Copyright: ©Author(s) 2026.
World J Gastroenterol. Apr 21, 2026; 32(15): 116679
Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.116679
Published online Apr 21, 2026. doi: 10.3748/wjg.v32.i15.116679
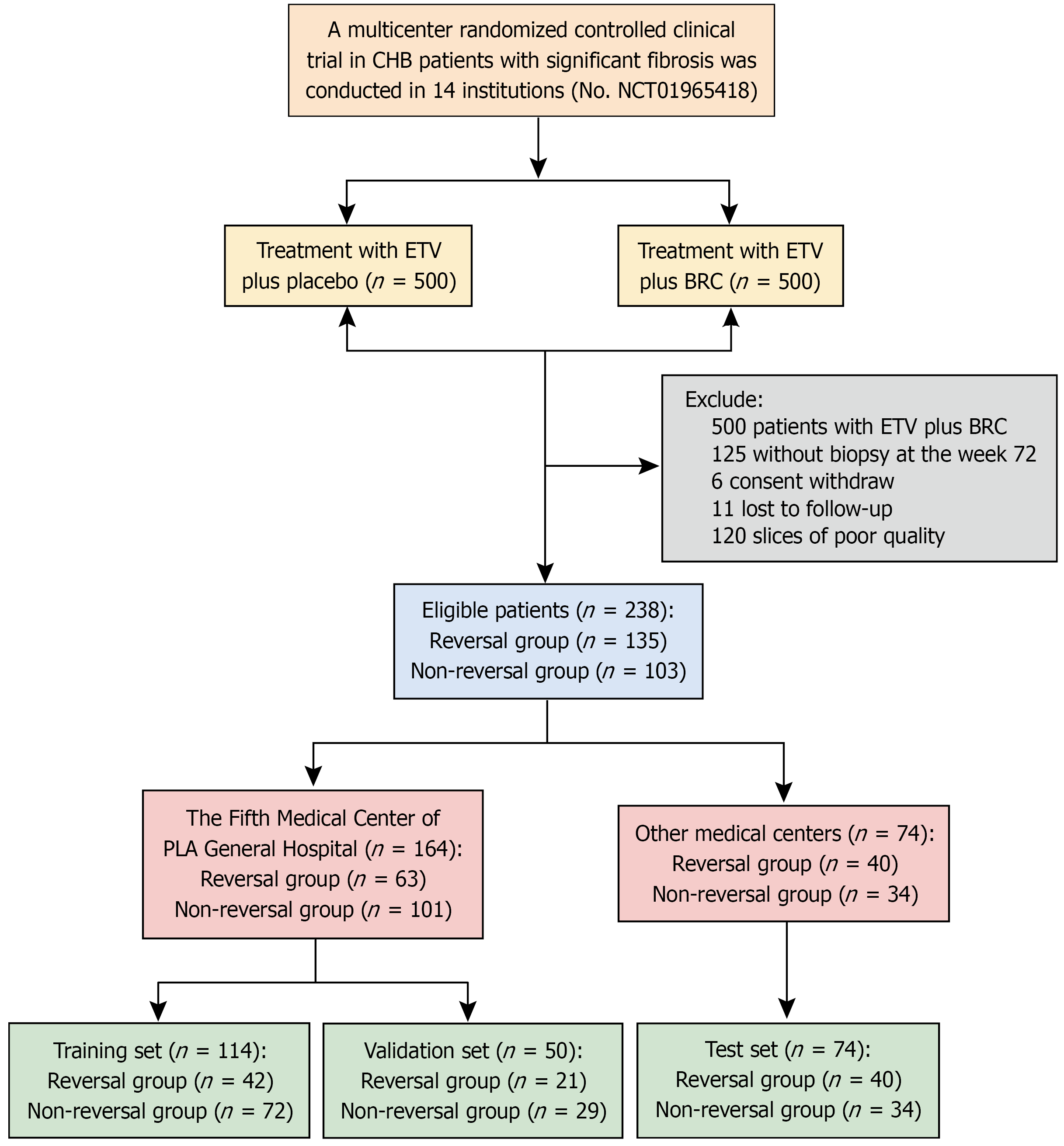
Figure 1 Flow diagram of the study population.
CHB: Chronic hepatitis B; ETV: Entecavir; BRC: Biejia-Ruangan tablet.
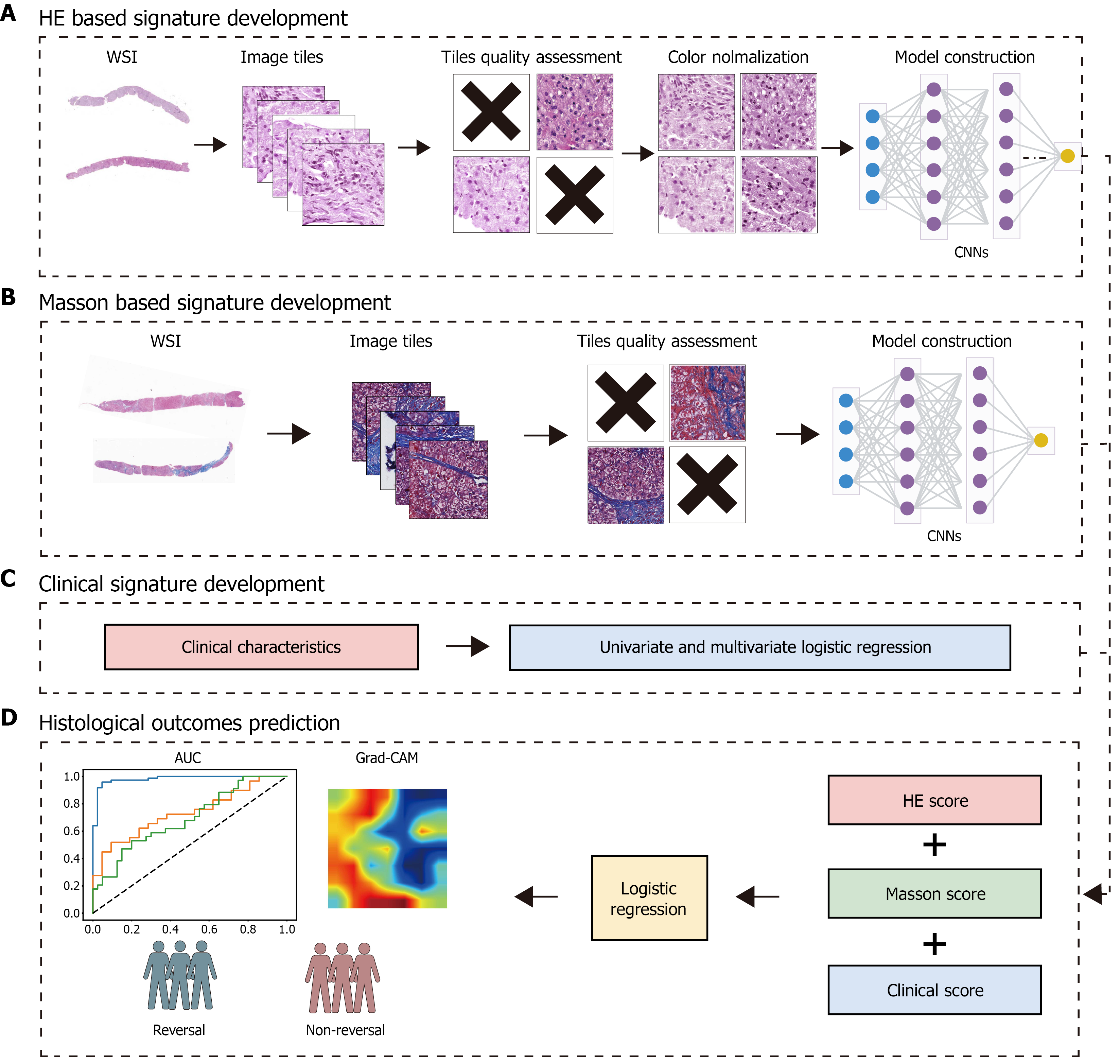
Figure 2 Workflow of the development and testing of deep learning model.
A and B: Digital hematoxylin and eosin (HE)- and Masson-stained slides were first categorized according to whether fibrosis reversal occurred. Whole-slide images were then partitioned into 512 × 512-pixel tiles and subjected to quality control to remove non-informative regions. Color normalization was applied to the HE tiles, whereas Masson-stained tiles were retained in their original color. The quality-controlled tiles, along with their corresponding labels, were used to train convolutional neural networks models; C: Univariate and multivariate logistic regression analyses were performed to identify clinical characteristics associated with histological outcomes; D: A logistic regression fusion model was developed using HE score, Masson score, and clinical score as input variables, followed by performance validation on internal and external validation sets. WSI: Whole-slide image; CNNs: Convolutional neural networks; Grad-CAM: Gradient-weighted Class Activation Mapping; HE: Hematoxylin and eosin; AUC: Area under the receiver operating characteristic curve.
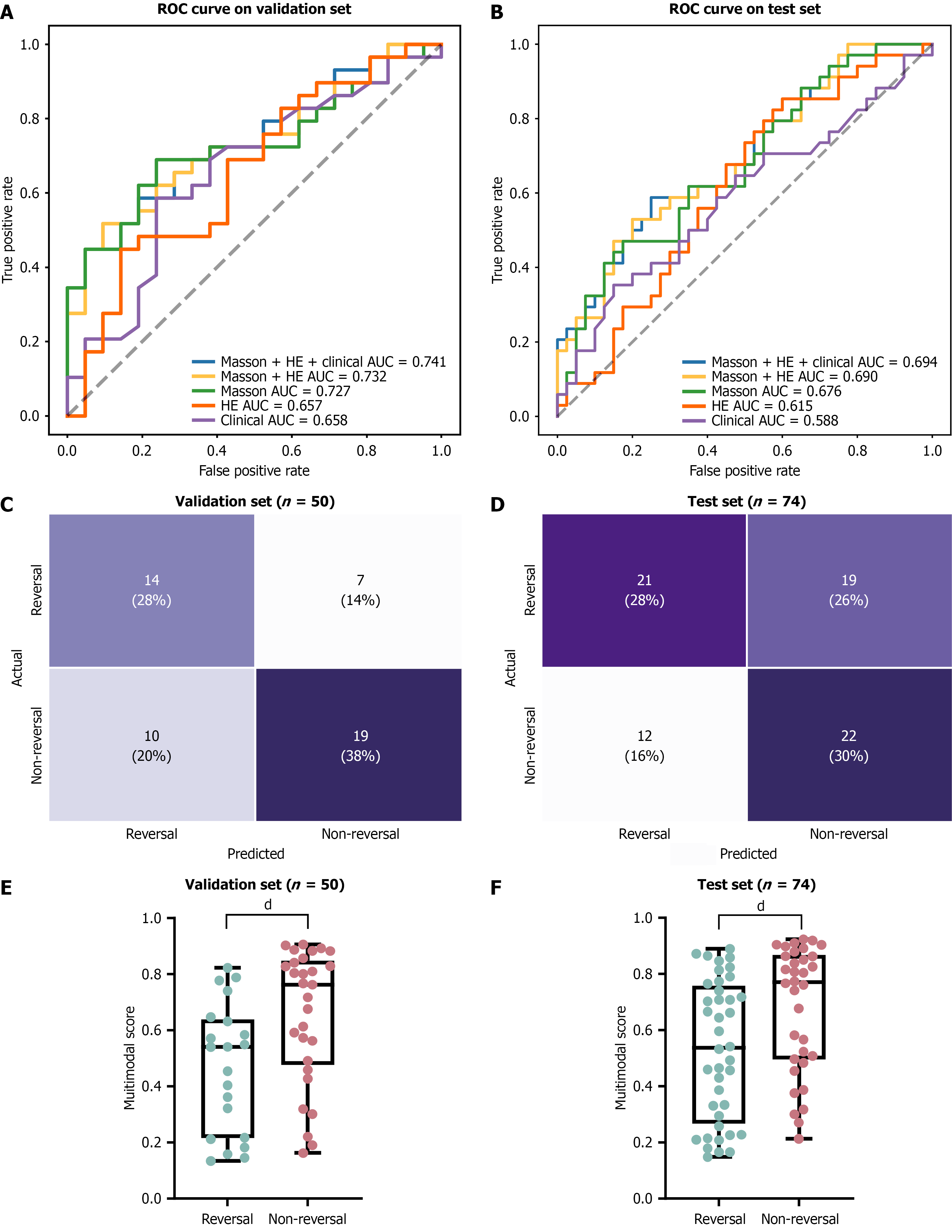
Figure 3 Model performance.
A and B: Performance of different model (purple for clinical-based, orange for hematoxylin and eosin-based, green for Masson-based, yellow for hematoxylin and eosin and Masson-based, blue for multimodal model) in the validation and test sets; C and D: Confusion matrix of the fusion model in the validation and test sets; E and F: Differences in model scores between the reversal and non-reversal groups in the validation and test sets. dP < 0.0001. ROC: Receiver operating characteristic; HE: Hematoxylin and eosin; AUC: Area under the receiver operating characteristic curve.
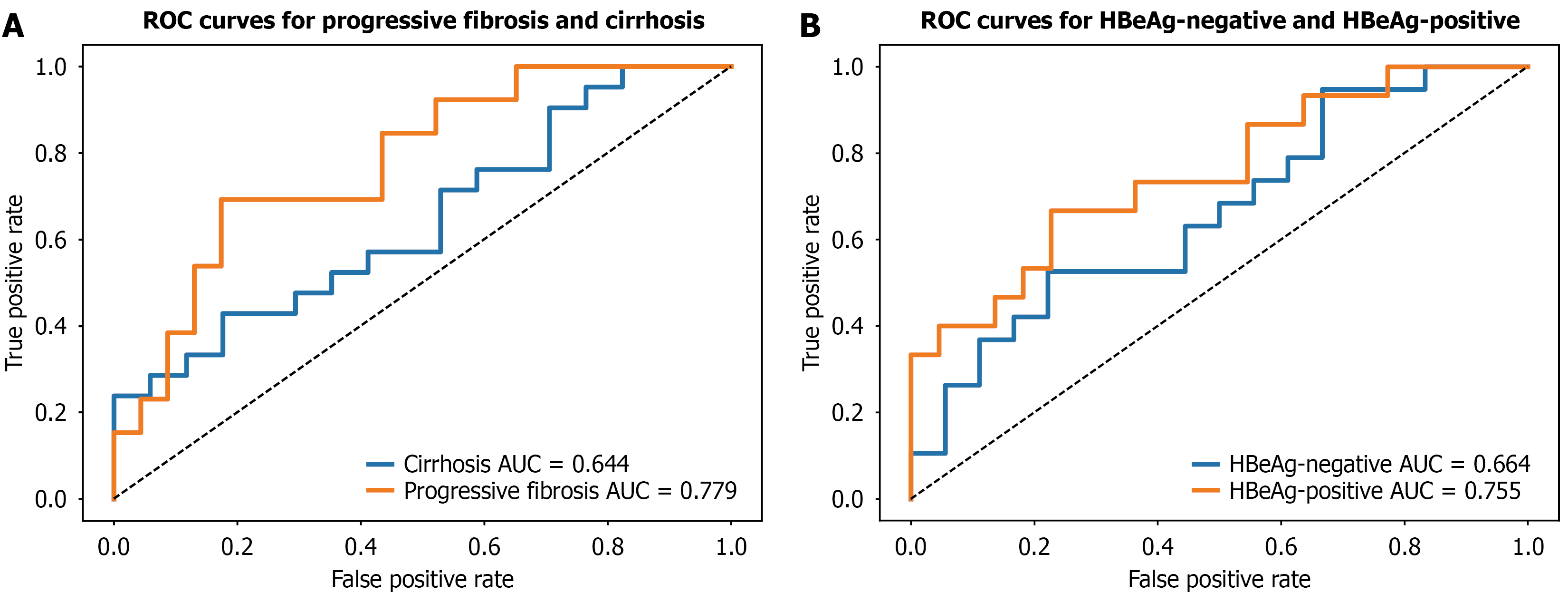
Figure 4 Subgroup analysis.
A: Multimodal model performance in the progressive fibrosis (orange) and cirrhosis (blue) subgroup; B: Multimodal model performance in the hepatitis B e antigen-positive (orange) and hepatitis B e antigen-negative subgroup (blue). HBeAg: Hepatitis B e antigen; ROC: Receiver operating characteristic; AUC: Area under the receiver operating characteristic curve.
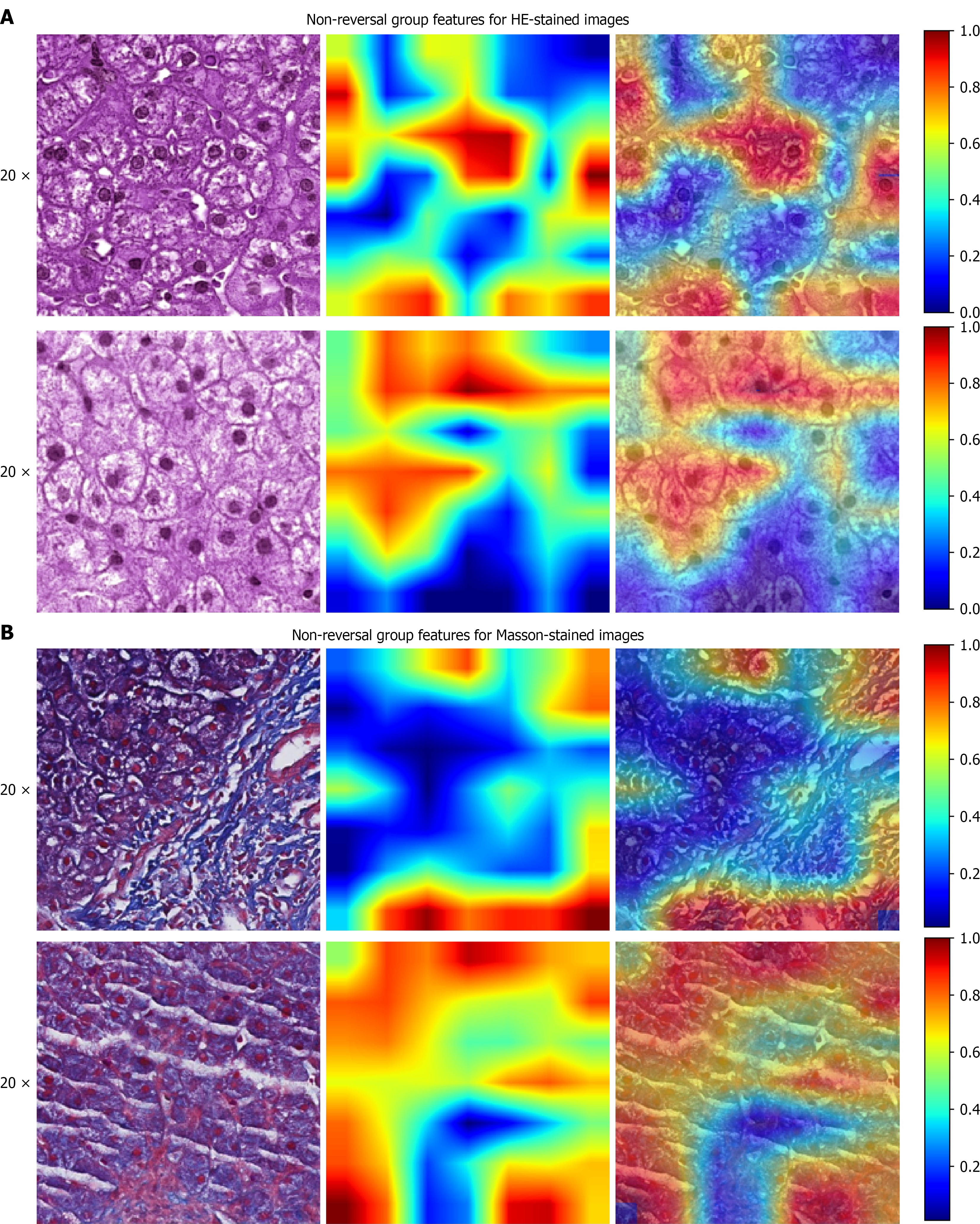
Figure 5 Saliency map.
A: Predicted non-reversal hematoxylin and eosin image (left), its corresponding saliency map (center), and the overlaid composite image (right). Darker red regions indicate more prominent features associated with non-reversal; B: Predicted non-reversal Masson-stained image (left), its corresponding saliency map (center), and the overlaid composite image (right). Darker red regions indicate more prominent features associated with non-reversal. HE: Hematoxylin and eosin.
- Citation: Han W, Cheng DY, He QW, Wang SH, Gong SJ, Chen Y, Yang YP. Deep learning-based multimodal model for predicting on-treatment histological outcomes in chronic hepatitis B-associated advanced liver fibrosis. World J Gastroenterol 2026; 32(15): 116679
- URL: https://www.wjgnet.com/1007-9327/full/v32/i15/116679.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i15.116679