©The Author(s) 2024.
World J Gastroenterol. May 14, 2024; 30(18): 2454-2466
Published online May 14, 2024. doi: 10.3748/wjg.v30.i18.2454
Published online May 14, 2024. doi: 10.3748/wjg.v30.i18.2454
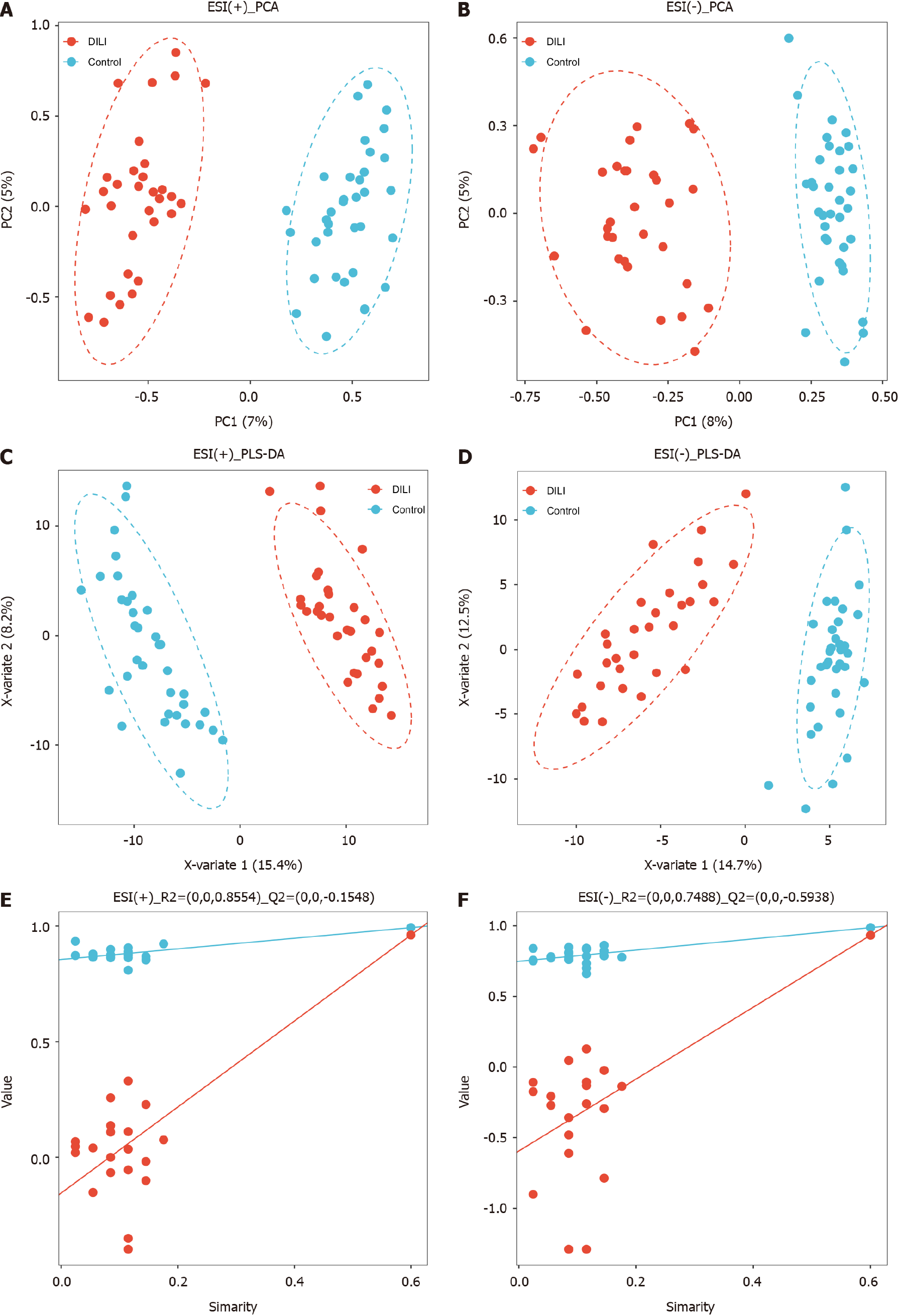
Figure 1 Multivariate statistical analysis.
A: Principal component analysis (PCA) scatter plots of the two groups in positive mode; B: PCA scatter plots of the two groups in negative mode; C: Partial least squares-discriminant analysis (PLS-DA) scatter plots of the two groups in positive mode; D: PLS-DA scatter plots of the two groups in negative mode; E: Cross-validation plot of the two groups with a permutation test repeated 200 times in positive mode; F: Cross-validation plot of the two groups with a permutation test repeated 200 times in negative mode. Healthy controls group (n = 35, blue circles), drug-induced liver injury group (n = 31, red circles). PCA: Principal component analysis; PLS-DA: Partial least squares-discriminant analysis.
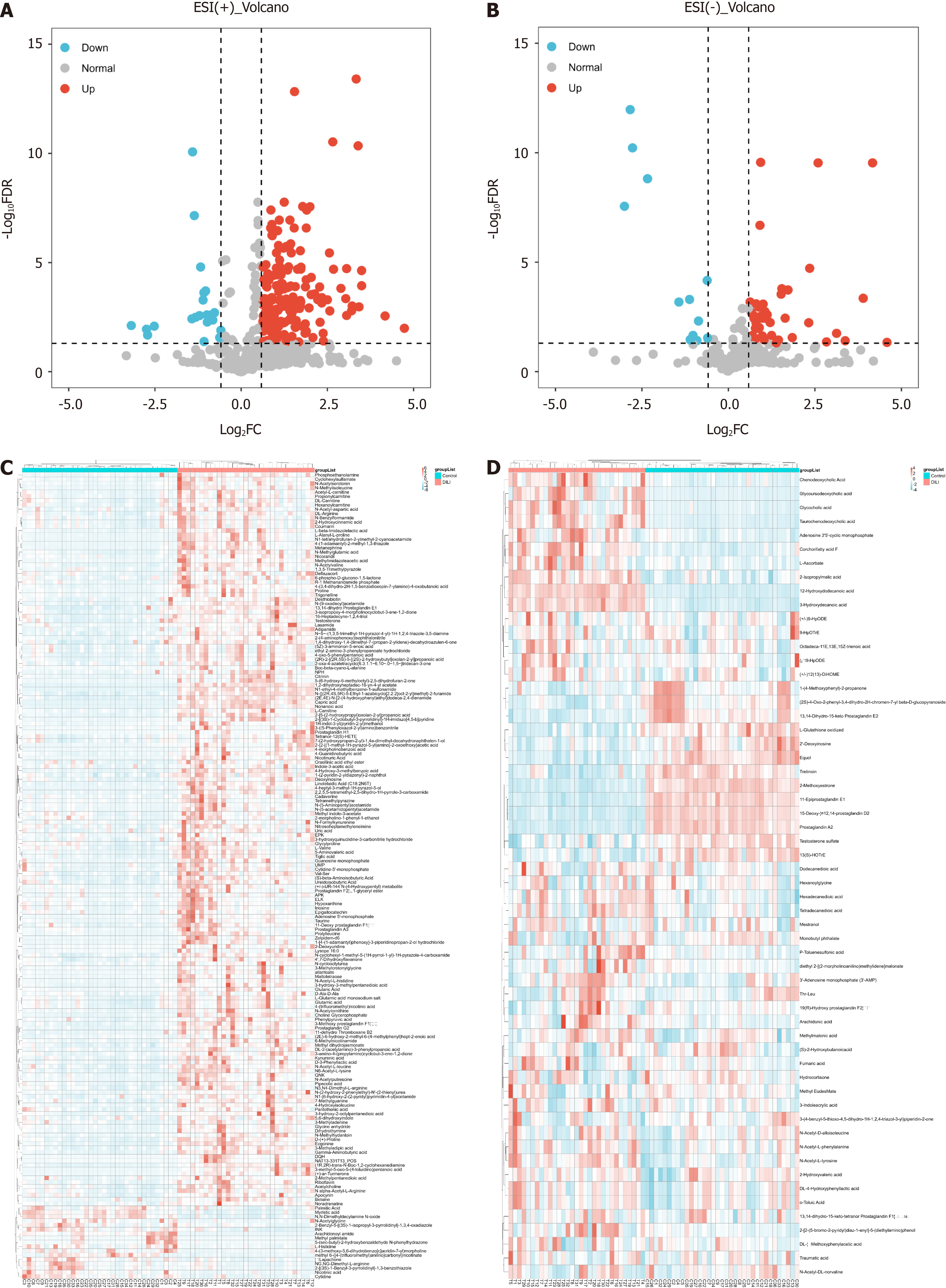
Figure 2 Differentially abundant metabolite analysis.
A: Volcanograms of differentially abundant metabolites between drug-induced liver injury (DILI) and healthy controls (HC) in positive mode; B: Volcanograms of differentially abundant metabolites between DILI and HC in negative mode; C: Heat plots of the differentially abundant metabolites between DILI and HC in positive mode; D: Heat plots of the differentially abundant metabolites between DILI and HC in negative mode.
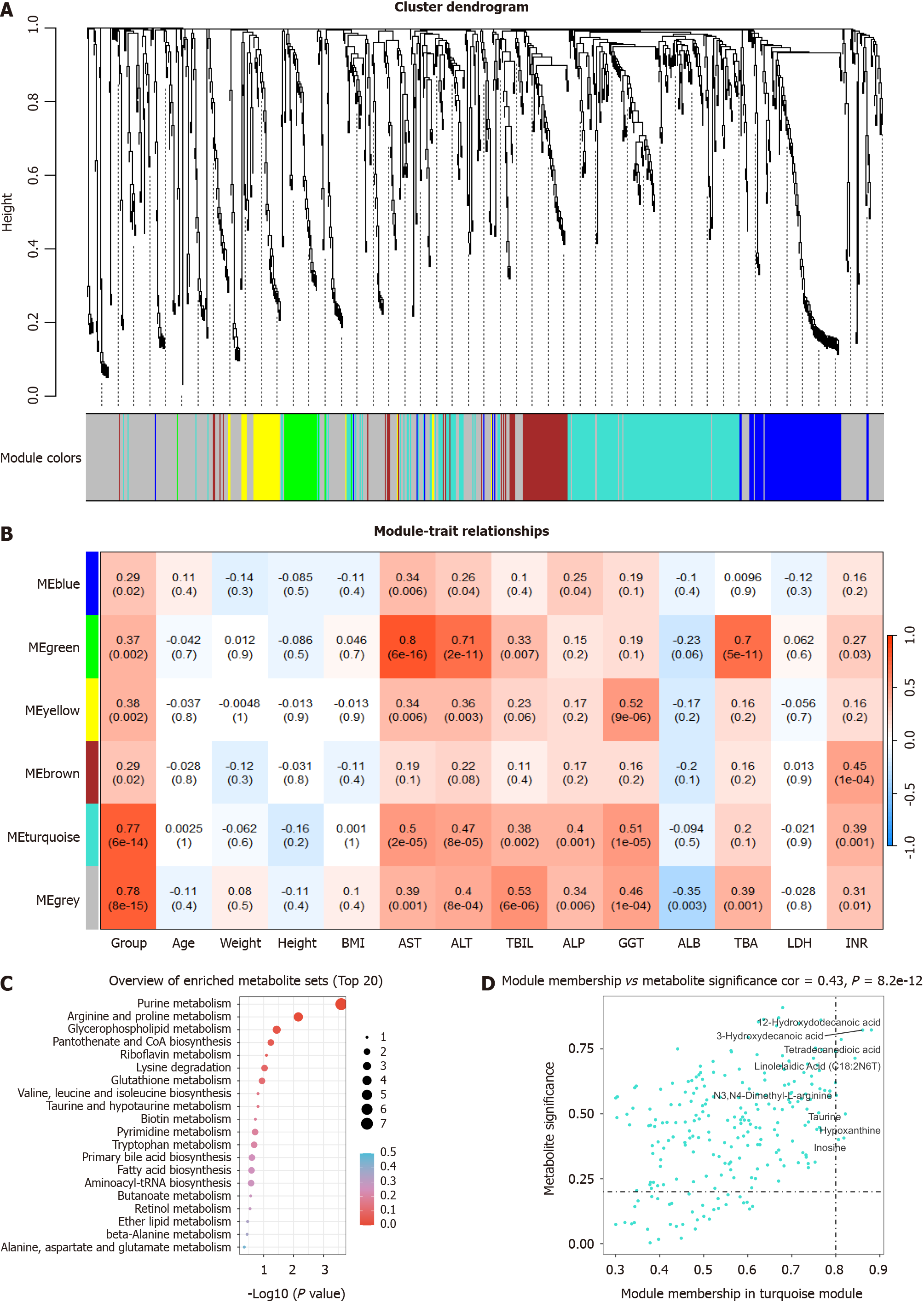
Figure 3 Identification of the hub metabolites using weighted metabolite coexpression network analysis.
A: The branches of the cluster dendrogram for metabolites related to drug-induced liver injury (DILI) traits; B: Heatmap depicting correlations between modules and DILI traits; C: Results of Kyoto Encyclopaedia of Genes and Genomes enrichment analysis for the turquoise module; D: Scatter plot of the turquoise module.
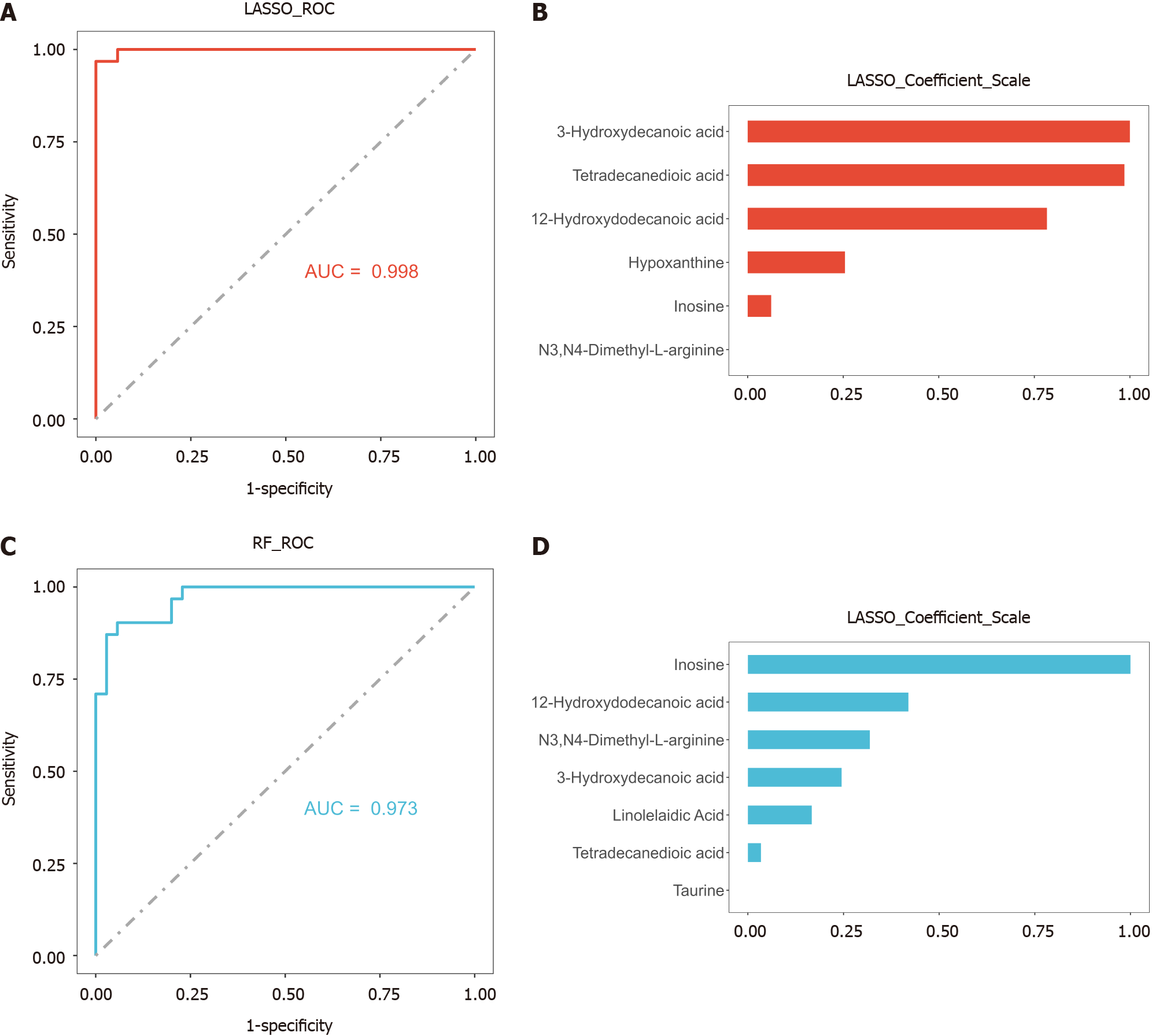
Figure 4 Area under the receiver operating characteristic curves and hub metabolite screening of machine learning algorithms for significantly different metabolites.
A: Area under the receiver operating characteristic (AUROC) curve of the least absolute shrinkage and selection operator (LASSO) model; B: AUROC curve of the random forest (RF) model; C: Hub metabolite screening of LASSO; D: Hub metabolite screening of RF. LASSO: Least absolute shrinkage and selection operator; RF: Random forest; AUC: Area under the curve; ROC: Receiver operating characteristic.
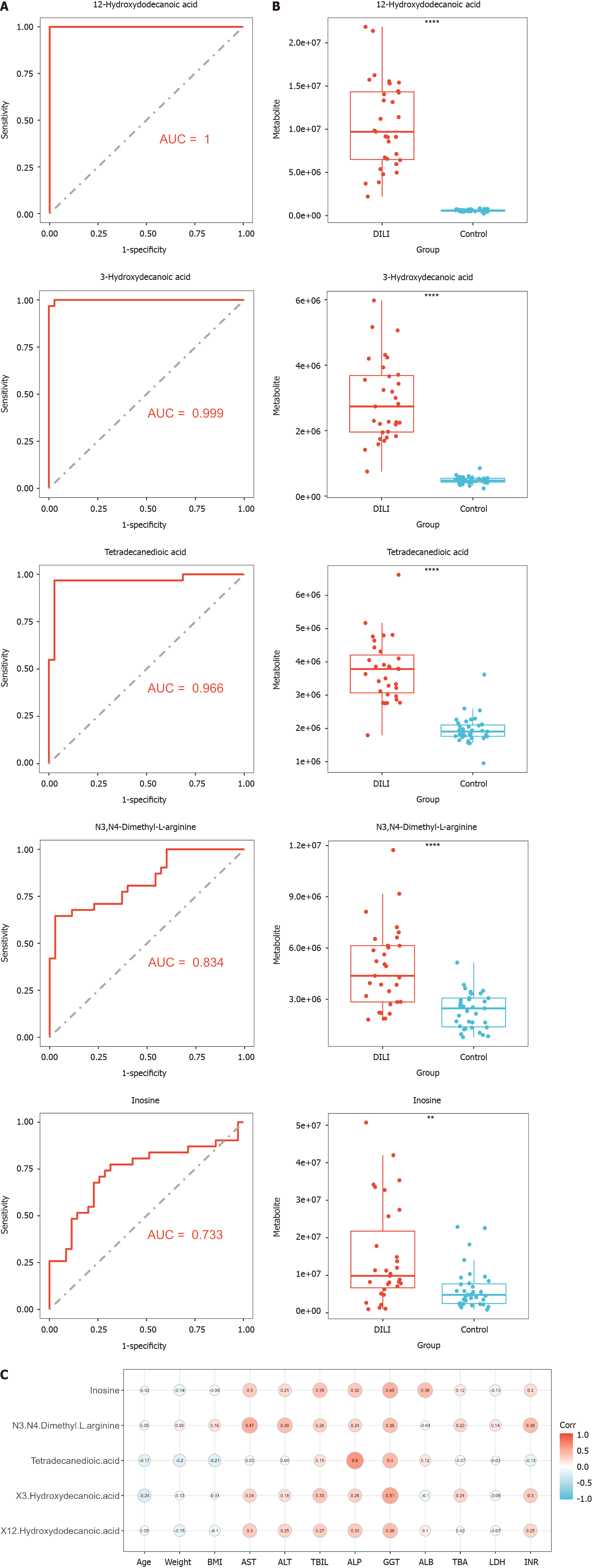
Figure 5 Diagnostic value of the hub metabolites.
A: Area under the receiver operating characteristic curve of hub metabolites; B: Expression of hub metabolites in both groups; C: Distance correlation matrix plots displaying the partial Spearman’s correlation among the five hub metabolites and 12 clinical indicators of drug-induced liver injury. AUC: Area under the curve.
- Citation: Yu SM, Zheng HC, Wang SC, Rong WY, Li P, Jing J, He TT, Li JH, Ding X, Wang RL. Salivary metabolites are promising noninvasive biomarkers of drug-induced liver injury. World J Gastroenterol 2024; 30(18): 2454-2466
- URL: https://www.wjgnet.com/1007-9327/full/v30/i18/2454.htm
- DOI: https://dx.doi.org/10.3748/wjg.v30.i18.2454