Copyright: ©Author(s) 2026.
World J Gastroenterol. Apr 7, 2026; 32(13): 115536
Published online Apr 7, 2026. doi: 10.3748/wjg.v32.i13.115536
Published online Apr 7, 2026. doi: 10.3748/wjg.v32.i13.115536
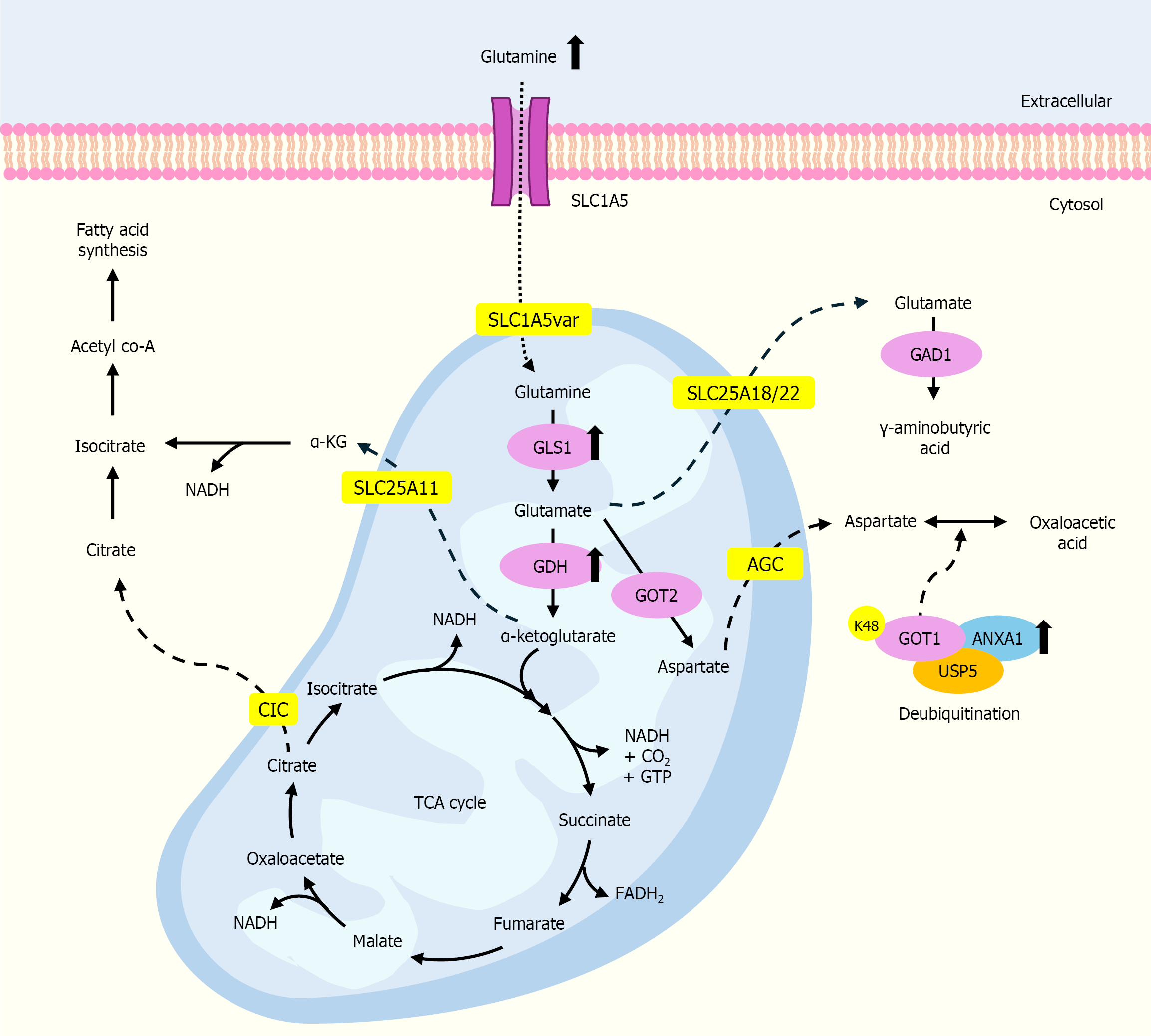
Figure 1 The alteration of glutamine metabolism in cholangiocarcinoma cells.
Glutaminolysis begins with the transportation of glutamine through the inner mitochondrial membrane through solute carrier family 1 member 5, followed by the conversion of glutamine to glutamate by GLSs. Mitochondrial glutamate can be converted to alpha-ketoglutarate by GDH1. Aspartate can be exported to the cytoplasm, undergo transamination, and converted to oxaloacetate by glutamate-oxaloacetate transaminase (GOT1). Annexin A1 generally functions as a scaffold protein, connecting ubiquitin-specific protease 5 with GOT1 and facilitating their interaction. This scaffolding activity promotes GOT1 deubiquitylation and stabilization, inhibiting its degradation via the ubiquitin-proteasome pathway. SLC1A5: Solute carrier family 1 member 5; AGC: Aspartate-glutamate carrier; GLS1: Glutaminase 1; GDH: Glutamate dehydrogenase; GOT: Glutamic-oxaloacetic tran
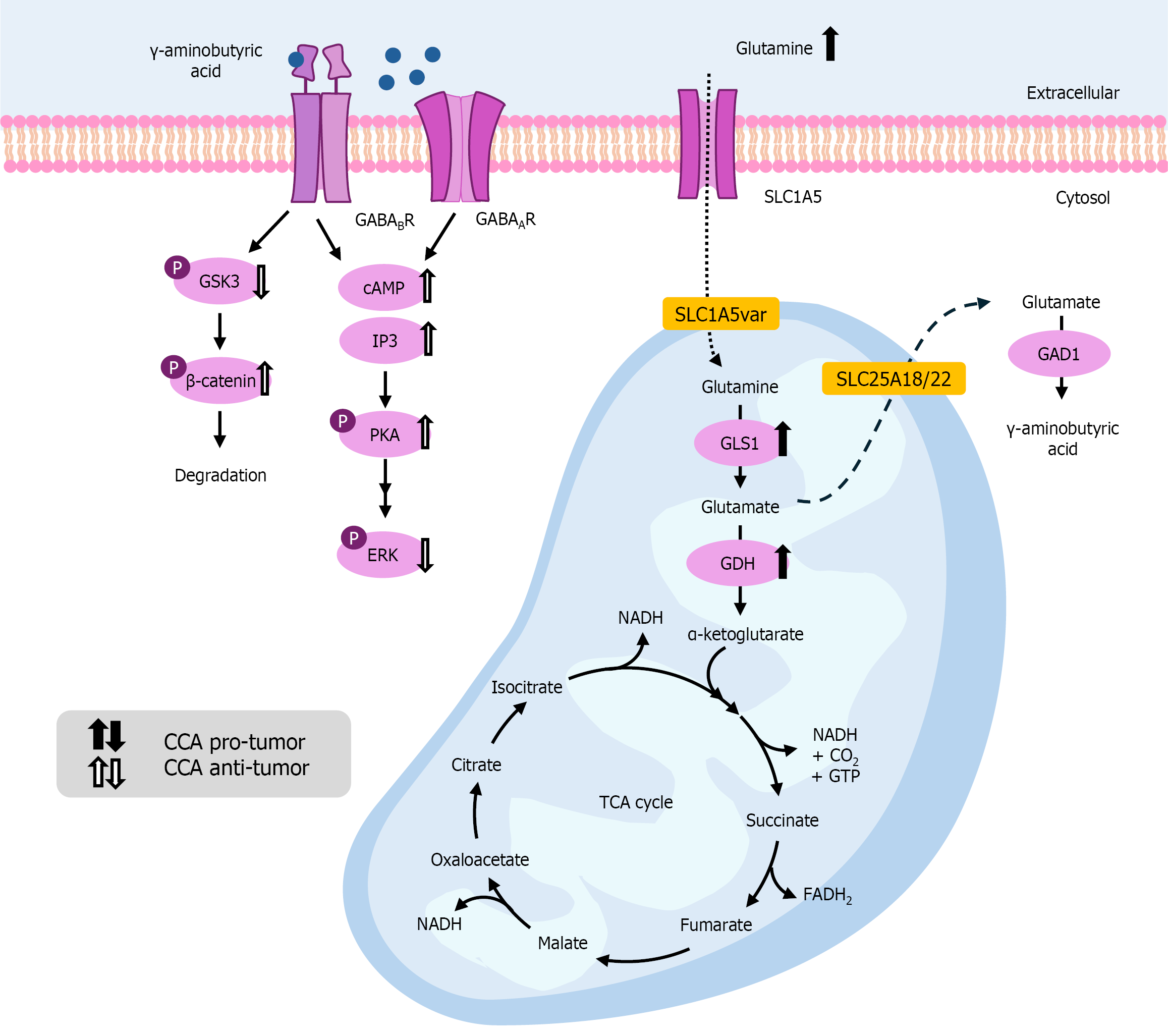
Figure 2 Metabolic pathways of gamma-aminobutyric acid and the effects of gamma-aminobutyric acid analogs on cholangiocarcinoma cell biology.
Gamma-aminobutyric acid (GABA) inhibits cholangiocarcinoma (CCA) cell proliferation via both cyclic AMP/protein kinase A and d-myo-inositol-1,4,5-triphosphate/Ca2+-dependent pathways, leading to downregulation of phosphorylated extracellular signal-regulated kinase 1/2. Moreover, an aminobutyric acid B2 receptor agonist can inhibit glycogen synthase kinase 3 phosphorylation, thereby further phosphorylating β-catenin, leading to its degradation. The up or down direction of the arrows indicates increased or decreased levels in CCA cells. Black arrows denote the pro-tumor effects of the genes or intermediates, while the white arrows denote their anti-tumor effects. GABA: γ-aminobutyric acid; GLS1: Glutaminase 1; GDH: Glutamate dehydrogenase; GAD1: Glutamate decarboxylase 1; cAMP: Cyclic adenosine monophosphate; IP3: Inositol trisphosphate; PKA: Protein kinase A; ERK: Extracellular signal-regulated kinase; GSK3: Glycogen synthase kinase-3.
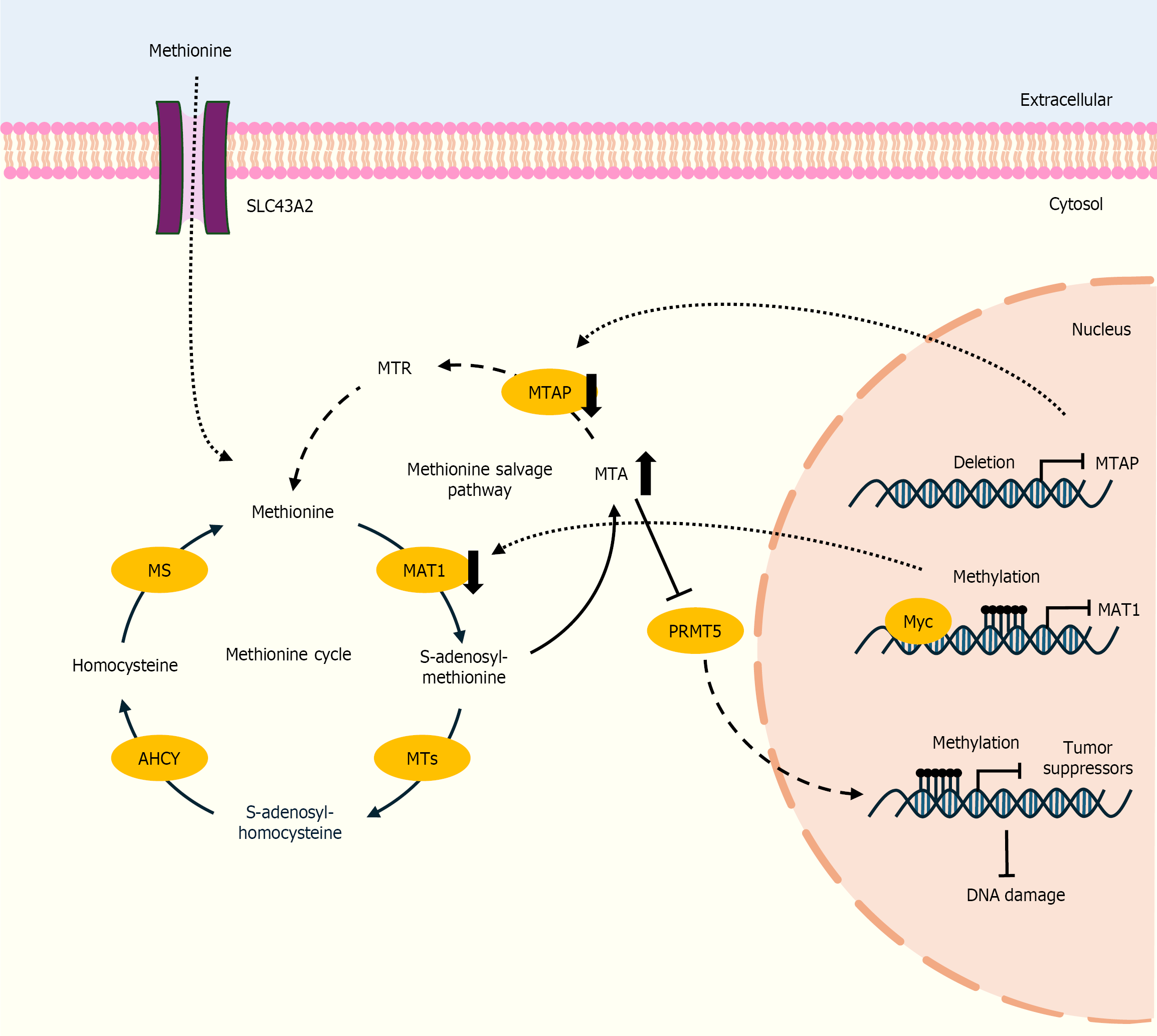
Figure 3 Metabolic pathways and the alteration of methionine metabolism in cholangiocarcinoma cells, including the methionine cycle and the methionine salvage pathway.
Loss of methylthioadenosine phosphorylase in the methionine salvage pathway leads to the accumulation of 5'-methylthioadenosine, thereby creating a hypomorphic state of protein arginine methyltransferase 5 (PRMT5), which selectively sensitizes cells to PRMT5 inhibition. The methionine adenosyltransferase 1 promoter is hypermethylated due to c-Myc-mediated transcriptional suppression, leading to increased tumor growth, invasion, and metastasis. The up or down direction of the arrows indicates whether levels in cholangiocarcinoma cells are increased or decreased. MTR: 5-methylthioribose; MTAP: Methylthioadenosine phosphorylase; MTA: 5'-methylthioadenosine; MAT1: Methionine adenosyltransferase 1; MS: Methionine synthase; AHCY: Ade
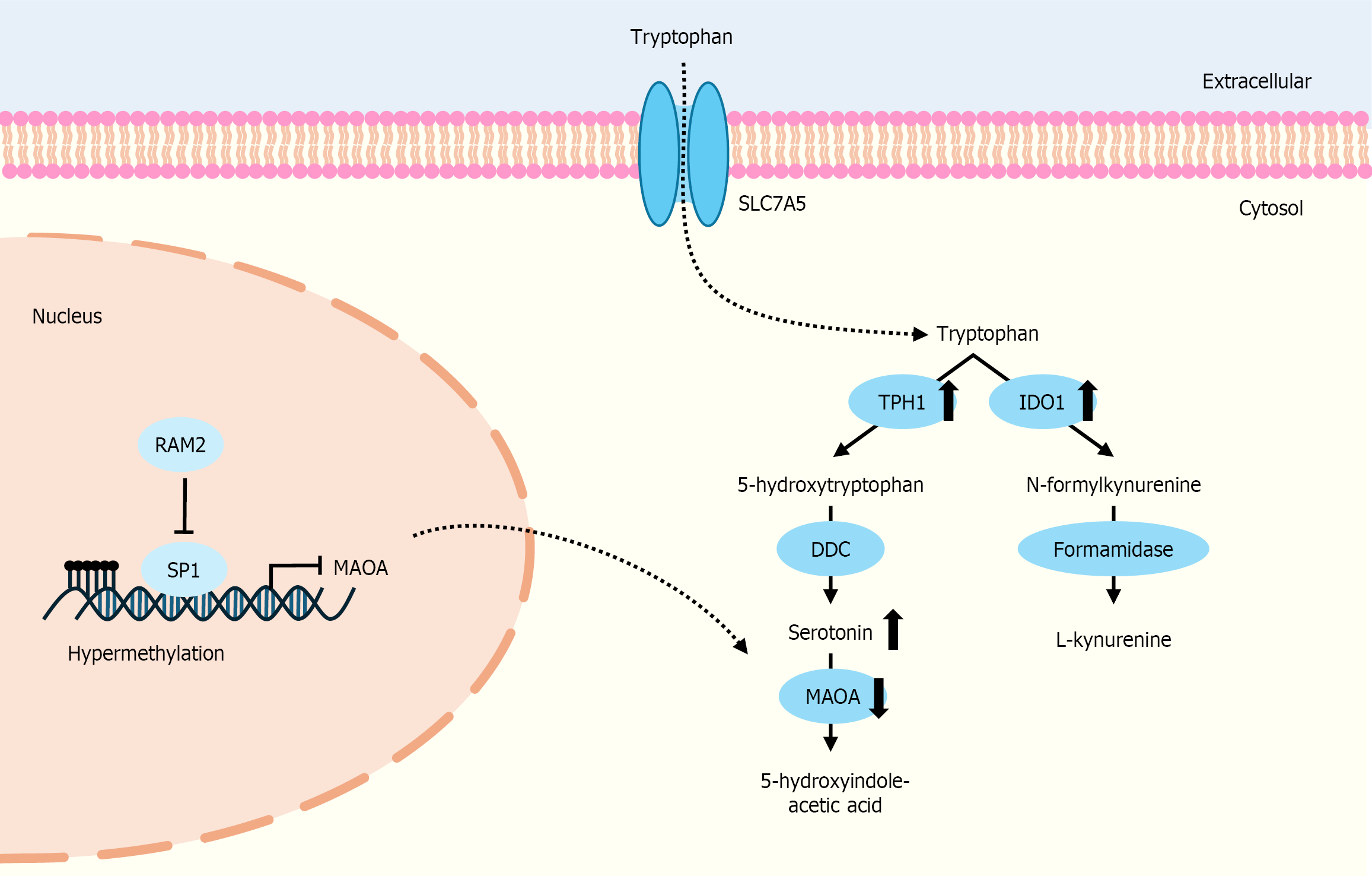
Figure 4 Metabolisms of tryptophan and its roles in cholangiocarcinoma cells, including the kynurenine pathway and serotonin pathway.
In the serotonin pathway, monoamine oxidase A (MAOA) expression and catalytic activity can be regulated by the transcription factor SP-1, which binds to the MAOA promoter. R1 repressor competitively binds to the SP-1 site on the MAOA core promoter, leading to suppression of MAOA gene expression. The up or down direction of the arrows indicates increased or decreased levels in cholangiocarcinoma cells. MAOA: Monoamine oxidase A; RAM2: R1 repressor; TPH1: Tryptophan hyd
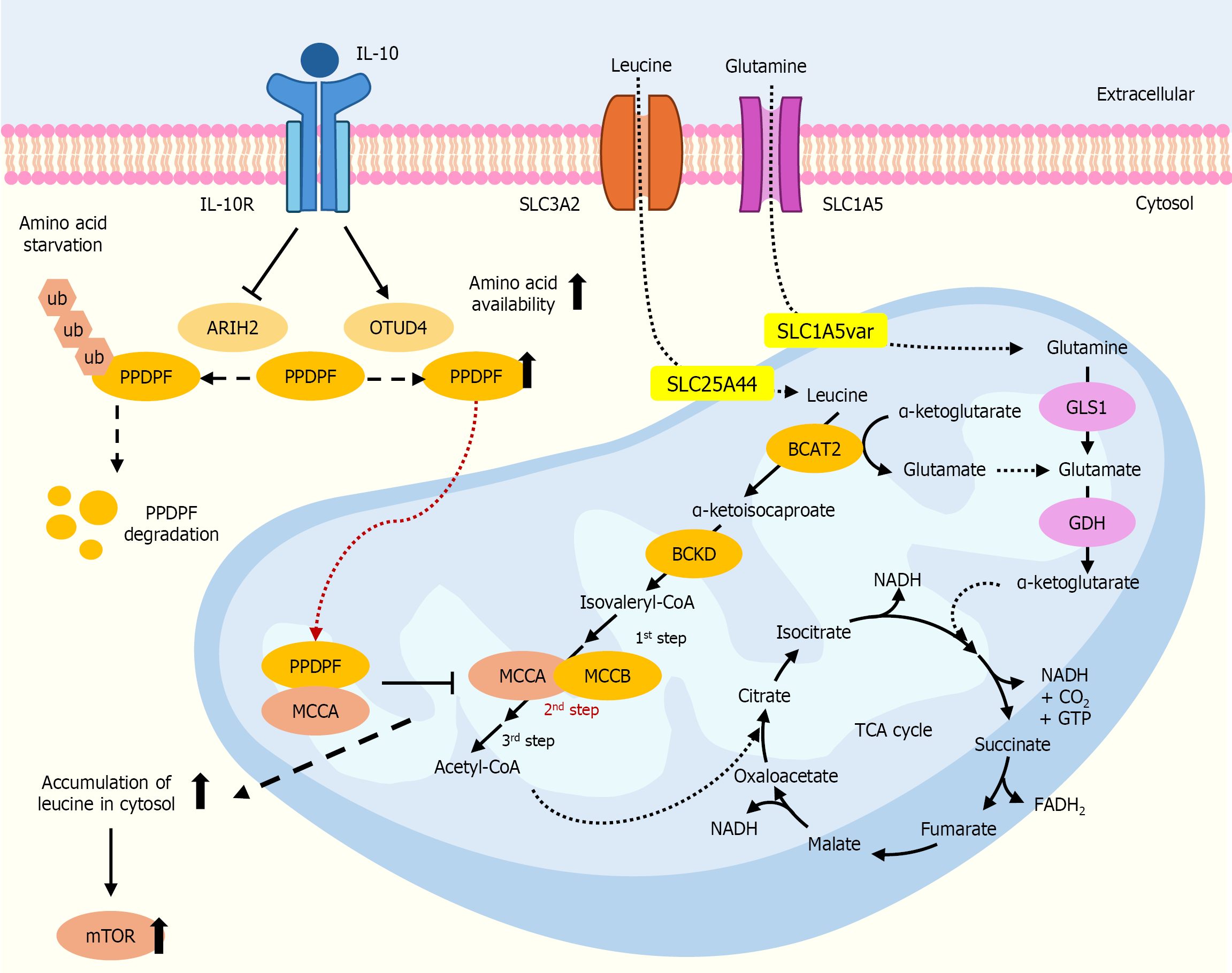
Figure 5 Branch chain amino acid metabolism and its alteration in cholangiocarcinoma cells.
The level of pancreatic progenitor cell differentiation and proliferation factor (PPDPF) protein is stabilized by OTU deubiquitinase 4, and leucine catabolism is inhibited, leading to its accumulation in the cytosol. Upon amino acid starvation, the PPDPF protein is ubiquitously degraded upon binding to ariadne homolog 2. The up or down direction of the arrows indicates increased or decreased levels in cholangiocarcinoma cells. The dashed arrow refers to unclarified mechanisms. IL-10: Interleukin-10; IL-10R: Interleukin-10 receptor; ARIH2: Ariadne homolog 2; OTUD4: OTU deubiquitinase 4; PPDPF: Pancreatic progenitor cell differentiation and proliferation factor; BCAT2: Branched chain amino acid transaminase 2; BCKD: Branched chain ketoacid dehydrogenase kinase; MCC: Methylcrotonyl-CoA carboxylase; mTOR: Mammalian target of rapamycin; GLS1: Glutaminase 1; GDH: Glutamate dehydrogenase.
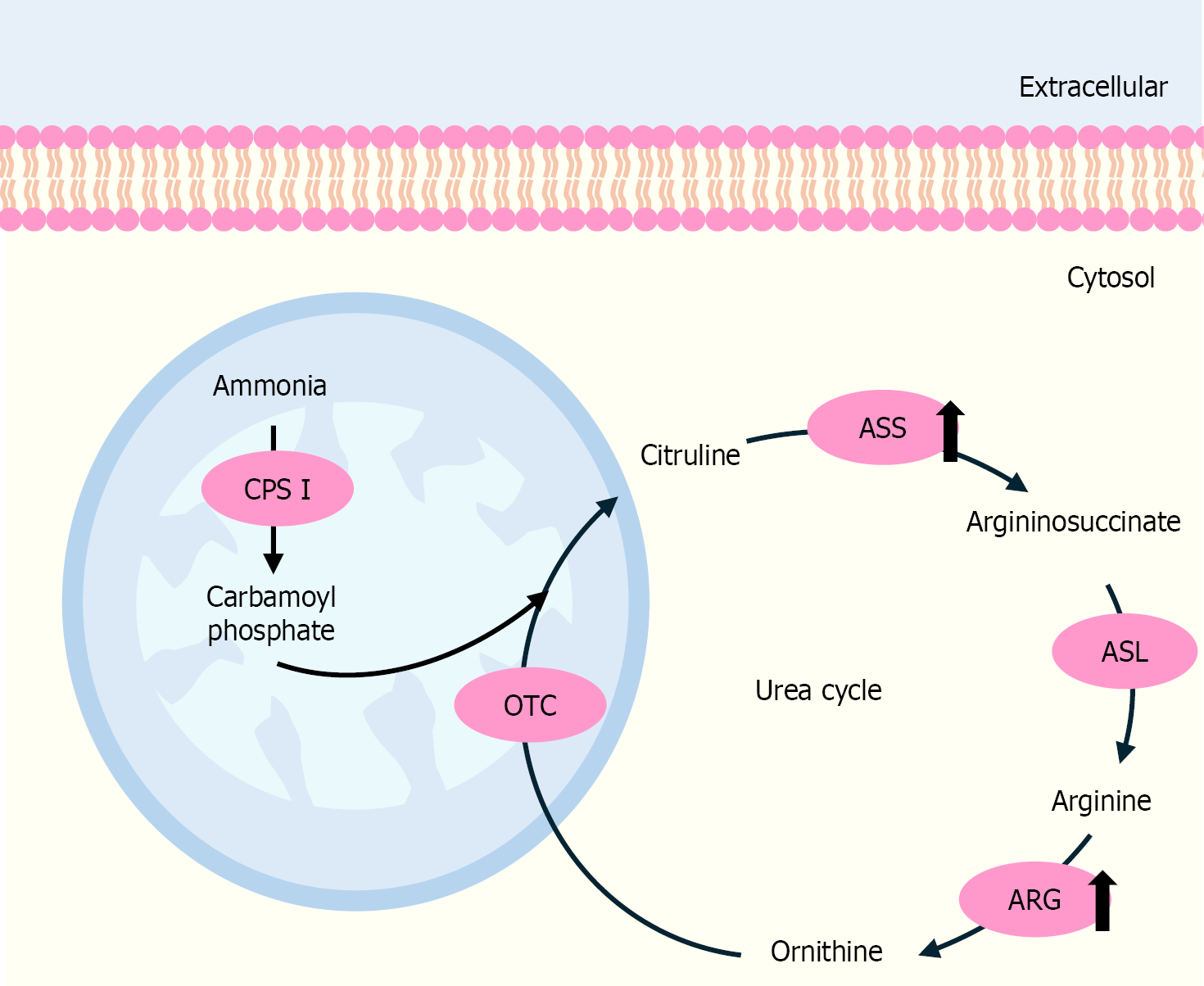
Figure 6 The alteration of the urea cycle’s enzymes in cholangiocarcinoma progression.
Alteration of enzymes in the urea cycle is associated with aggressive phenotypes of cholangiocarcinoma (CCA) cells and poor clinicopathological characteristics of patients. The up or down direction of the arrows indicates increased or decreased levels in CCA cells. CPS1: Carbamoyl phosphate synthetase 1; OTC: Ornithine transcarbamylase; ASS: Argininosuccinate synthetase; ASL: Argininosuccinate lyase; ARG: Arginase.
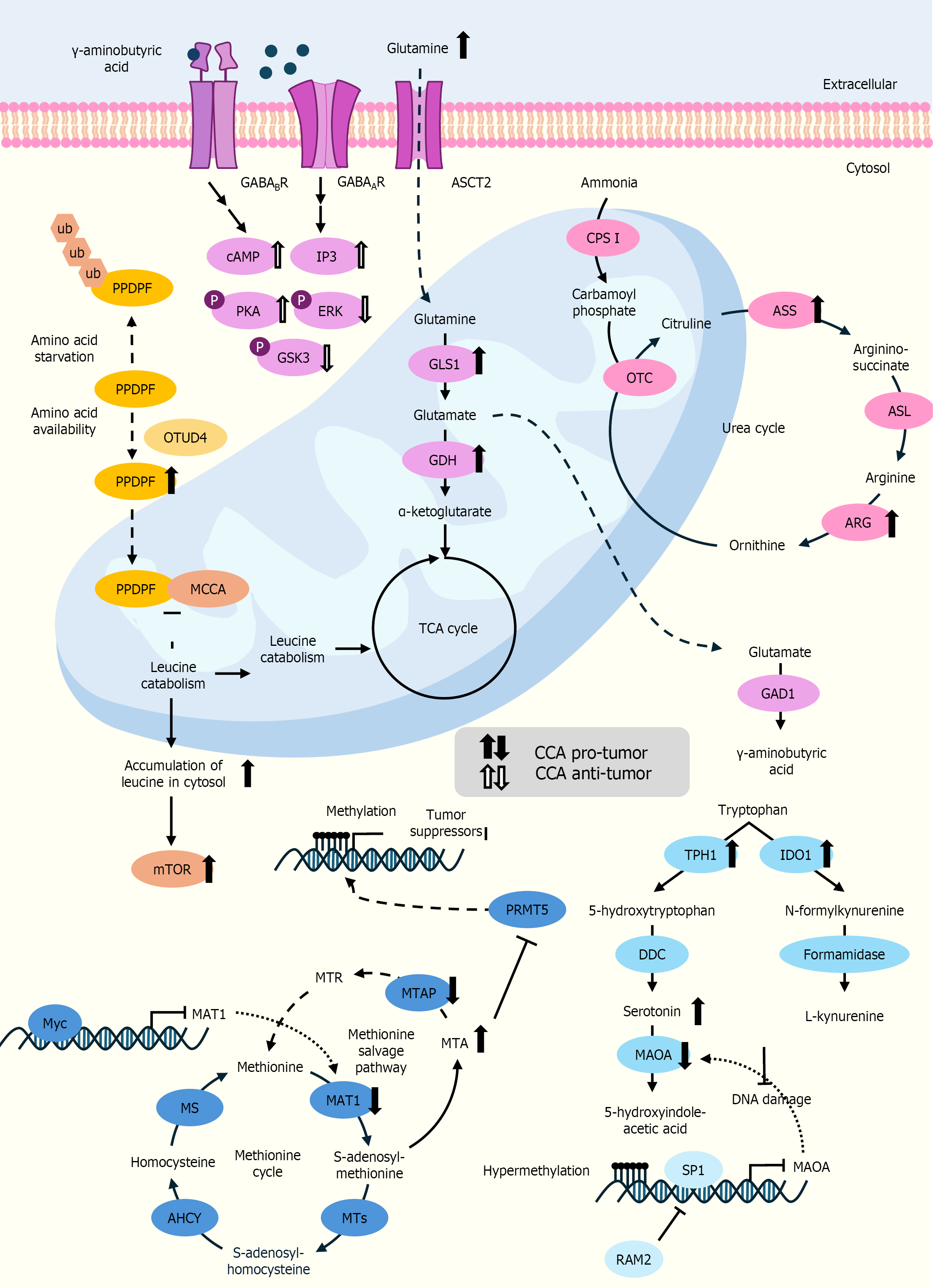
Figure 7 A comprehensive view of amino acid metabolism alteration in cholangiocarcinoma.
The up or down direction of the arrows indicates increased or decreased levels in cholangiocarcinoma cells. The black arrows denote the pro-tumor effects of the genes or intermediates, while the white arrow denotes their anti-tumor effects. AGC: Aspartate-glutamate carrier; GLS1: Glutaminase 1; GDH: Glutamate dehydrogenase; GOT: Glutamic-oxaloacetic transa
- Citation: Khawkhiaw K, Lert-Itthiporn W, Seyedasli N, Chiu CF, Saengboonmee C. Reprogramming of amino acid metabolism in cholangiocarcinoma: A potential target for metabolic-targeted therapy. World J Gastroenterol 2026; 32(13): 115536
- URL: https://www.wjgnet.com/1007-9327/full/v32/i13/115536.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i13.115536