together, these analyses involving databases, cells, and animal experiments demonstrated that RHEB expression promotes PC proliferation, migration, and invasion, and as such, it is not only strongly linked to cancer progression, but also leads to poor prognosis in PC patients. The small G protein RHEB can respond to growth factor signals and activate the mTOR complex 1 (mTORC1), which is a key regulator of cell growth and metabolism. Abnormal mTORC1 signaling is associated with fibrosis, metabolic and neurodegenerative diseases, cancer, rare diseases, and so on. When there is a point mutation in the switch II domain of RHEB, its ability to activate mTORC1 will be weakened[30]. The occurrence and development of many cancers are related to the abnormal activation of the RHEB/mTORC1 signaling pathway. In in vitro cell experiments, the use of RHEB inhibitors can significantly inhibit the proliferation, migration, and invasion abilities of tumor cells. For example, in cancer cell lines such as PC and colorectal cancer, after adding the P1_wt peptide inhibitor, the growth rate of cancer cells is significantly slowed down, the cell cycle is blocked, and at the same time, the migration and invasion abilities of the cells are also significantly inhibited, indicating that RHEB inhibitors have potential application value in cancer treatment.
Early metastasis is often responsible for the unsatisfactory prognosis of PC patients[1]. Therefore, subsequent experiments focused on the mechanism through which RHEB regulates PC metastasis. Different studies have identified translationally controlled tumor protein, phospholipase D1, and FK506 binding protein 38 as those proteins which interact with RHEB[31-33]. Translationally controlled tumor protein can interact with RHEB to accelerate the release of GDP and activate the mTORC1 signaling pathway[31]. Furthermore, Shen et al[32] showed that phospholipase D1 was required for activating RHEB within the mTOR signaling pathway, while RHEB bind to FK506 binding protein 38 to release mTOR and lead to its activation[33]. This study is the first one to identify RHEB and CSF1R as interacting proteins based on CO-IP and bioinformatics analyses, but the link between the two is yet to be reported.
CSF1R is part of the RTK family type III receptors, and when associated with tumors, it has been linked with poor survival[34]. Previous studies have found that in both hepatocellular carcinoma[35] and breast cancer[36], CSF1R played a role in promoting tumor cell metastasis. In this study, WB assays confirmed the upregulation of CSF1R through RHEB, while in replenishment experiments, CSF1R overexpression in RHEB-silenced PC cell lines highlighted the enhanced migratory and invasive potentials of PC cells after overexpressing CSF1R. In summary, the results confirmed that RHEB could upregulate and interact with CSF1R to influence PC metastasis.
To investigate the mechanism through which RHEB influences PC metastasis, functional enrichment analyses (GO, KEGG, GSEA and protein-protein interaction) of RHEB and CSF1R in PC were performed. The findings demonstrated that the mTOR, PI3K-AKT and insulin signaling pathways were enriched. In PC, the PI3K/AKT/mTOR pathway activation through PI3K, AKT and mTOR phosphorylation, promoted cell proliferation, growth and angiogenesis, thereby participating in cell biological activities such as invasion, apoptosis and metabolism[37-39]. This pathway has also been explored in autophagy[40], with the different results being consistent. Therefore, the ability of RHEB, as an autophagy-related gene, to regulate PC metastasis through this signaling pathway was explored. In this study, silencing/overexpressing RHEB or overexpressing CSF1R caused notable p-PI3K, p-AKT and p-mTOR levels. In vivo experiments showed consistent results with in vitro experiments in variations in p-AKT and p-mTOR levels. In Transwell rescue assays, overexpression of CSF1R and the use of PI3K activator/inhibitor influenced cell migration and invasion. RHEB’s potential to affect autophagy was then verified by TEM and IF. Autophagy marker (Beclin1, LC3, and ATG5) expressions were also observed using WB assays after silencing/overexpressing RHEB. Finally, in Transwell rescue assays, overexpression of CSF1R and the use of PI3K activator/inhibitor impacted cell migration and invasion. Altogether, the experimental results demonstrated that the RHEB-CSF1R complex may inhibit autophagy by phosphorylating PI3K, AKT and mTOR. RHEB is a GTPase that activates mTORC1 and has been identified as an oncoprotein[41]. Previous studies have demonstrated that silencing RHEB inhibits colon tumor cell growth by suppressing the mTOR pathway and accelerates apoptosis[29]. Additionally, studies have shown that elevated RHEB levels are associated with poor prognosis in PC patients[42]. Similar to other studies, Deng et al’s research[43] confirms the role of RHEB in driving PC progression and metastasis.
EMT represents a hallmark event of tumor metastasis[23], and autophagy is vital in regulating this EMT process[24]. Therefore, this work also explored whether RHEB, as an autophagy-related gene, could regulate EMT through the autophagy process and thus, regulate PC metastasis. In this study, WB assays first confirmed that RHEB influenced the expression patterns of the dominating EMT indicators E-cadherin, N-cadherin, and vimentin. Moreover, in replenishment experiments that involved chloroquine as an inhibitor of CSF1R overexpression and autophagy, upregulated N-cadherin and vimentin expressions, and downregulated E-cadherin expression were confirmed. In Transwell assays, chloroquine exerted in vitro pro-invasion and pro-migration effects. These different results revealed that the RHEB-CSF1R complex may promote EMT marker expression through autophagy inhibitor.
Despite the innovative findings of this study, this work was not without limitations. Firstly, the specific binding site through which RHEB and CSF1R interact needs to be further explored. While Co-IP experiments confirm their binding, without establishing specific binding domains (e.g., whether RHEB’s GTP-binding domain or CSF1R’s intracellular kinase domain participates) through truncated mutant constructs. The effect of GTP-binding status on RHEB-CSF1R interaction was also not examined. Secondly, the underlying mechanism that leads to RHEB’s upregulation in PC metastatic tissues needs to be elucidated before exploring the potential of upstream regulation of RHEB as a novel therapeutic strategy for PC. The specific role of PI3K subtypes (e.g., P110β) was not thoroughly explored, leaving unclear whether other subtypes also participate in this pathway. Finally, additional in vivo experiments are required to determine whether RHEB influences specific mechanisms of PC metastasis via the CSF1R-phosphorylated PI3K/AKT/mTOR axis. Also, in vivo experiments employed a liver metastasis model, but lacked quantitative analysis of RHEB/CSF1R expression in metastatic lesions. Therefore, a well-designed comprehensive study needs to further confirm the results in this study.
In conclusion, this study revealed a potential mechanism through which RHEB enhanced PC metastasis, with increased RHEB expression also reducing PC patients’ survival. In addition, RHEB expression promoted PC proliferation, migration, and invasive ability. RHEB was also able to inhibit autophagy by upregulating and interacting with CSF1R, leading to phosphorylation of PI3K, AKT, and mTOR, which further promoted PC metastasis by elevating the EMT marker expression. These findings suggest that RHEB can be a promising prognostic biomarker and a possible target for developing new prevention and treatment strategies in PC.
Experiments of the current study were supported by the First Affiliated Hospital of Chongqing Medical University Key Laboratory of Molecular Oncology and Epigenetics.
| 1. | Klein AP. Pancreatic cancer epidemiology: understanding the role of lifestyle and inherited risk factors. Nat Rev Gastroenterol Hepatol. 2021;18:493-502. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1039] [Cited by in RCA: 915] [Article Influence: 183.0] [Reference Citation Analysis (0)] |
| 2. | Halbrook CJ, Lyssiotis CA, Pasca di Magliano M, Maitra A. Pancreatic cancer: Advances and challenges. Cell. 2023;186:1729-1754. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1043] [Cited by in RCA: 910] [Article Influence: 303.3] [Reference Citation Analysis (2)] |
| 3. | Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72:7-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13024] [Cited by in RCA: 12107] [Article Influence: 3026.8] [Reference Citation Analysis (6)] |
| 4. | Global Burden of Disease 2019 Cancer Collaboration, Kocarnik JM, Compton K, Dean FE, Fu W, Gaw BL, Harvey JD, Henrikson HJ, Lu D, Pennini A, Xu R, Ababneh E, Abbasi-Kangevari M, Abbastabar H, Abd-Elsalam SM, Abdoli A, Abedi A, Abidi H, Abolhassani H, Adedeji IA, Adnani QES, Advani SM, Afzal MS, Aghaali M, Ahinkorah BO, Ahmad S, Ahmad T, Ahmadi A, Ahmadi S, Ahmed Rashid T, Ahmed Salih Y, Akalu GT, Aklilu A, Akram T, Akunna CJ, Al Hamad H, Alahdab F, Al-Aly Z, Ali S, Alimohamadi Y, Alipour V, Aljunid SM, Alkhayyat M, Almasi-Hashiani A, Almasri NA, Al-Maweri SAA, Almustanyir S, Alonso N, Alvis-Guzman N, Amu H, Anbesu EW, Ancuceanu R, Ansari F, Ansari-Moghaddam A, Antwi MH, Anvari D, Anyasodor AE, Aqeel M, Arabloo J, Arab-Zozani M, Aremu O, Ariffin H, Aripov T, Arshad M, Artaman A, Arulappan J, Asemi Z, Asghari Jafarabadi M, Ashraf T, Atorkey P, Aujayeb A, Ausloos M, Awedew AF, Ayala Quintanilla BP, Ayenew T, Azab MA, Azadnajafabad S, Azari Jafari A, Azarian G, Azzam AY, Badiye AD, Bahadory S, Baig AA, Baker JL, Balakrishnan S, Banach M, Bärnighausen TW, Barone-Adesi F, Barra F, Barrow A, Behzadifar M, Belgaumi UI, Bezabhe WMM, Bezabih YM, Bhagat DS, Bhagavathula AS, Bhardwaj N, Bhardwaj P, Bhaskar S, Bhattacharyya K, Bhojaraja VS, Bibi S, Bijani A, Biondi A, Bisignano C, Bjørge T, Bleyer A, Blyuss O, Bolarinwa OA, Bolla SR, Braithwaite D, Brar A, Brenner H, Bustamante-Teixeira MT, Butt NS, Butt ZA, Caetano Dos Santos FL, Cao Y, Carreras G, Catalá-López F, Cembranel F, Cerin E, Cernigliaro A, Chakinala RC, Chattu SK, Chattu VK, Chaturvedi P, Chimed-Ochir O, Cho DY, Christopher DJ, Chu DT, Chung MT, Conde J, Cortés S, Cortesi PA, Costa VM, Cunha AR, Dadras O, Dagnew AB, Dahlawi SMA, Dai X, Dandona L, Dandona R, Darwesh AM, das Neves J, De la Hoz FP, Demis AB, Denova-Gutiérrez E, Dhamnetiya D, Dhimal ML, Dhimal M, Dianatinasab M, Diaz D, Djalalinia S, Do HP, Doaei S, Dorostkar F, Dos Santos Figueiredo FW, Driscoll TR, Ebrahimi H, Eftekharzadeh S, El Tantawi M, El-Abid H, Elbarazi I, Elhabashy HR, Elhadi M, El-Jaafary SI, Eshrati B, Eskandarieh S, Esmaeilzadeh F, Etemadi A, Ezzikouri S, Faisaluddin M, Faraon EJA, Fares J, Farzadfar F, Feroze AH, Ferrero S, Ferro Desideri L, Filip I, Fischer F, Fisher JL, Foroutan M, Fukumoto T, Gaal PA, Gad MM, Gadanya MA, Gallus S, Gaspar Fonseca M, Getachew Obsa A, Ghafourifard M, Ghashghaee A, Ghith N, Gholamalizadeh M, Gilani SA, Ginindza TG, Gizaw ATT, Glasbey JC, Golechha M, Goleij P, Gomez RS, Gopalani SV, Gorini G, Goudarzi H, Grosso G, Gubari MIM, Guerra MR, Guha A, Gunasekera DS, Gupta B, Gupta VB, Gupta VK, Gutiérrez RA, Hafezi-Nejad N, Haider MR, Haj-Mirzaian A, Halwani R, Hamadeh RR, Hameed S, Hamidi S, Hanif A, Haque S, Harlianto NI, Haro JM, Hasaballah AI, Hassanipour S, Hay RJ, Hay SI, Hayat K, Heidari G, Heidari M, Herrera-Serna BY, Herteliu C, Hezam K, Holla R, Hossain MM, Hossain MBH, Hosseini MS, Hosseini M, Hosseinzadeh M, Hostiuc M, Hostiuc S, Househ M, Hsairi M, Huang J, Hugo FN, Hussain R, Hussein NR, Hwang BF, Iavicoli I, Ibitoye SE, Ida F, Ikuta KS, Ilesanmi OS, Ilic IM, Ilic MD, Irham LM, Islam JY, Islam RM, Islam SMS, Ismail NE, Isola G, Iwagami M, Jacob L, Jain V, Jakovljevic MB, Javaheri T, Jayaram S, Jazayeri SB, Jha RP, Jonas JB, Joo T, Joseph N, Joukar F, Jürisson M, Kabir A, Kahrizi D, Kalankesh LR, Kalhor R, Kaliyadan F, Kalkonde Y, Kamath A, Kameran Al-Salihi N, Kandel H, Kapoor N, Karch A, Kasa AS, Katikireddi SV, Kauppila JH, Kavetskyy T, Kebede SA, Keshavarz P, Keykhaei M, Khader YS, Khalilov R, Khan G, Khan M, Khan MN, Khan MAB, Khang YH, Khater AM, Khayamzadeh M, Kim GR, Kim YJ, Kisa A, Kisa S, Kissimova-Skarbek K, Kopec JA, Koteeswaran R, Koul PA, Koulmane Laxminarayana SL, Koyanagi A, Kucuk Bicer B, Kugbey N, Kumar GA, Kumar N, Kumar N, Kurmi OP, Kutluk T, La Vecchia C, Lami FH, Landires I, Lauriola P, Lee SW, Lee SWH, Lee WC, Lee YH, Leigh J, Leong E, Li J, Li MC, Liu X, Loureiro JA, Lunevicius R, Magdy Abd El Razek M, Majeed A, Makki A, Male S, Malik AA, Mansournia MA, Martini S, Masoumi SZ, Mathur P, McKee M, Mehrotra R, Mendoza W, Menezes RG, Mengesha EW, Mesregah MK, Mestrovic T, Miao Jonasson J, Miazgowski B, Miazgowski T, Michalek IM, Miller TR, Mirzaei H, Mirzaei HR, Misra S, Mithra P, Moghadaszadeh M, Mohammad KA, Mohammad Y, Mohammadi M, Mohammadi SM, Mohammadian-Hafshejani A, Mohammed S, Moka N, Mokdad AH, Molokhia M, Monasta L, Moni MA, Moosavi MA, Moradi Y, Moraga P, Morgado-da-Costa J, Morrison SD, Mosapour A, Mubarik S, Mwanri L, Nagarajan AJ, Nagaraju SP, Nagata C, Naimzada MD, Nangia V, Naqvi AA, Narasimha Swamy S, Ndejjo R, Nduaguba SO, Negoi I, Negru SM, Neupane Kandel S, Nguyen CT, Nguyen HLT, Niazi RK, Nnaji CA, Noor NM, Nuñez-Samudio V, Nzoputam CI, Oancea B, Ochir C, Odukoya OO, Ogbo FA, Olagunju AT, Olakunde BO, Omar E, Omar Bali A, Omonisi AEE, Ong S, Onwujekwe OE, Orru H, Ortega-Altamirano DV, Otstavnov N, Otstavnov SS, Owolabi MO, P A M, Padubidri JR, Pakshir K, Pana A, Panagiotakos D, Panda-Jonas S, Pardhan S, Park EC, Park EK, Pashazadeh Kan F, Patel HK, Patel JR, Pati S, Pattanshetty SM, Paudel U, Pereira DM, Pereira RB, Perianayagam A, Pillay JD, Pirouzpanah S, Pishgar F, Podder I, Postma MJ, Pourjafar H, Prashant A, Preotescu L, Rabiee M, Rabiee N, Radfar A, Radhakrishnan RA, Radhakrishnan V, Rafiee A, Rahim F, Rahimzadeh S, Rahman M, Rahman MA, Rahmani AM, Rajai N, Rajesh A, Rakovac I, Ram P, Ramezanzadeh K, Ranabhat K, Ranasinghe P, Rao CR, Rao SJ, Rawassizadeh R, Razeghinia MS, Renzaho AMN, Rezaei N, Rezaei N, Rezapour A, Roberts TJ, Rodriguez JAB, Rohloff P, Romoli M, Ronfani L, Roshandel G, Rwegerera GM, S M, Sabour S, Saddik B, Saeed U, Sahebkar A, Sahoo H, Salehi S, Salem MR, Salimzadeh H, Samaei M, Samy AM, Sanabria J, Sankararaman S, Santric-Milicevic MM, Sardiwalla Y, Sarveazad A, Sathian B, Sawhney M, Saylan M, Schneider IJC, Sekerija M, Seylani A, Shafaat O, Shaghaghi Z, Shaikh MA, Shamsoddin E, Shannawaz M, Sharma R, Sheikh A, Sheikhbahaei S, Shetty A, Shetty JK, Shetty PH, Shibuya K, Shirkoohi R, Shivakumar KM, Shivarov V, Siabani S, Siddappa Malleshappa SK, Silva DAS, Singh JA, Sintayehu Y, Skryabin VY, Skryabina AA, Soeberg MJ, Sofi-Mahmudi A, Sotoudeh H, Steiropoulos P, Straif K, Subedi R, Sufiyan MB, Sultan I, Sultana S, Sur D, Szerencsés V, Szócska M, Tabarés-Seisdedos R, Tabuchi T, Tadbiri H, Taherkhani A, Takahashi K, Talaat IM, Tan KK, Tat VY, Tedla BAA, Tefera YG, Tehrani-Banihashemi A, Temsah MH, Tesfay FH, Tessema GA, Thapar R, Thavamani A, Thoguluva Chandrasekar V, Thomas N, Tohidinik HR, Touvier M, Tovani-Palone MR, Traini E, Tran BX, Tran KB, Tran MTN, Tripathy JP, Tusa BS, Ullah I, Ullah S, Umapathi KK, Unnikrishnan B, Upadhyay E, Vacante M, Vaezi M, Valadan Tahbaz S, Velazquez DZ, Veroux M, Violante FS, Vlassov V, Vo B, Volovici V, Vu GT, Waheed Y, Wamai RG, Ward P, Wen YF, Westerman R, Winkler AS, Yadav L, Yahyazadeh Jabbari SH, Yang L, Yaya S, Yazie TSY, Yeshaw Y, Yonemoto N, Younis MZ, Yousefi Z, Yu C, Yuce D, Yunusa I, Zadnik V, Zare F, Zastrozhin MS, Zastrozhina A, Zhang J, Zhong C, Zhou L, Zhu C, Ziapour A, Zimmermann IR, Fitzmaurice C, Murray CJL, Force LM. Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life Years for 29 Cancer Groups From 2010 to 2019: A Systematic Analysis for the Global Burden of Disease Study 2019. JAMA Oncol. 2022;8:420-444. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1649] [Cited by in RCA: 1454] [Article Influence: 363.5] [Reference Citation Analysis (4)] |
| 5. | Mizushima N, Levine B. Autophagy in Human Diseases. N Engl J Med. 2020;383:1564-1576. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 995] [Cited by in RCA: 868] [Article Influence: 144.7] [Reference Citation Analysis (3)] |
| 6. | Klionsky DJ, Petroni G, Amaravadi RK, Baehrecke EH, Ballabio A, Boya P, Bravo-San Pedro JM, Cadwell K, Cecconi F, Choi AMK, Choi ME, Chu CT, Codogno P, Colombo MI, Cuervo AM, Deretic V, Dikic I, Elazar Z, Eskelinen EL, Fimia GM, Gewirtz DA, Green DR, Hansen M, Jäättelä M, Johansen T, Juhász G, Karantza V, Kraft C, Kroemer G, Ktistakis NT, Kumar S, Lopez-Otin C, Macleod KF, Madeo F, Martinez J, Meléndez A, Mizushima N, Münz C, Penninger JM, Perera RM, Piacentini M, Reggiori F, Rubinsztein DC, Ryan KM, Sadoshima J, Santambrogio L, Scorrano L, Simon HU, Simon AK, Simonsen A, Stolz A, Tavernarakis N, Tooze SA, Yoshimori T, Yuan J, Yue Z, Zhong Q, Galluzzi L, Pietrocola F. Autophagy in major human diseases. EMBO J. 2021;40:e108863. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1393] [Cited by in RCA: 1224] [Article Influence: 244.8] [Reference Citation Analysis (5)] |
| 7. | Ferretti GDS, Quaas CE, Bertolini I, Zuccotti A, Saatci O, Kashatus JA, Sharmin S, Lu DY, Poli ANR, Quesnelle AF, Rodriguez-Blanco J, de Cubas AA, Hobbs GA, Liu Q, O'Bryan JP, Salvino JM, Kashatus DF, Sahin O, Barnoud T. HSP70-mediated mitochondrial dynamics and autophagy represent a novel vulnerability in pancreatic cancer. Cell Death Differ. 2024;31:881-896. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 32] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 8. | Kalli M, Mpekris F, Charalambous A, Michael C, Stylianou C, Voutouri C, Hadjigeorgiou AG, Papoui A, Martin JD, Stylianopoulos T. Mechanical forces inducing oxaliplatin resistance in pancreatic cancer can be targeted by autophagy inhibition. Commun Biol. 2024;7:1581. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 9. | Oyama K, Nakata K, Tsutsumi C, Hayashi M, Zhang B, Mochida Y, Shinkawa T, Hirotaka K, Zhong P, Date S, Luo H, Kubo A, Higashijima N, Yamada Y, Abe T, Ideno N, Koikawa K, Iwamoto C, Ikenaga N, Ohuchida K, Onishi H, Morisaki T, Kuba K, Oda Y, Nakamura M. Combined Autophagy Inhibition and Dendritic Cell Recruitment Induces Antitumor Immunity and Enhances Immune Checkpoint Blockade Sensitivity in Pancreatic Cancer. Cancer Res. 2024;84:4214-4232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 10. | Liang C, Xu J, Meng Q, Zhang B, Liu J, Hua J, Zhang Y, Shi S, Yu X. TGFB1-induced autophagy affects the pattern of pancreatic cancer progression in distinct ways depending on SMAD4 status. Autophagy. 2020;16:486-500. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 79] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 11. | Yamamoto K, Venida A, Yano J, Biancur DE, Kakiuchi M, Gupta S, Sohn ASW, Mukhopadhyay S, Lin EY, Parker SJ, Banh RS, Paulo JA, Wen KW, Debnath J, Kim GE, Mancias JD, Fearon DT, Perera RM, Kimmelman AC. Autophagy promotes immune evasion of pancreatic cancer by degrading MHC-I. Nature. 2020;581:100-105. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1047] [Cited by in RCA: 980] [Article Influence: 163.3] [Reference Citation Analysis (4)] |
| 12. | Rahman M, Nguyen TM, Lee GJ, Kim B, Park MK, Lee CH. Unraveling the Role of Ras Homolog Enriched in Brain (Rheb1 and Rheb2): Bridging Neuronal Dynamics and Cancer Pathogenesis through Mechanistic Target of Rapamycin Signaling. Int J Mol Sci. 2024;25:1489. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 13. | Foerster EG, Mukherjee T, Cabral-Fernandes L, Rocha JDB, Girardin SE, Philpott DJ. How autophagy controls the intestinal epithelial barrier. Autophagy. 2022;18:86-103. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 338] [Cited by in RCA: 307] [Article Influence: 76.8] [Reference Citation Analysis (0)] |
| 14. | Zhang H, Xia P, Liu J, Chen Z, Ma W, Yuan Y. ATIC inhibits autophagy in hepatocellular cancer through the AKT/FOXO3 pathway and serves as a prognostic signature for modeling patient survival. Int J Biol Sci. 2021;17:4442-4458. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 50] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 15. | Liu X, Wang X, Chai B, Wu Z, Gu Z, Zou H, Zhang H, Li Y, Sun Q, Fang W, Ma Z. miR-199a-3p/5p regulate tumorgenesis via targeting Rheb in non-small cell lung cancer. Int J Biol Sci. 2022;18:4187-4202. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 29] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 16. | Hu D, Jiang L, Luo S, Zhao X, Hu H, Zhao G, Tang W. Development of an autophagy-related gene expression signature for prognosis prediction in prostate cancer patients. J Transl Med. 2020;18:160. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 81] [Cited by in RCA: 77] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 17. | Chun YS, Pawlik TM, Vauthey JN. 8th Edition of the AJCC Cancer Staging Manual: Pancreas and Hepatobiliary Cancers. Ann Surg Oncol. 2018;25:845-847. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 637] [Cited by in RCA: 600] [Article Influence: 75.0] [Reference Citation Analysis (4)] |
| 18. | Sule R, Rivera G, Gomes AV. Western blotting (immunoblotting): history, theory, uses, protocol and problems. Biotechniques. 2023;75:99-114. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 101] [Article Influence: 33.7] [Reference Citation Analysis (0)] |
| 19. | Shi X, Kaller M, Rokavec M, Kirchner T, Horst D, Hermeking H. Characterization of a p53/miR-34a/CSF1R/STAT3 Feedback Loop in Colorectal Cancer. Cell Mol Gastroenterol Hepatol. 2020;10:391-418. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 81] [Article Influence: 13.5] [Reference Citation Analysis (4)] |
| 20. | Cersosimo F, Lonardi S, Ulivieri C, Martini P, Morrione A, Vermi W, Giordano A, Giurisato E. CSF-1R in Cancer: More than a Myeloid Cell Receptor. Cancers (Basel). 2024;16:282. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 27] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 21. | Mehta AK, Cheney EM, Hartl CA, Pantelidou C, Oliwa M, Castrillon JA, Lin JR, Hurst KE, de Oliveira Taveira M, Johnson NT, Oldham WM, Kalocsay M, Berberich MJ, Boswell SA, Kothari A, Johnson S, Dillon DA, Lipschitz M, Rodig S, Santagata S, Garber JE, Tung N, Yélamos J, Thaxton JE, Mittendorf EA, Sorger PK, Shapiro GI, Guerriero JL. Targeting immunosuppressive macrophages overcomes PARP inhibitor resistance in BRCA1-associated triple-negative breast cancer. Nat Cancer. 2021;2:66-82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 110] [Cited by in RCA: 203] [Article Influence: 40.6] [Reference Citation Analysis (0)] |
| 22. | Fattahi S, Amjadi-Moheb F, Tabaripour R, Ashrafi GH, Akhavan-Niaki H. PI3K/AKT/mTOR signaling in gastric cancer: Epigenetics and beyond. Life Sci. 2020;262:118513. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 343] [Cited by in RCA: 293] [Article Influence: 48.8] [Reference Citation Analysis (7)] |
| 23. | Bakir B, Chiarella AM, Pitarresi JR, Rustgi AK. EMT, MET, Plasticity, and Tumor Metastasis. Trends Cell Biol. 2020;30:764-776. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 835] [Cited by in RCA: 777] [Article Influence: 129.5] [Reference Citation Analysis (7)] |
| 24. | Gundamaraju R, Lu W, Paul MK, Jha NK, Gupta PK, Ojha S, Chattopadhyay I, Rao PV, Ghavami S. Autophagy and EMT in cancer and metastasis: Who controls whom? Biochim Biophys Acta Mol Basis Dis. 2022;1868:166431. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 120] [Article Influence: 30.0] [Reference Citation Analysis (0)] |
| 25. | Deng J, Zhang Q, Lv L, Ma P, Zhang Y, Zhao N, Zhang Y. Identification of an autophagy-related gene signature for predicting prognosis and immune activity in pancreatic adenocarcinoma. Sci Rep. 2022;12:7006. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 26. | Turcu S, Grama F, Gazouli M. Gut Microbiome-Mediated Genetic and Epigenetic Alterations in Colorectal Cancer: Population-Specific Insights. Biomedicines. 2025;13:2262. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 27. | Liu B, Zhang J, Meng X, Xie SM, Liu F, Chen H, Yao D, Li M, Guo M, Shen H, Zhang X, Xing L. HDAC6-G3BP2 promotes lysosomal-TSC2 and suppresses mTORC1 under ETV4 targeting-induced low-lactate stress in non-small cell lung cancer. Oncogene. 2023;42:1181-1195. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 21] [Reference Citation Analysis (0)] |
| 28. | Dong P, Wang X, Liu L, Tang W, Ma L, Zeng W, Sun S, Zhang L, Zhang N, Shen X, Janssen HLA, Dong L, Zhang S, Chen S. Dampened VEPH1 activates mTORC1 signaling by weakening the TSC1/TSC2 association in hepatocellular carcinoma. J Hepatol. 2020;73:1446-1459. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 29. | Tan J, Liu W, Li J, Zhang X, Liu Y, Yuan Y, Song Z. Over-expressed RHEB promotes the progression of pancreatic adenocarcinoma. Life Sci. 2021;277:119462. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 30. | Fu W, Wu G. Design of negative-regulating proteins of Rheb/mTORC1 with much-reduced sizes of the tuberous sclerosis protein complex. Protein Sci. 2023;32:e4731. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 31. | Rahman M, Nguyen TM, Lee GJ, Kim B, Park MK, Lee CH. Role of Rheb1 and Rheb2 in Cancer Development and Therapeutic Prospects. 2023 Preprint. [DOI] [Full Text] |
| 32. | Shen JZ, Wu G, Guo S. Amino Acids in Autophagy: Regulation and Function. Adv Exp Med Biol. 2021;1332:51-66. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 33. | De Cicco M, Kiss L, Dames SA. NMR analysis of the backbone dynamics of the small GTPase Rheb and its interaction with the regulatory protein FKBP38. FEBS Lett. 2018;592:130-146. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 34. | Larroquette M, Guegan JP, Besse B, Cousin S, Brunet M, Le Moulec S, Le Loarer F, Rey C, Soria JC, Barlesi F, Bessede A, Scoazec JY, Soubeyran I, Italiano A. Spatial transcriptomics of macrophage infiltration in non-small cell lung cancer reveals determinants of sensitivity and resistance to anti-PD1/PD-L1 antibodies. J Immunother Cancer. 2022;10:e003890. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 114] [Article Influence: 28.5] [Reference Citation Analysis (4)] |
| 35. | He Q, Liu M, Huang W, Chen X, Zhang B, Zhang T, Wang Y, Liu D, Xie M, Ji X, Sun M, Tian D, Xia L. IL-1β-Induced Elevation of Solute Carrier Family 7 Member 11 Promotes Hepatocellular Carcinoma Metastasis Through Up-regulating Programmed Death Ligand 1 and Colony-Stimulating Factor 1. Hepatology. 2021;74:3174-3193. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 131] [Article Influence: 26.2] [Reference Citation Analysis (0)] |
| 36. | Klemm F, Möckl A, Salamero-Boix A, Alekseeva T, Schäffer A, Schulz M, Niesel K, Maas RR, Groth M, Elie BT, Bowman RL, Hegi ME, Daniel RT, Zeiner PS, Zinke J, Harter PN, Plate KH, Joyce JA, Sevenich L. Compensatory CSF2-driven macrophage activation promotes adaptive resistance to CSF1R inhibition in breast-to-brain metastasis. Nat Cancer. 2021;2:1086-1101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 106] [Article Influence: 21.2] [Reference Citation Analysis (0)] |
| 37. | Chen J, Liu F, Wu J, Yang Y, He J, Wu F, Yang K, Li J, Jiang Z, Jiang Z. Effect of STK3 on proliferation and apoptosis of pancreatic cancer cells via PI3K/AKT/mTOR pathway. Cell Signal. 2023;106:110642. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 38. | Cui J, Guo Y, Wu H, Xiong J, Peng T. Everolimus regulates the activity of gemcitabine-resistant pancreatic cancer cells by targeting the Warburg effect via PI3K/AKT/mTOR signaling. Mol Med. 2021;27:38. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 36] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 39. | Mizrahi JD, Surana R, Valle JW, Shroff RT. Pancreatic cancer. Lancet. 2020;395:2008-2020. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2196] [Cited by in RCA: 1909] [Article Influence: 318.2] [Reference Citation Analysis (4)] |
| 40. | Xu Z, Han X, Ou D, Liu T, Li Z, Jiang G, Liu J, Zhang J. Targeting PI3K/AKT/mTOR-mediated autophagy for tumor therapy. Appl Microbiol Biotechnol. 2020;104:575-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 544] [Cited by in RCA: 494] [Article Influence: 82.3] [Reference Citation Analysis (4)] |
| 41. | Tian Y, Shen L, Li F, Yang J, Wan X, Ouyang M. Silencing of RHEB inhibits cell proliferation and promotes apoptosis in colorectal cancer cells via inhibition of the mTOR signaling pathway. J Cell Physiol. 2020;235:442-453. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 30] [Article Influence: 4.3] [Reference Citation Analysis (3)] |
| 42. | Zhang Q, Lv L, Ma P, Zhang Y, Deng J, Zhang Y. Identification of an Autophagy-Related Pair Signature for Predicting Prognoses and Immune Activity in Pancreatic Adenocarcinoma. Front Immunol. 2021;12:743938. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 43. | Deng Q, Wu L, He J, Wu F, Jiang Z. Identification of autophagy-related immune targets for enhancing immunotherapy in pancreatic cancer aggressiveness. Discov Oncol. 2025;16:382. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |