Published online Feb 28, 2020. doi: 10.3748/wjg.v26.i8.789
Peer-review started: October 3, 2019
First decision: November 10, 2019
Revised: December 20, 2019
Accepted: February 12, 2020
Article in press: February 12, 2020
Published online: February 28, 2020
Processing time: 147 Days and 18.3 Hours
Despite significant advances in multimodality treatments, hepatocellular carcinoma (HCC) remains one of the most common malignant tumors. Identification of novel prognostic biomarkers and molecular targets is urgently needed.
To identify potential key genes associated with tumor microenvironments and the prognosis of HCC.
The infiltration levels of immune cells and stromal cells were calculated and quantified based on the ESTIMATE algorithm. Differentially expressed genes (DEGs) between high and low groups according to immune or stromal scores were screened using the gene expression profile of HCC patients in The Cancer Genome Atlas and were further linked to the prognosis of HCC. These genes were validated in four independent HCC cohorts. Survival-related key genes were identified by a LASSO Cox regression model.
HCC patients with a high immune/stromal score had better survival benefits than patients with a low score. A total of 899 DEGs were identified and found to be involved in immune responses and extracellular matrices, 147 of which were associated with overall survival. Subsequently, 52 of 147 survival-related DEGs were validated in additional cohorts. Finally, ten key genes (STSL2, TMC5, DOK5, RASGRP2, NLRC3, KLRB1, CD5L, CFHR3, ADH1C, and UGT2B15) were selected and used to construct a prognostic gene signature, which presented a good performance in predicting overall survival.
This study extracted a list of genes associated with tumor microenvironments and the prognosis of HCC, thereby providing several valuable directions for the prognostic prediction and molecular targeted therapy of HCC in the future.
Core tip: We performed an integrated bioinformatics analysis to assess the influence of tumor microenvironment on the prognosis of patients with hepatocellular carcinoma (HCC). The results found that HCC patients with a high immune/stromal infiltration level had better survival benefits than patients with a low infiltration level. Ten microenvironment-related key genes were screened and used to construct a prognostic gene signature, which presented a good performance in predicting overall survival. Our study provided novel insight into the potential association of tumor microenvironment with HCC prognosis and molecular targeted therapy of HCC in the future.
- Citation: Pan L, Fang J, Chen MY, Zhai ST, Zhang B, Jiang ZY, Juengpanich S, Wang YF, Cai XJ. Promising key genes associated with tumor microenvironments and prognosis of hepatocellular carcinoma. World J Gastroenterol 2020; 26(8): 789-803
- URL: https://www.wjgnet.com/1007-9327/full/v26/i8/789.htm
- DOI: https://dx.doi.org/10.3748/wjg.v26.i8.789
Core tip: We performed an integrated bioinformatics analysis to assess the influence of tumor microenvironment on the prognosis of patients with hepatocellular carcinoma (HCC). The results found that HCC patients with a high immune/stromal infiltration level had better survival benefits than patients with a low infiltration level. Ten microenvironment-related key genes were screened and used to construct a prognostic gene signature, which presented a good performance in predicting overall survival. Our study provided novel insight into the potential association of tumor microenvironment with HCC prognosis and molecular targeted therapy of HCC in the future.
Hepatocellular carcinoma (HCC) is a common malignant tumor and the fifth leading cause of cancer-related death in the United States[1]. Due to frequent recurrences and metastases, existing treatments show unsatisfactory efficacy. The pathogenesis of HCC is extremely intricate, involving various biological processes, such as signal transduction (e.g., Ras signaling pathways and WNT/β-catenin pathway) and cell cycle regulation, which displays the interaction among multiple genes in a complex signal network[2,3]. Recently, numerous studies have demonstrated that tumor microenvironments (TMEs) play an important role in cancer development, including HCC, hence affecting the clinical outcomes[4-6]. TMEs consist of immune cells, stromal cells, normal endothelial cells, and vascular cells. Stromal and immune cells are two major components of nontumor cells in the TMEs and are considered to be valuable for the prognostic evaluation of tumors[7,8].
Gene expression profiling has been widely used to identify the potential biomarkers and subsequently construct prognosis models, and it has enriched our knowledge of the underlying molecular mechanisms of tumorigenesis[9-11]. Yoshihara et al[12] designed the ESTIMATE method using gene expression data to calculate the stromal and immune scores, which can predict infiltration levels of two major nontumor cells in tumor issues. The ESTIMATE algorithm has been applied to breast cancer and prostate cancer, showing its feasibility and effectiveness[13,14].
Here, taking advantage of the HCC cohort in The Cancer Genome Atlas (TCGA) database, we used the ESTIMATE method to obtain a list of genes associated with TMEs that predict a poor prognosis in HCC patients. Importantly, such correlation was validated in four different HCC cohorts available from the integrative molecular database of HCC (HCCDB). Key genes were screened by utilizing the LASSO algorithm, and a multigene-based classifier was constructed to further evaluate the relationship between these key genes and the prognosis of HCC.
The gene expression profile (level 3 data) for HCC patients was acquired from the TCGA data portal (https://xenabrowser.net/datapages/), in which the RNA expression in HCC was analyzed using Illumina HiSeq 2000 RNA Sequencing (October 13, 2017). Clinical data, such as gender, age, grade, tumor stage, survival, and outcome, were also downloaded from TCGA data portal. The ESTIMATE algorithm was used to calculate immune scores and stromal scores, and the score data were obtained from the official data portal (https://bioinformatics.mdanderson.org/ estimate/)[12]. The abundances of six types of immune cells (B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells) in the TCGA were acquired via the TIMER data portal, a web server for comprehensive analysis of tumor-infiltrating immune cells (https:// cistrome.shinyapps.io/timer/)[15,16]. For validation, four HCC datasets were obtained from the HCCDB database (http://lifeome.net/database/hccdb/ home.html), including three Gene Expression Omnibus datasets (HCCDB6, HCCDB7, and HCCDB17) and one Liver Cancer-RIKEN, Japan Project from International Cancer Genome Consortium (HCCDB18)[17].
Data analysis was conducted using the “limma” package in R[18]. |log2FC| ≥ 1 and FDR < 0.05 were considered statistically significant to screen for differentially expressed genes (DEGs).
Gene annotation and functional analysis of DEGs were performed by utilizing the “clusterProfiler” in R and Metascape website (a gene annotation & analysis resource) (http://metascape.org)[19,20]. The STRING database was applied to construct potential protein-protein interactions among the DEGs[21]. PPIs with a confidence score ≥ 0.4 were downloaded and reconstructed via Cytoscape[22]. Small networks with less than ten nodes were removed. The connectivity degree of each node in the network was calculated. Moreover, molecular complex detection (MCODE) was performed to detect hub clusters in the PPI network. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were also conducted for significant modules.
Kaplan-Meier curves were used to estimate the survival differences for patients between the high and low score/risk groups. Univariate Cox proportional hazards regression analyses were conducted to determine the survival-related DEGs. LASSO regression was used to further screen the most powerful prognostic genes. A λ value of 0.066 with log (λ) = -2.72 was selected by 5-fold cross-validation via 1-SE criteria (Supplementary Figure 1). A prognostic gene signature was constructed according to a linear combination of the relative expression values (Expi) and LASSO coefficients (Li) using the following formula: Risk score = L1 × Exp1 + L2 × Exp2 +…+ Ln × Expn. To classify these HCC patients into the high- or low-risk group, the best cutoff value was calculated when the specificity and sensitivity in the receiver operating characteristic (ROC) curve achieved their maximum for predicting 3-year survival using TCGA data. To further evaluate if the prognostic classifier is an independent indicator in patients with HCC, the effect of each clinicopathologic factor on OS was analyzed by univariate Cox regression. Factors whose P value < 0.05 in the univariate analysis were selected for multivariate analysis. Additionally, to test the performance of the prognostic classifier in predicting HCC clinical outcomes, we calculated the area under the curve (AUC) to measure the predictive ability and compared gene signatures with other clinicopathologic features, including TNM stages. All statistical analyses were carried out with R (version 3.6.0) and GraphPad Prism 7 (GraphPad Software Inc., United States).
Gene expression profiles and clinical data of 373 HCC patients with their initial pathologic diagnosis made between 1995 and 2013 were downloaded from the TCGA database. Twenty-nine patients with an overall survival (OS) less than 30 d were excluded. Among the remaining patients, 109 (31.7%) were female, and 235 (68.3%) were male. Based on the ESTIMATE method, immune scores ranged from -1209.16 to 2934.36, and stromal scores were distributed between -1741.56 and 1195.07. Of note, 258 patients with scores higher than the 25th percentile were defined as a “high score group”, while 86 patients with scores lower than the 25th percentile were defined as a “low score group”. The detailed clinical information of HCC patients is shown in Supplementary Table 1. Kaplan-Meier survival curves showed that the mean recurrence-free survival (RFS) and OS in the high-immune score group were better than those in the low-immune score group (RFS: 1032 d vs 478 d, P = 0.0022; OS: 2116 d vs 848 d, P = 0. 0272; Figure 1A and B). Patients with high-stromal scores also had consistently better survival benefit in comparison to patients with low-stromal scores (RFS: 990 d vs 384 d, P = 0.0007; OS: 2116 d vs 1005 d, P = 0.0213; Figure 1C and D). Furthermore, detailed information about the infiltration levels of B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells in the high/low immune score groups is shown in Figure 1E-J (P < 0.0001).
| Times of gene verification by HCC cohorts from HCCDB | Down-regulated gene in low score group | Up-regulated gene in low score group |
| 2 | MS4A1, KLRB1, ITK, GZMH, ADH1C, UGT2B15, GPR171, CD40LG, CD5, FBLN2, CCL21, ATP2A3, IL16, ELN | |
| 1 | SELL, CD5L, KLRK1, CXCR6, TRAF3IP3, CD79A, FLT3, GZMA, SPN, PRKCQ, SIDT1, RERGL, INMT, C11orf21, ZAP70, FLI1, DOK5, ITM2A, PHLDA3, APLNR, CFHR3, NLRC3, GIMAP1, FBLN5, DARC, SLFN12L, RASGRP2, SELP, ACAP1, C7, HTR2B, TNXB, SAMD3, PDZD4 | CTSL2, MFAP2, TMC5, BACE2 |
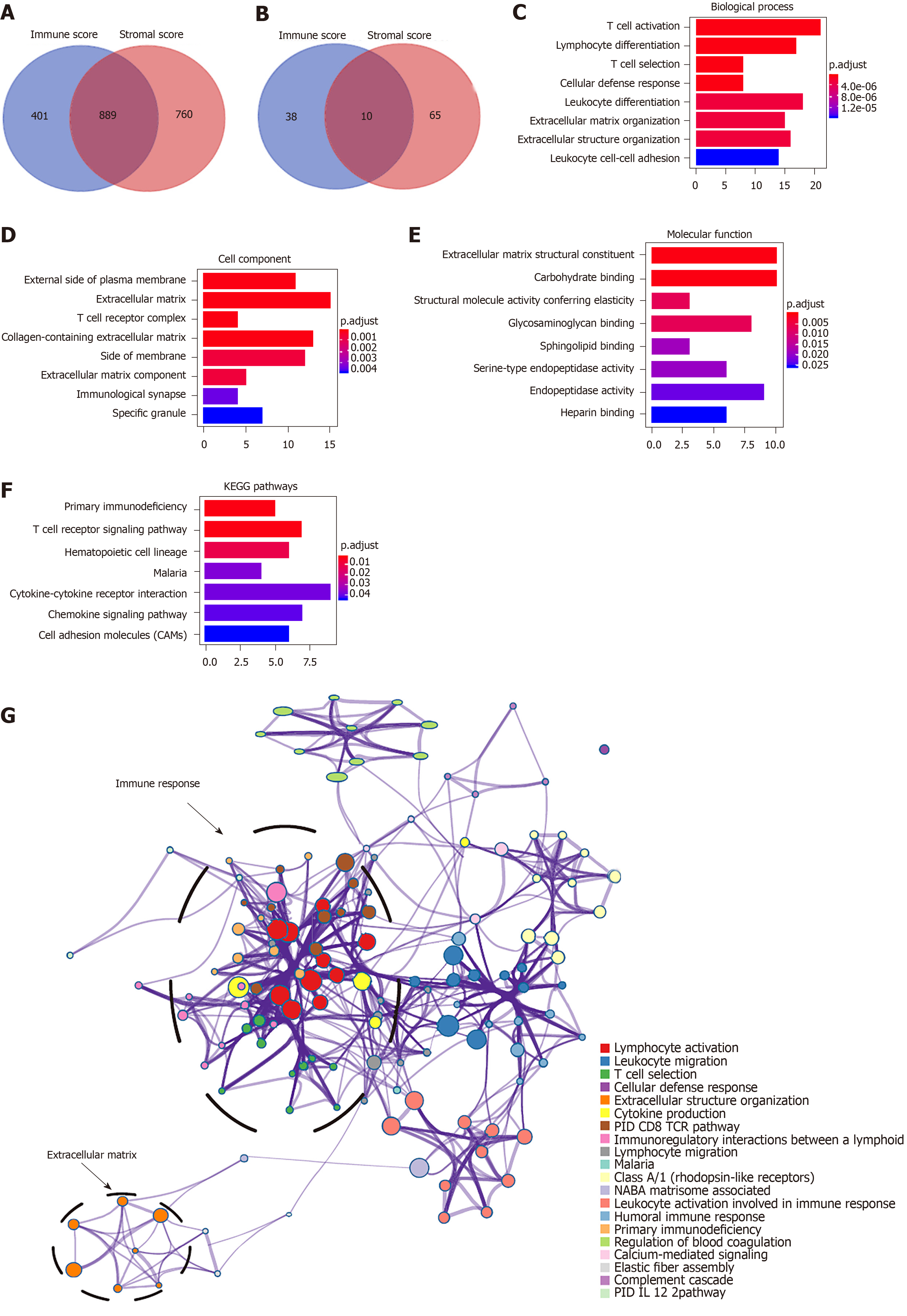
To explore the survival differences between the high and low score groups, we compared Illumina HiSeq 2000 RNA-seq data of all 373 HCC patients obtained in the TCGA database. For the high and low groups according to immune scores, 1290 genes were downregulated and 48 were upregulated in the low score group (fold change > 2; P < 0.05). Similarly, for comparison based on stromal scores, 1649 genes were downregulated and 75 were upregulated in the low score group (fold change > 2; P < 0.05). Furthermore, 889 genes were downregulated in both the low immune score group and low stromal score group, while 10 genes were commonly upregulated in the low immune/stromal score group (Figure 2A and B). To investigate the potential roles of individual DEGs in OS, we performed univariate Cox proportional hazards regression analysis using the TCGA database data. Among the 899 DEGs in the low immune/stromal score group, 147 were associated with a significantly poorer prognosis (P < 0.05, Supplementary Table 2). Therefore, we focused on these DEGs for subsequent analyses.
| Gene | Univariate Cox regression analysis | LASSO coefficient | ||
| HR | 95%CI | P value | ||
| NLRC3 | 0.72 | 0.60-0.87 | 0.0008 | -0.0780 |
| CFHR3 | 0.89 | 0.84-0.94 | 0.0001 | -0.0375 |
| KLRB1 | 0.84 | 0.76-0.93 | 0.0008 | -0.0276 |
| CTSL2 | 1.13 | 1.05-1.21 | 0.0005 | 0.0173 |
| ADH1C | 0.91 | 0.87-0.96 | 0.0002 | -0.0149 |
| CD5L | 0.90 | 0.85-0.95 | 0.0004 | -0.0131 |
| TMC5 | 1.06 | 1.01-1.12 | 0.0255 | 0.0106 |
| DOK5 | 0.84 | 0.75-0.95 | 0.0041 | -0.0082 |
| UGT2B15 | 0.91 | 0.86-0.96 | 0.0007 | -0.0072 |
| RASGRP2 | 0.82 | 0.72-0.93 | 0.0016 | -0.0062 |
To summarize the potential functions of 147 DEGs, functional enrichment analyses were performed. GO enrichment analysis indicated the following top GO terms: T cell activation, lymphocyte differentiation, and T cell selection in biological process (Figure 2C); external side of plasma membrane, extracellular matrix, and T cell receptor complex in cell component (Figure 2D); and extracellular matrix structural constituent, carbohydrate binding, and structural molecule activity conferring elasticity in molecular function (Figure 2E). Pathways identified by KEGG analysis were related to the immune response (Figure 2F). Moreover, we performed a network of enriched terms to capture the relationships between the terms by the metascape data portal and found that terms related to the adaptive immune process were in a central position in the network (Figure 2G).
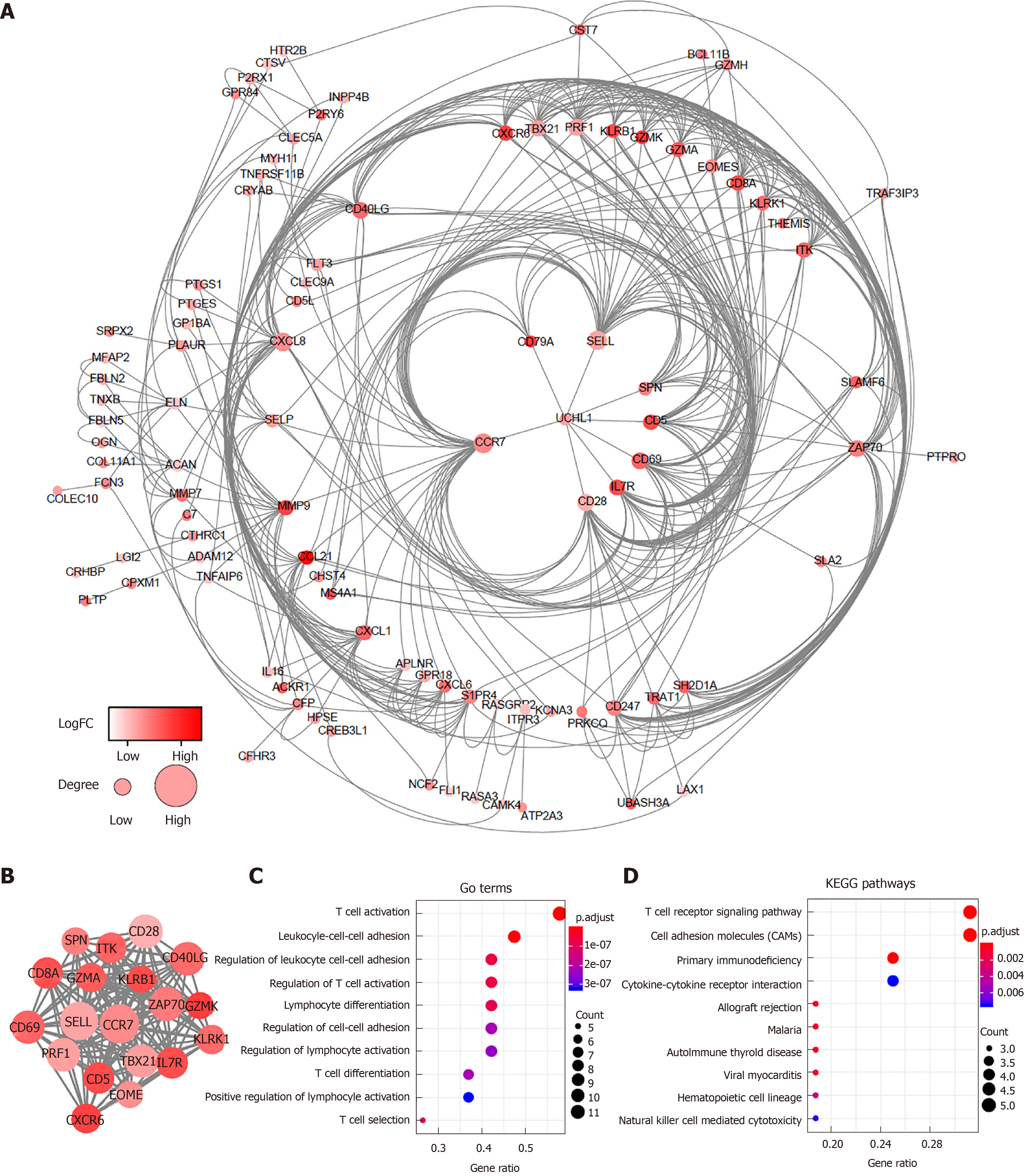
To better explore the interplay among the DEGs of prognostic significance, we generated PPI networks using the STRING data portal. The network consisted of six modules, which included 97 nodes and 420 interactions (Figure 3A). The top module (MCODE score 15.78; 19 nodes and 142 edges) was selected for further analysis (Figure 3B). Significant GO enrichment analysis showed that the module closely correlated with T cell activation, leukocyte cell-cell adhesion, and regulation of leukocyte cell-cell adhesion (Figure 3C). With respect to the KEGG pathway enrichment analysis, DEGs in the module were mainly located in the T cell receptor signaling pathway, cell adhesion molecules (CAMs), and primary immunodeficiency (Figure 3D).
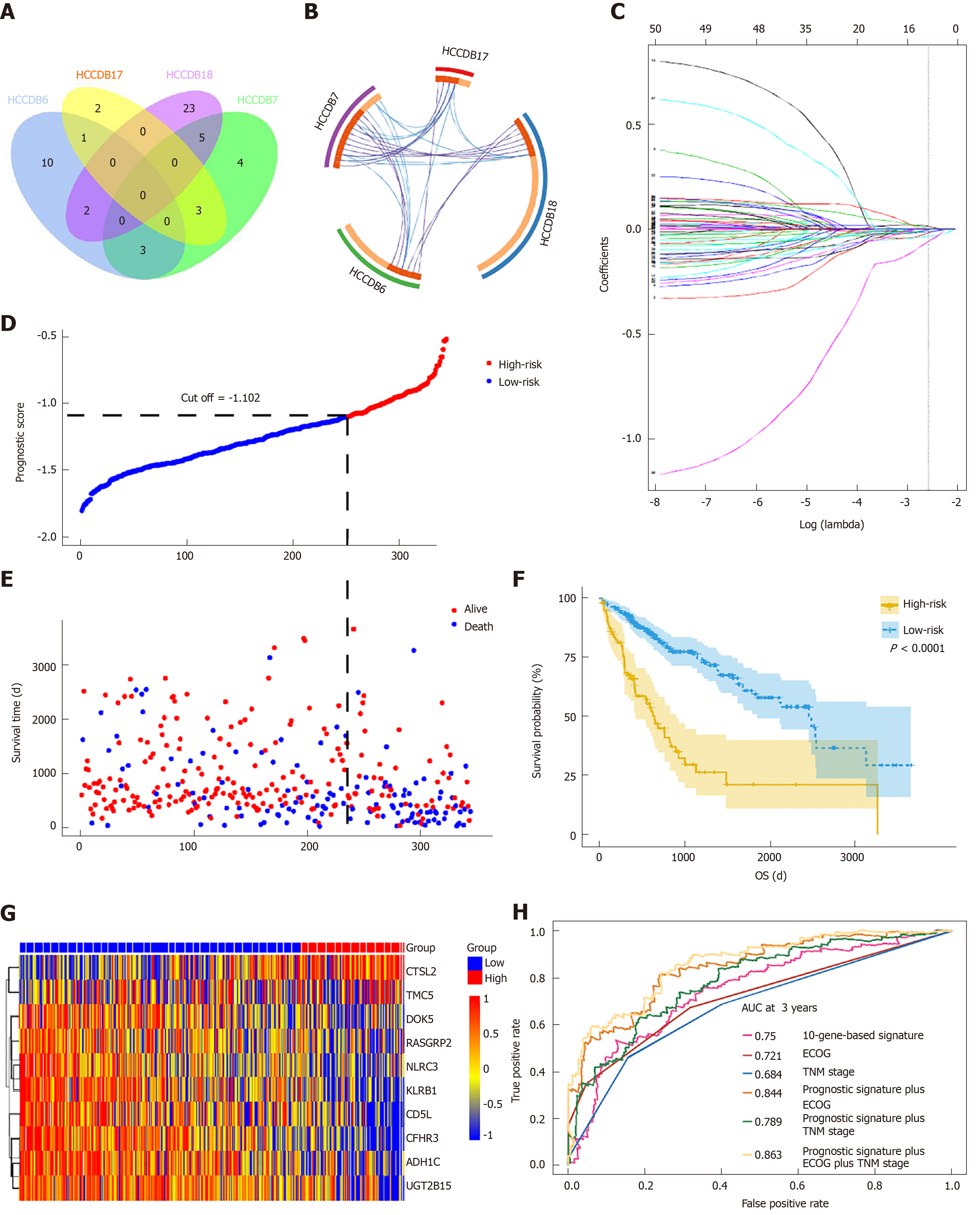
To determine if the genes identified from TCGA-HCC cohort are also of prognostic significance to other HCC-related cohorts, we downloaded the gene expression profile and clinical data of four HCC cohorts from the HCCDB database. A total of 52 genes were validated to be associated with a poor prognosis. Of these genes, 14 were verified by two HCCDB cohorts, and 38were verified by one HCCDB cohort (Figure 4A and B, Table 1). In addition, 38 genes generated particular interest as they have not been reported to be associated with poor outcomes in HCC patients previously.
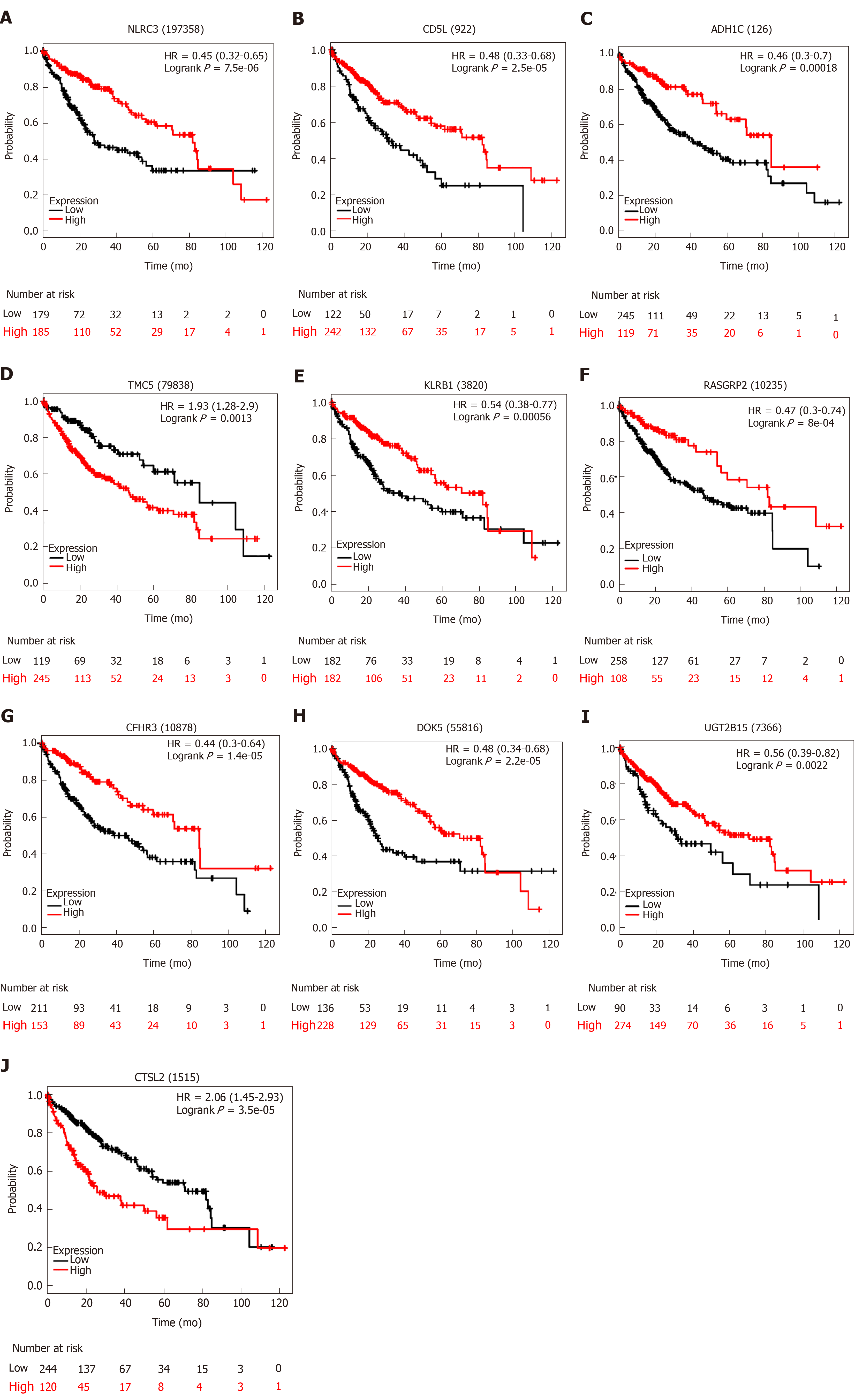
We utilized LASSO regression to screen ten key genes from the set of 52 genes that are most relevant to OS (Figure 4C, Table 2). The Kaplan-Meier survival curves of ten key genes are shown in Figure 5 and they presented great prediction performance of OS using the TCGA-HCC cohort. The risk score of patients was calculated according to the expression levels of the ten key genes and LASSO coefficients. A total of 251 patients with prognostic scores less than the cutoff value (-1.102) were categorized into the “low-risk group”, while the remaining 93 patients were categorized into the “high-risk group” (Figure 4D and E). An extremely significant difference in OS was detected between the two groups (P < 0.0001, Figure 4F). Figure 4G shows that STSL2 and TMC5 were highly expressed in the high-risk group, whereas the remaining eight genes (DOK5, RASGRP2, NLRC3, KLRB1, CD5L, CFHR3, ADH1C, and UGT2B15) were highly expressly in the low-risk group. To determine if the gene signature is an independent risk factor in patients with HCC, univariate and multivariate Cox regression analyses were conducted. After the multivariable adjustment, the ten-gene-based signature remained a significantly independent factor (P = 0.001, Table 3). In addition, the prognostic gene classifier showed a good performance in survival prediction, which was better than TNM staging (AUC for 3-year survival: 0.75 vs 0.684, Figure 4H). More importantly, the ten-gene-based signature combined with clinical features, such as Eastern Cooperative Oncology Group performance status and TNM stage, achieved a greater AUC, which was significantly better than the classifier alone. These results suggested that combining the ten-gene-based classifier with clinical features may further improve the ability to predict OS.
| Variable | Univariate analysis | Multivariate analysis | ||||
| HR | 95%CI | P value | HR | 95%CI | P value | |
| Risk score | 11.37 | 5.49-23.56 | < 0.001 | 8.99 | 2.53-32.01 | 0.001 |
| TNM stage | 1.79 | 1.45-2.22 | < 0.001 | 1.24 | 0.89-1.75 | 0.209 |
| Age | 1.01 | 0.99-1.02 | 0.254 | |||
| Gender | 0.79 | 0.55-1.13 | 0.192 | |||
| Race | 1.13 | 0.94-1.37 | 0.196 | |||
| Child-Pugh score | 1.69 | 0.93-3.07 | 0.084 | |||
| ECGO performance status | 2.41 | 1.91-3.04 | < 0.001 | 1.74 | 1.16-2.63 | 0.008 |
| AFP | 1 | 1-1 | 0.443 | |||
| Fibrosis ishak score | 0.91 | 0.78-1.07 | 0.253 | |||
| Histologic grade | 1.1 | 0.86-1.4 | 0.445 | |||
| Vascular invasion | 1.47 | 1.06-2.06 | 0.022 | 1.16 | 0.75-1.79 | 0.511 |
| Hepatitis virus infection | 0.45 | 0.3-0.68 | < 0.001 | 0.81 | 0.45-1.46 | 0.477 |
| Alcohol consumption | 1.1 | 0.75-1.62 | 0.616 | |||
In this study, low infiltration levels of immune and stromal cells in HCC microenvironments were associated with poor outcomes, and we confirmed the TME-related genes responsible for OS in the TCGA-HCC cohort. By investigating the gene expression profiles in 373 cases with low vs high immune/stromal scores, 147 prognosis-related DEGs were identified to be involved in immune response and extracellular matrix. More importantly, 52 genes were validated in additional four HCC cohorts from HCCDB, an integrative HCC database containing Gene Expression Omnibus data and International Cancer Genome Consortium data. Ten key genes were screened by the LASSO algorithm and used to construct a prognostic classifier, which showed good performance for predicting OS, indicating a close correlation between these key genes and the prognosis of HCC.
Based on the survival difference between the high and low score groups, 899 DEGs were obtained from the comparison. Among these DEGs, 147 were identified to be related to poor outcomes, many of which are involved in TMEs (Figure 2). These findings are consistent with previous studies showing that TMEs have a critical influence on HCC[4,23,24].
Second, we also generated a PPI network to investigate the correlation among the 147 genes. Ubiquitin carboxyl-terminal hydrolase L1 (UCHL1), which was at the center of the network, has been reported to be a tumor suppressor and play a vital role in the tumorigenesis of HCC[25]. Densely connected regions in the network determined by MCODE module analysis are also involved in immune responses and extracellular matrices, which is consistent with the overall enrichment results of all 147 genes. The nodes with a highly connectivity degree in the module containing CXCR6, CCR7, and CD69 have been demonstrated to promote the progression of HCC and be involved in dysfunctions of immune cells[26-28].
By cross validation with four independent HCC cohorts from the HCCDB database, 52 TME-related genes were identified, showing a significant correlation between gene expression and OS. Of the 52 genes confirmed, 12 (e.g., CD5, CCL21, CXCR6, GZMA, and SELP) have been reported to be involved in hepatocarcinogenesis or significant in predicting prognosis, indicating that our bioinformatical analysis using TCGA and HCCDB cohorts has reliable prognostic values. The remaining 38 genes have not been previously studied for their effects on hepatocarcinogenesis, suggesting that they may serve as novel therapeutic targets or potential biomarkers for HCC.
Finally, we used LASSO regression analysis to identify ten key genes and subsequently constructed a gene signature showing great capacity for predicting OS. According to our study, CTSL2 and TMC5 were regarded as tumor promoters, while DOK5, CFHR3, ADH1C, UGT2B15, CD5L, RASGRP2, KLRB1, and NLRC3 were regarded as tumor suppressors. NLRC3, a member of the nucleotide-binding domain and leucine-rich repeat (NLR) family protein, plays an important role in immunity and inflammation[29,30]. Ma et al[31] reported that high expression levels of NLRC3 correlate with a favorable clinical prognosis in a Chinese HCC population and that downregulation of NLRC3 expression inhibits cell apoptosis and contributes to cell proliferation in vitro. Maehara et al[32] investigated CD5L, which is the apoptosis inhibitor of macrophages, and they found that CD5L prevents hepatocellular carcinoma through complement activation. Two other studies have also shown that CD5L favors overall HCC survival, which agreed with our study[33,34]. Therefore, the prognostic value of the additional eight key genes reported in this study may provide a valuable direction for future research.
The interactions of HCC and the TME significantly affect tumor evolution, hence influencing tumor escape, tumor recurrence, drug resistance, and patient clinical outcomes[4,35-37]. Remarkable progress has been made in the research of the interplay between HCC and TME, and most of these experiments have been performed in cell lines, animal models, or small cohorts based on patient samples. However, the complexity of HCC and TME requires additional comprehensive analysis. Fortunately, the development of tumor databases, including TCGA and HCCDB, has provided a solid foundation for high-throughput screening of large-scale HCC cohorts. Our results provide novel data revealing the complex interplay of tumor and TME in HCC.
There are several limitations in our study. First, due to the current study being implemented based on bioinformatics analysis, biological experiments are urgently needed for validation. Second, as the data used were from public databases, their quality was not effectively assessed. Third, the clinical characteristic of patients were not taken into account in survival analysis since identification of the prognosis-related genes from multiple datasets was the primary focus.
In conclusion, this study extracted a series of genes associated with TME and prognosis from the TCGA database using the ESTIMATE method. These genes were validated in four independent HCC cohorts. After screening by LASSO regression, ten key genes were used to construct a gene signature, showing good performance in predicting OS. These findings contribute to outlining the prognosis and molecular targeted therapy of HCC. Further investigations of these key genes may bring a novel insight into the potential association of TME with HCC prognosis in a more comprehensive pattern.
Tumor microenvironments (TMEs) play an important role in cancer development, including hepatocellular carcinoma (HCC), hence affecting clinical outcomes. Stromal and immune cells are two major components of nontumor cells in TMEs and are considered to be valuable for the prognostic evaluation of tumors. Nonetheless, the infiltration level of stromal/immune cells and specific roles of TME-related genes in HCC have not yet been clarified.
More biomarkers are required for the diagnosis and treatment of HCC.
The present study investigated potential key genes associated with tumor microenvironments and the prognosis of HCC.
The ESTIMATE method was used to predict the infiltration levels of stromal and immune cells in HCC. Differentially expressed genes (DEGs) between patients with high and low infiltration levels were screened using the TCGA database and linked to overall survival of HCC. DEGs were verified by four other independent HCC cohorts and further selected by LASSO Cox regression analysis.
HCC patients with high immune/stromal infiltration levels had better survival benefits than patients with low infiltration levels. A total of 147 DEGs were identified to be associated with overall survival. Moreover, 52 survival-related DEGs were validated, and ten key genes were selected by LASSO algorithm, which were further used to construct a prognostic classifier, showing a good performance in predicting overall survival.
Our study screened a series of TME-related genes and provided a novel insight into the potential association of TME with HCC prognosis and molecular targeted therapy of HCC in the future.
HCC and TMEs are an integral whole. More in-depth research should be conducted to reveal the important role of components of TMEs in HCC to develop novel anti-cancer treatments via targeting TME-related cells and the extracellular matrix.
We thank Yi-Dong Pan (Guy's, King's & St Thomas's (GKT) School of Medical Education, King's College London, London, United Kingdom) and AJE (http://www.aje.com) for their linguistic assistance during the preparation of this manuscript.
| 1. | Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11530] [Cited by in RCA: 13111] [Article Influence: 1638.9] [Reference Citation Analysis (3)] |
| 2. | Qu C, He, Lu X, Dong L, Zhu Y, Zhao Q, Jiang X, Chang P, Jiang X, Wang L, Zhang Y, Bi L, He J, Peng Y, Su J, Zhang H, Huang H, Li Y, Zhou S, Qu Y, Zhao Y, Zhang Z. Salt-inducible Kinase (SIK1) regulates HCC progression and WNT/β-catenin activation. J Hepatol. 2016;64:1076-1089. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 73] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 3. | Levrero M, Zucman-Rossi J. Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol. 2016;64:S84-S101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 814] [Cited by in RCA: 775] [Article Influence: 77.5] [Reference Citation Analysis (4)] |
| 4. | Giannelli G, Rani B, Dituri F, Cao Y, Palasciano G. Moving towards personalised therapy in patients with hepatocellular carcinoma: the role of the microenvironment. Gut. 2014;63:1668-1676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 87] [Article Influence: 7.3] [Reference Citation Analysis (3)] |
| 5. | Muppa P, Parrilha Terra SBS, Sharma A, Mansfield AS, Aubry MC, Bhinge K, Asiedu MK, de Andrade M, Janaki N, Murphy SJ, Nasir A, Van Keulen V, Vasmatzis G, Wigle DA, Yang P, Yi ES, Peikert T, Kosari F. Immune Cell Infiltration May Be a Key Determinant of Long-Term Survival in Small Cell Lung Cancer. J Thorac Oncol. 2019;14:1286-1295. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 88] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 6. | Wu T, Dai Y. Tumor microenvironment and therapeutic response. Cancer Lett. 2017;387:61-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1494] [Cited by in RCA: 1338] [Article Influence: 148.7] [Reference Citation Analysis (0)] |
| 7. | Fridman WH, Pagès F, Sautès-Fridman C, Galon J. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12:298-306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4103] [Cited by in RCA: 3750] [Article Influence: 267.9] [Reference Citation Analysis (8)] |
| 8. | Mlecnik B, Tosolini M, Kirilovsky A, Berger A, Bindea G, Meatchi T, Bruneval P, Trajanoski Z, Fridman WH, Pagès F, Galon J. Histopathologic-based prognostic factors of colorectal cancers are associated with the state of the local immune reaction. J Clin Oncol. 2011;29:610-618. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 858] [Cited by in RCA: 792] [Article Influence: 52.8] [Reference Citation Analysis (3)] |
| 9. | Villanueva A, Hoshida Y, Battiston C, Tovar V, Sia D, Alsinet C, Cornella H, Liberzon A, Kobayashi M, Kumada H, Thung SN, Bruix J, Newell P, April C, Fan JB, Roayaie S, Mazzaferro V, Schwartz ME, Llovet JM. Combining clinical, pathology, and gene expression data to predict recurrence of hepatocellular carcinoma. Gastroenterology. 2011;140:1501-12.e2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 345] [Cited by in RCA: 344] [Article Influence: 22.9] [Reference Citation Analysis (5)] |
| 10. | Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O'Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN; Cancer Genome Atlas Research Network. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 2010;17:98-110. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6344] [Cited by in RCA: 5777] [Article Influence: 361.1] [Reference Citation Analysis (5)] |
| 11. | Shedden K, Taylor JM, Enkemann SA, Tsao MS, Yeatman TJ, Gerald WL, Eschrich S, Jurisica I, Giordano TJ, Misek DE, Chang AC, Zhu CQ, Strumpf D, Hanash S, Shepherd FA, Ding K, Seymour L, Naoki K, Pennell N, Weir B, Verhaak R, Ladd-Acosta C, Golub T, Gruidl M, Sharma A, Szoke J, Zakowski M, Rusch V, Kris M, Viale A, Motoi N, Travis W, Conley B, Seshan VE, Meyerson M, Kuick R, Dobbin KK, Lively T, Jacobson JW, Beer DG; Director's Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med. 2008;14:822-827. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 909] [Cited by in RCA: 886] [Article Influence: 49.2] [Reference Citation Analysis (0)] |
| 12. | Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7649] [Cited by in RCA: 7125] [Article Influence: 548.1] [Reference Citation Analysis (5)] |
| 13. | Alonso MH, Aussó S, Lopez-Doriga A, Cordero D, Guinó E, Solé X, Barenys M, de Oca J, Capella G, Salazar R, Sanz-Pamplona R, Moreno V. Comprehensive analysis of copy number aberrations in microsatellite stable colon cancer in view of stromal component. Br J Cancer. 2017;117:421-431. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 81] [Cited by in RCA: 104] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 14. | Shah N, Wang P, Wongvipat J, Karthaus WR, Abida W, Armenia J, Rockowitz S, Drier Y, Bernstein BE, Long HW, Freedman ML, Arora VK, Zheng D, Sawyers CL. Regulation of the glucocorticoid receptor via a BET-dependent enhancer drives antiandrogen resistance in prostate cancer. Elife. 2017;6. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 126] [Cited by in RCA: 145] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 15. | Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017;77:e108-e110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4366] [Cited by in RCA: 4366] [Article Influence: 485.1] [Reference Citation Analysis (3)] |
| 16. | Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, Jiang P, Shen H, Aster JC, Rodig S, Signoretti S, Liu JS, Liu XS. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 2016;17:174. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1888] [Cited by in RCA: 1764] [Article Influence: 176.4] [Reference Citation Analysis (4)] |
| 17. | Lian Q, Wang S, Zhang G, Wang D, Luo G, Tang J, Chen L, Gu J. HCCDB: A Database of Hepatocellular Carcinoma Expression Atlas. Genomics Proteomics Bioinformatics. 2018;16:269-275. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 118] [Cited by in RCA: 236] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 18. | Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35052] [Cited by in RCA: 29373] [Article Influence: 2670.3] [Reference Citation Analysis (7)] |
| 19. | Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, Benner C, Chanda SK. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11806] [Cited by in RCA: 10245] [Article Influence: 1463.6] [Reference Citation Analysis (5)] |
| 20. | Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284-287. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30559] [Cited by in RCA: 26105] [Article Influence: 1864.6] [Reference Citation Analysis (5)] |
| 21. | Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45:D362-D368. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5872] [Cited by in RCA: 5195] [Article Influence: 577.2] [Reference Citation Analysis (4)] |
| 22. | Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498-2504. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43766] [Cited by in RCA: 36496] [Article Influence: 1586.8] [Reference Citation Analysis (4)] |
| 23. | Giannelli G, Fransvea E, Bergamini C, Marinosci F, Antonaci S. Laminin-5 chains are expressed differentially in metastatic and nonmetastatic hepatocellular carcinoma. Clin Cancer Res. 2003;9:3684-3691. [PubMed] |
| 24. | Govaere O, Komuta M, Berkers J, Spee B, Janssen C, de Luca F, Katoonizadeh A, Wouters J, van Kempen LC, Durnez A, Verslype C, De Kock J, Rogiers V, van Grunsven LA, Topal B, Pirenne J, Vankelecom H, Nevens F, van den Oord J, Pinzani M, Roskams T. Keratin 19: a key role player in the invasion of human hepatocellular carcinomas. Gut. 2014;63:674-685. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 239] [Cited by in RCA: 232] [Article Influence: 19.3] [Reference Citation Analysis (4)] |
| 25. | Yu J, Tao Q, Cheung KF, Jin H, Poon FF, Wang X, Li H, Cheng YY, Röcken C, Ebert MP, Chan AT, Sung JJ. Epigenetic identification of ubiquitin carboxyl-terminal hydrolase L1 as a functional tumor suppressor and biomarker for hepatocellular carcinoma and other digestive tumors. Hepatology. 2008;48:508-518. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 125] [Cited by in RCA: 136] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 26. | Mossanen JC, Kohlhepp M, Wehr A, Krenkel O, Liepelt A, Roeth AA, Möckel D, Heymann F, Lammers T, Gassler N, Hermann J, Jankowski J, Neumann UP, Luedde T, Trautwein C, Tacke F. CXCR6 Inhibits Hepatocarcinogenesis by Promoting Natural Killer T- and CD4+ T-Cell-Dependent Control of Senescence. Gastroenterology. 2019;156:1877-1889.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 96] [Article Influence: 13.7] [Reference Citation Analysis (3)] |
| 27. | Duan M, Goswami S, Shi JY, Wu LJ, Wang XY, Ma JQ, Zhang Z, Shi Y, Ma LJ, Zhang S, Xi RB, Cao Y, Zhou J, Fan J, Zhang XM, Gao Q. Activated and Exhausted MAIT Cells Foster Disease Progression and Indicate Poor Outcome in Hepatocellular Carcinoma. Clin Cancer Res. 2019;25:3304-3316. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 136] [Article Influence: 19.4] [Reference Citation Analysis (3)] |
| 28. | Easom NJW, Stegmann KA, Swadling L, Pallett LJ, Burton AR, Odera D, Schmidt N, Huang WC, Fusai G, Davidson B, Maini MK. IL-15 Overcomes Hepatocellular Carcinoma-Induced NK Cell Dysfunction. Front Immunol. 2018;9:1009. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 80] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 29. | Gültekin Y, Eren E, Özören N. Overexpressed NLRC3 acts as an anti-inflammatory cytosolic protein. J Innate Immun. 2015;7:25-36. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 47] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 30. | Lv Z, Wang Y, Liu YJ, Mao YF, Dong WW, Ding ZN, Meng GX, Jiang L, Zhu XY. NLRP3 Inflammasome Activation Contributes to Mechanical Stretch-Induced Endothelial-Mesenchymal Transition and Pulmonary Fibrosis. Crit Care Med. 2018;46:e49-e58. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 81] [Article Influence: 10.1] [Reference Citation Analysis (1)] |
| 31. | Ma YY, Zhang GH, Li J, Wang SB, Hu ZM, Zhang CW, Li E. The correlation of NLRC3 expression with the progression and prognosis of hepatocellular carcinoma. Hum Pathol. 2018;82:273-281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 32. | Maehara N, Arai S, Mori M, Iwamura Y, Kurokawa J, Kai T, Kusunoki S, Taniguchi K, Ikeda K, Ohara O, Yamamura KI, Miyazaki T. Circulating AIM prevents hepatocellular carcinoma through complement activation. Cell Rep. 2014;9:61-74. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 65] [Article Influence: 5.4] [Reference Citation Analysis (1)] |
| 33. | Zhang X, Kang C, Li N, Liu X, Zhang J, Gao F, Dai L. Identification of special key genes for alcohol-related hepatocellular carcinoma through bioinformatic analysis. PeerJ. 2019;7:e6375. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 24] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 34. | Brunner SM, Itzel T, Rubner C, Kesselring R, Griesshammer E, Evert M, Teufel A, Schlitt HJ, Fichtner-Feigl S. Tumor-infiltrating B cells producing antitumor active immunoglobulins in resected HCC prolong patient survival. Oncotarget. 2017;8:71002-71011. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 31] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 35. | Tian XP, Wang CY, Jin XH, Li M, Wang FW, Huang WJ, Yun JP, Xu RH, Cai QQ, Xie D. Acidic Microenvironment Up-Regulates Exosomal miR-21 and miR-10b in Early-Stage Hepatocellular Carcinoma to Promote Cancer Cell Proliferation and Metastasis. Theranostics. 2019;9:1965-1979. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 94] [Cited by in RCA: 200] [Article Influence: 28.6] [Reference Citation Analysis (4)] |
| 36. | Liu M, Zhou J, Liu X, Feng Y, Yang W, Wu F, Cheung OK, Sun H, Zeng X, Tang W, Mok MTS, Wong J, Yeung PC, Lai PBS, Chen Z, Jin H, Chen J, Chan SL, Chan AWH, To KF, Sung JJY, Chen M, Cheng AS. Targeting monocyte-intrinsic enhancer reprogramming improves immunotherapy efficacy in hepatocellular carcinoma. Gut. 2019;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 80] [Cited by in RCA: 143] [Article Influence: 23.8] [Reference Citation Analysis (0)] |
| 37. | Kim W, Khan SK, Liu Y, Xu R, Park O, He Y, Cha B, Gao B, Yang Y. Hepatic Hippo signaling inhibits protumoural microenvironment to suppress hepatocellular carcinoma. Gut. 2018;67:1692-1703. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 125] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
Open-Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See:
Manuscript source: Invited Manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Kang KJ, Mizuguchi T, Sung WW S-Editor: Gong ZM L-Editor: Wang TQ E-Editor: Ma YJ