Copyright: ©Author(s) 2026.
World J Gastroenterol. Apr 28, 2026; 32(16): 115969
Published online Apr 28, 2026. doi: 10.3748/wjg.v32.i16.115969
Published online Apr 28, 2026. doi: 10.3748/wjg.v32.i16.115969
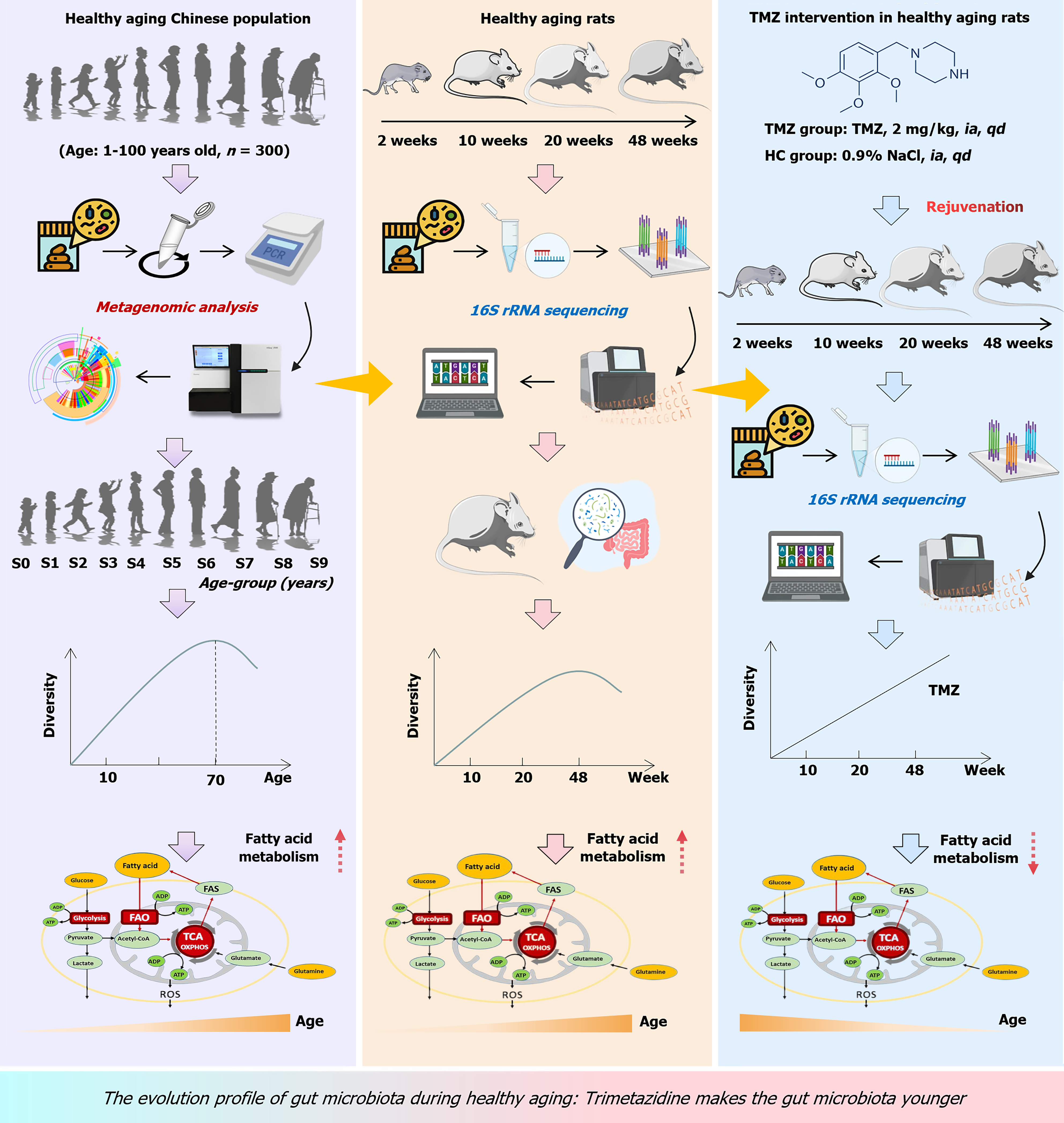
Figure 1 Design flowchart of this study.
S0-S9 indicate the healthy natural aging population group was categorized into ten subgroups based on age (n = 30 for each group). TMZ: Trimetazidine; FAO: Fatty acid oxidation; TCA: Tricarboxylic acid cycle; ADP: Adenosine diphosphate; ATP: Adenosine triphosphate; ROS: Reactive oxygen species; HC: Healthy control; NaCl: Sodium chloride.
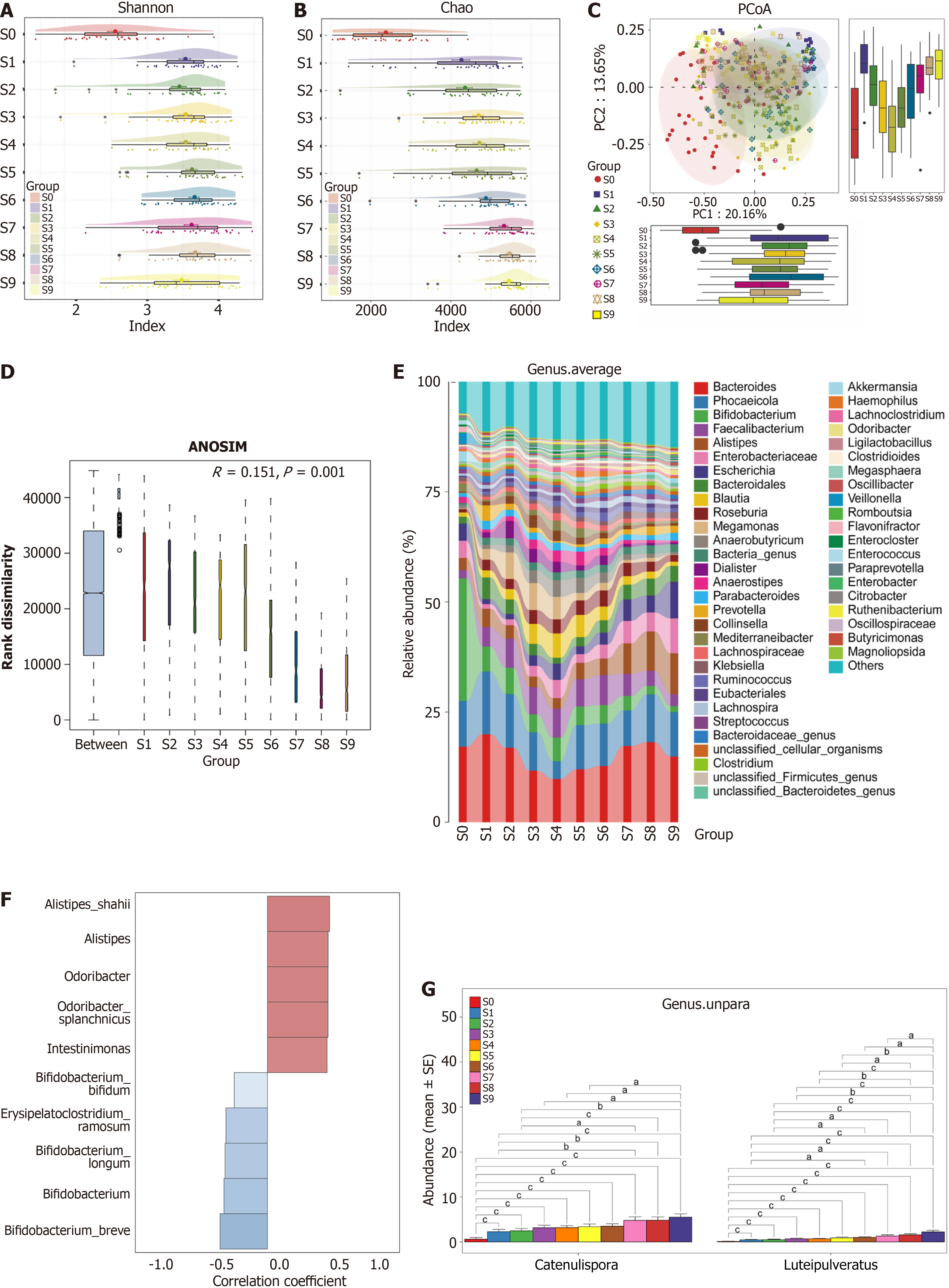
Figure 2 Characterization of age-associated gut microbiome alterations in the healthy natural aging population group.
A and B: The Shannon index (A) and Chao index (B) were used to determine the alpha diversity changes during the healthy aging process. The gut microbial diversity significantly increased from the S0 group (n = 30) to the S1 group (n = 30), and from S2 (n = 30) to S7, the diversity remained relatively stable but decreased from the S7 group (n = 30) (all P < 0.05); C: Principal coordinate analysis distribution revealed that the gut taxonomic composition significantly differed among the ten groups; D: As estimated by analysis of similarities analysis, the corresponding statistical significance of the beta diversity was measured separately; E: The average compositions and relative abundance of the gut microbial community in groups at the genus level; F: Correlation analysis of the abundance of the major species with age. Heatmap showing the abundance-fold change in bacteria with age. In red: Bacteria that are more abundant; in blue: Bacteria that are less abundant. Scale: Log2 (fold change) = -1 (blue) < 0 (white) < 1 (red); G: Changes in the abundances of Catenulispora and Luteipulveratus across the human lifespan. S0-S9 indicate the healthy natural aging population group was categorized into ten subgroups based on age (n = 30 for each group). aP < 0.05. bP < 0.01. cP < 0.001. PCoA: Principal coordinate analysis; PC: Principal component; ANOSIM: Analysis of similarities.
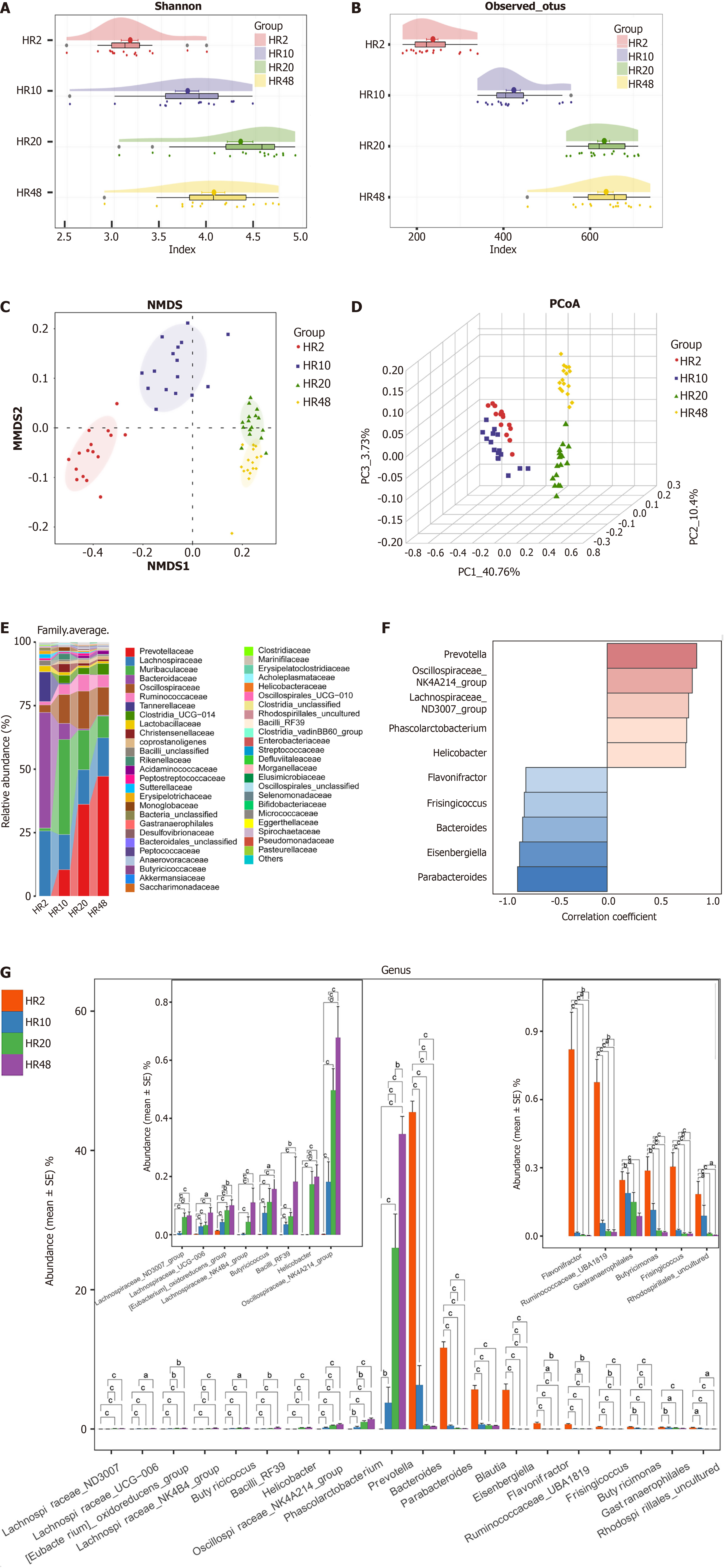
Figure 3 Gut microbiome profiles across different life stages in healthy rat cohort.
A and B: The Shannon index (A) and observed operational taxonomic units diversity indices (B) demonstrate the alpha diversity of the ten groups of the healthy rat cohort (HR) cohort; C and D: The nonmetric multidimensional scaling (C) and principal coordinate analysis (D) revealed differences in the gut microbiome among aged 2 weeks (n = 16), aged 10 weeks (n = 16), aged 20 weeks (n = 16) and aged 48 weeks (n = 16) groups; E: Average compositions and relative abundance of the bacterial community among groups at the genus level; F: Correlation analysis of the abundance of the major species with age. Heatmap showing the abundance-fold change in bacteria with age. In red: Bacteria that are more abundant; in blue: Bacteria that are less abundant. Scale: Log2 (fold change) = -1 (blue) < 0 (white) < 1 (red); G: Age-associated changes in differential gut microbial abundance in rats. aP < 0.05. bP < 0.01. cP < 0.001. HR: Healthy rat cohort; HR2: Aged 2 weeks; HR10: Aged 10 weeks; HR20: Aged 20 weeks; HR48: Aged 48 weeks; PCoA: Principal coordinate analysis; PC: Principal component; NMDS: Nonmetric multidimensional scaling.
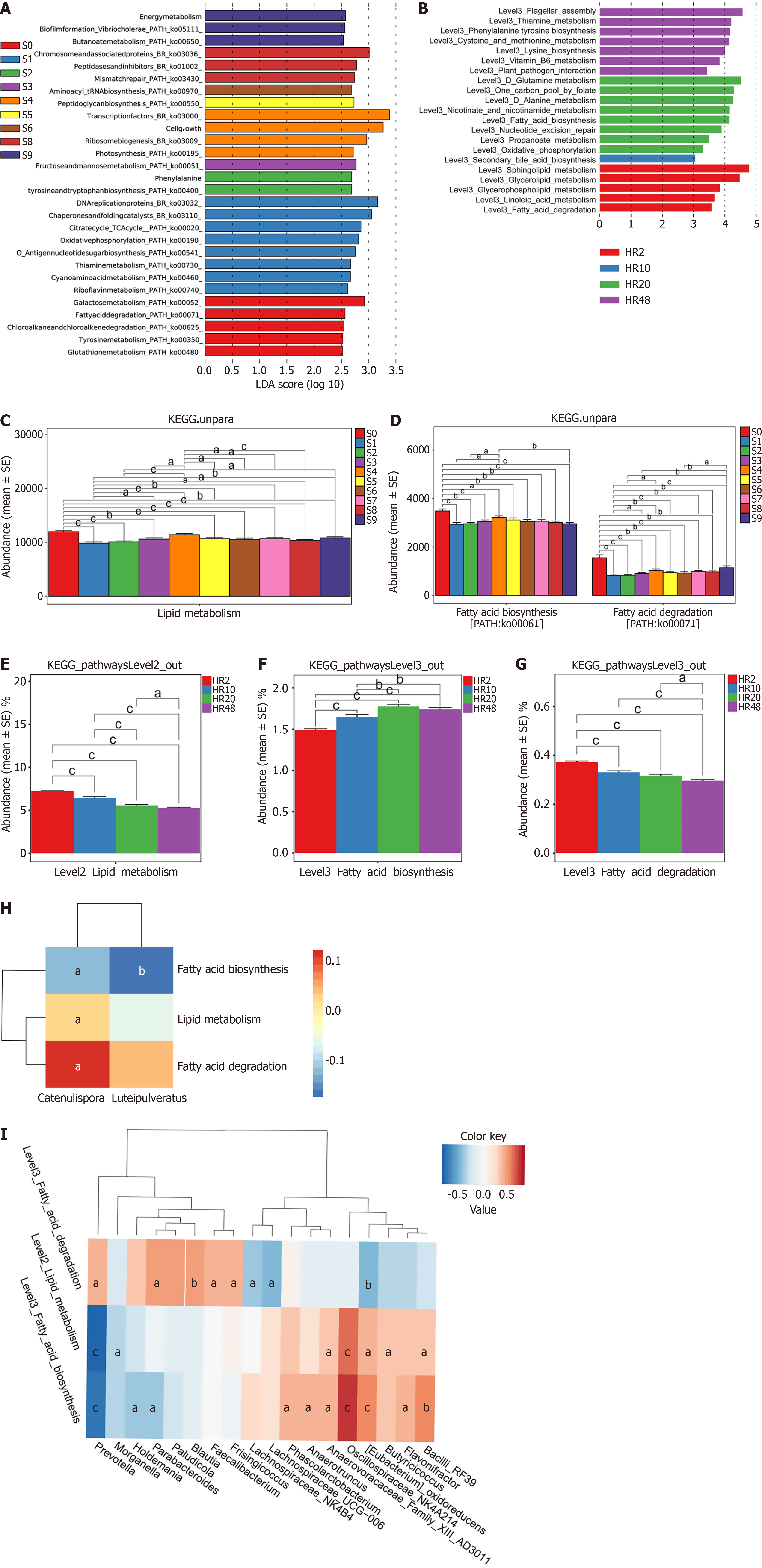
Figure 4 Lipid metabolism associated gut microbiota is significantly enriched in both healthy natural aging population and healthy rat cohorts.
A: Functional prediction of the gut microbiota based on Kyoto Encyclopedia of Genes and Genomes pathways in the healthy natural aging population (HP) group, highlighting key metabolic modules; B: The bar plot of microbial abundance shows a progressive enrichment of lipid metabolism-associated microbiota; C: Gut microbiota lipid metabolism pathway analysis of the HP group; D: The abundance of lipid metabolism-associated microbes in the HP group; E-G: Bar chart showing the abundance of microbes enriched in lipid metabolism, fatty acid biosynthesis and degradation metabolic pathways; H and I: Correlation analysis between the gut microbiota and lipid metabolism-associated pathways of the HP group (H) and the healthy rat cohort (P < 0.05, false discovery rate corrected) (I). aP < 0.05. bP < 0.01. cP < 0.001. S0-S9 indicate the healthy natural aging population group was categorized into ten subgroups based on age (n = 30 for each group). HR: Healthy rat cohort; HR2: Aged 2 weeks; HR10: Aged 10 weeks; HR20: Aged 20 weeks; HR48: Aged 48 weeks; LDA: Linear discriminant analysis; KEGG: Kyoto Encyclopedia of Genes and Genomes.
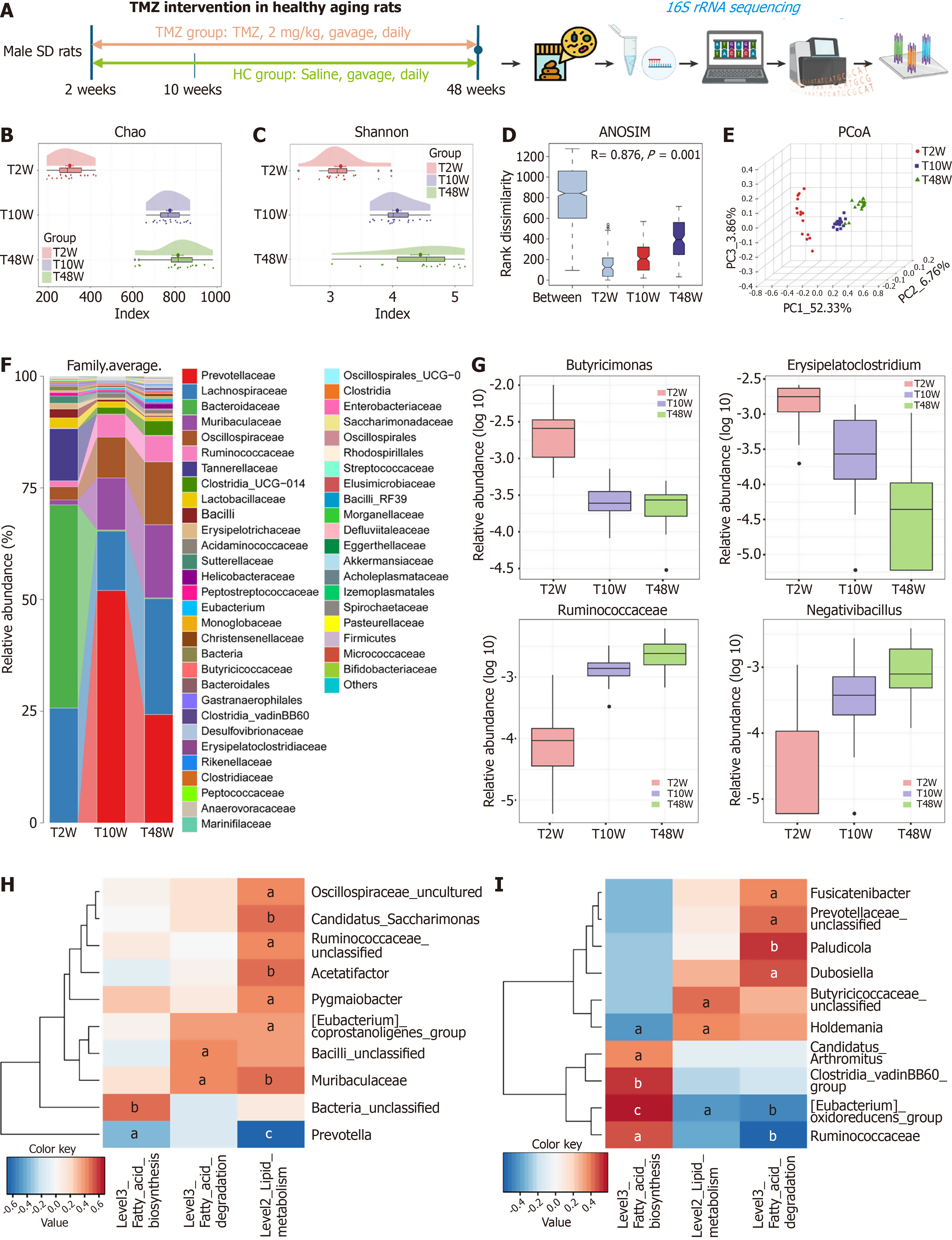
Figure 5 Trimetazidine intervention restored gut microbiota composition of healthy rats.
A: Schematic of the experimental intervention timeline for the animals; B and C: The Chao index (B) and Shannon index (C) indicated that the gut microbial community gradually increased from trimetazidine intervention group collected at 2, 10, and 48 weeks; D: Analysis of similarities; E: The principal coordinate analysis revealed the gut microbiome separated; F: Average compositions and relative abundance of the bacterial community among groups at the genus level; G: Relative abundances of Erysipelatoclostridium, Butyricimonas, Ruminococcaceae, and Negativibacillus; H and I: Correlation analysis between the gut microbiota and lipid metabolism-associated pathways at 10 weeks (H) and 48 weeks (I) of trimetazidine treatment (P < 0.05, false discovery rate corrected). aP < 0.05. bP < 0.01. cP < 0.001. TMZ: Trimetazidine; HC: Healthy control; T2W: Trimetazidine intervention group collected at 2 weeks; T10W: Trimetazidine intervention group collected at 10 weeks; T48W: Trimetazidine intervention group collected at 48 weeks; PCoA: Principal coordinate analysis; PC: Principal component; ANOSIM: Analysis of similarities.
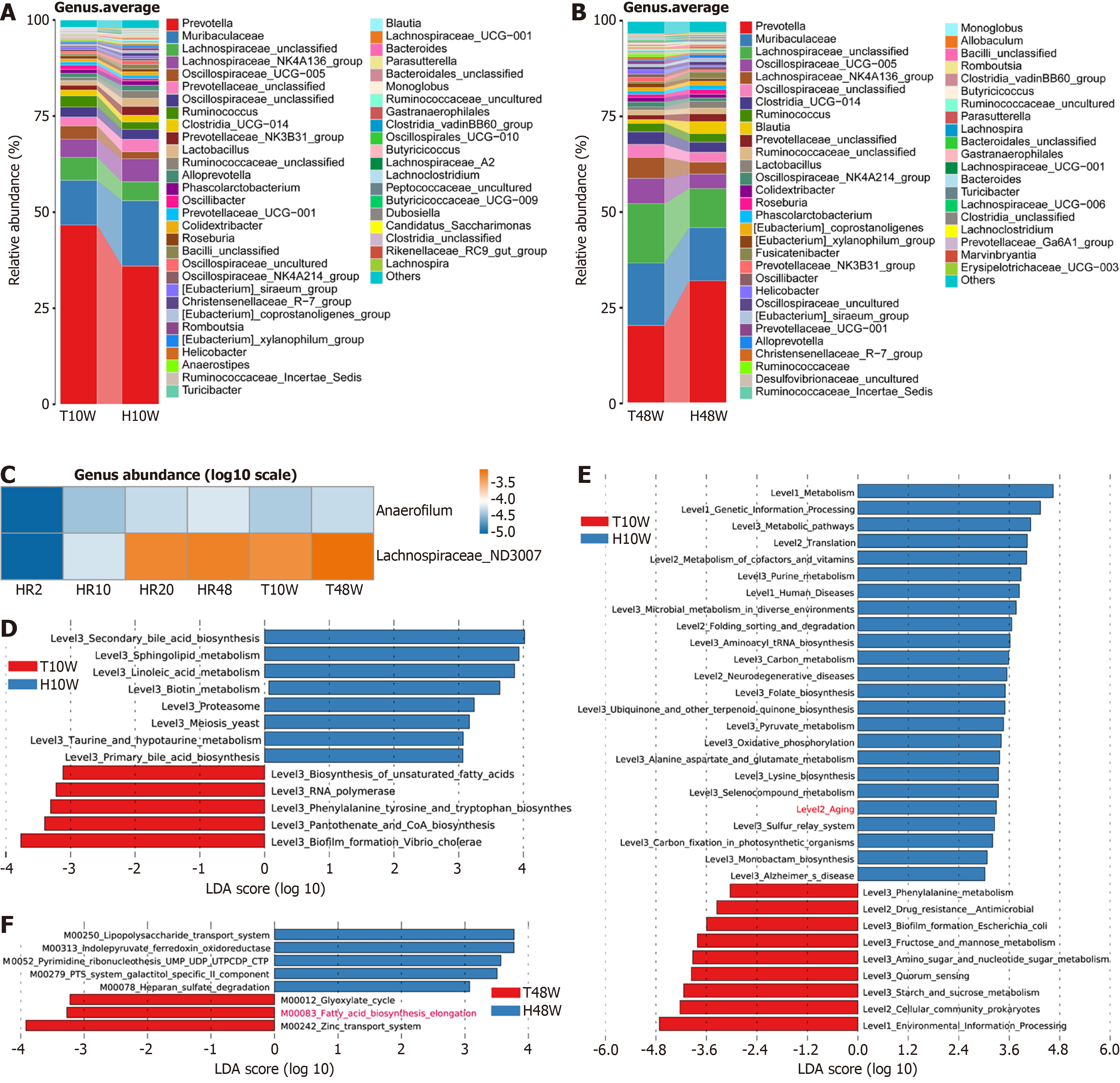
Figure 6 Trimetazidine intervention modulates gut microbiota composition and lipid metabolism of healthy aging rats.
A and B: At the genus level, average relative abundance analysis of differential microbiota after treatment with trimetazidine for 10 weeks (A) and 48 weeks (B); C: At the genus level, correlation analysis between microbial abundance and age; D and E: Kyoto Encyclopedia of Genes and Genomes (KEGG) module analysis in the trimetazidine intervention group collected at 10 weeks and healthy control group collected at 10 weeks (P < 0.05, linear discriminant analysis > 3.0); F: Gut microbe functional potential in the trimetazidine intervention group collected at 48 weeks and healthy control group collected at 48 weeks determined by KEGG analysis. T10W: Trimetazidine intervention group collected at 10 weeks; T48W: Trimetazidine intervention group collected at 48 weeks; H10W: Healthy control group collected at 10 weeks; H48W: Healthy control group collected at 48 weeks; HR: Healthy rat cohort; HR2: Aged 2 weeks; HR10: Aged 10 weeks; HR20: Aged 20 weeks; HR48: Aged 48 weeks; LDA: Linear discriminant analysis.
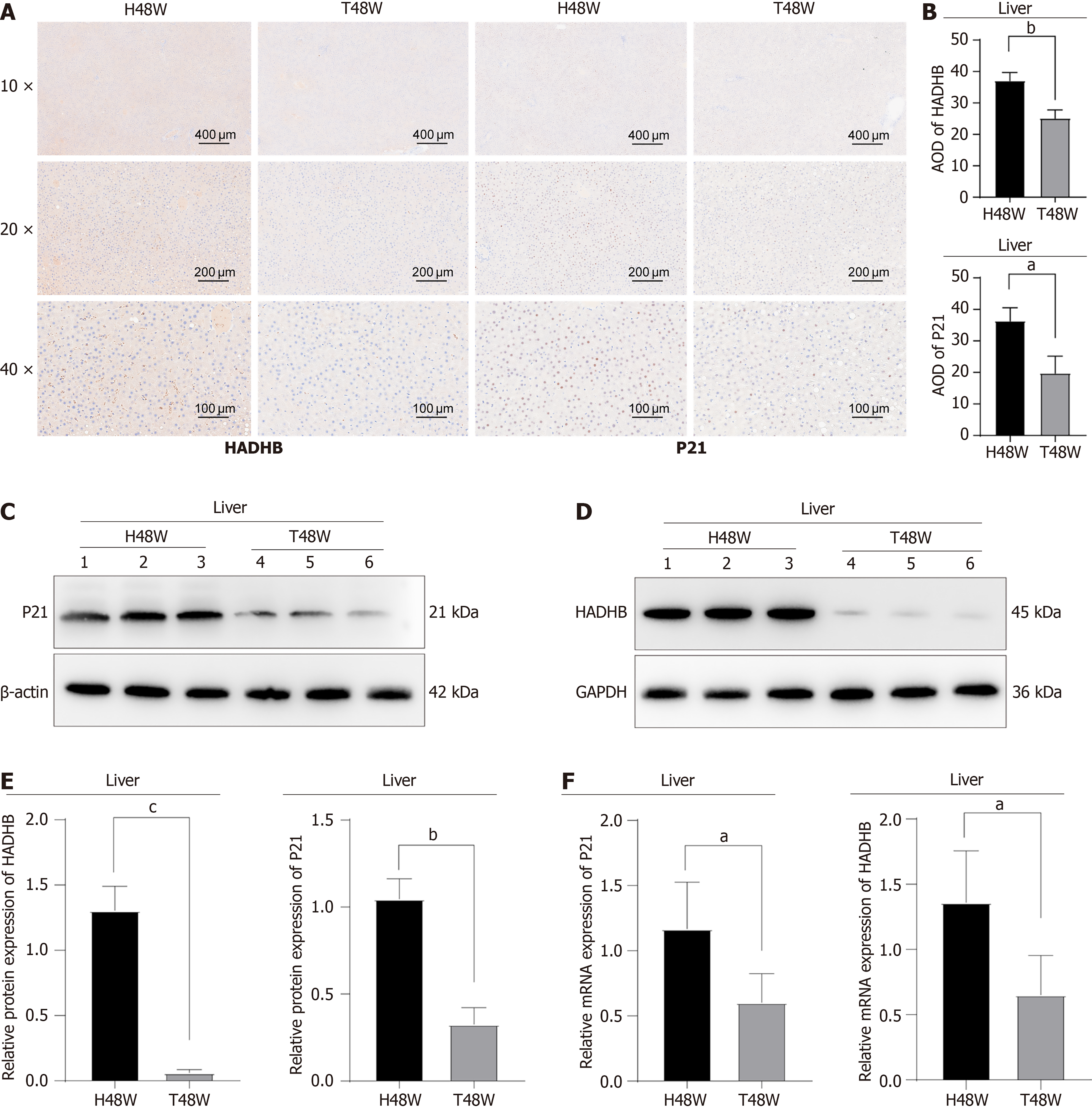
Figure 7 Phenotypic studies of trimetazidine regulation of fatty acid oxidation and aging.
A: Immunohistochemical staining of P21 and HADHB in liver tissues; B: Statistical analysis; C-F: The protein and messenger RNA levels of P21 and HADHB in liver tissues (n = 3 biological replicates) and statistical analysis (mean ± SEM, n = 3). aP < 0.05. bP < 0.01. cP < 0.001. T48W: Trimetazidine intervention group collected at 48 weeks; H48W: Healthy control group collected at 48 weeks; AOD: Average optical density; GAPDH: Glyceraldehyde-3-phosphate dehydrogenase; mRNA: Messenger RNA.
- Citation: Wang XM, Rao BC, Su GY, Wang HY, Zhang GZ, Ge FL, Yu ZJ, Ren ZG, Liang HX. Age-related gut microbiome profile and reversal of microbial imbalance by trimetazidine intervention. World J Gastroenterol 2026; 32(16): 115969
- URL: https://www.wjgnet.com/1007-9327/full/v32/i16/115969.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i16.115969