Published online Mar 8, 2026. doi: 10.37126/aige.v7.i1.114426
Revised: November 10, 2025
Accepted: December 10, 2025
Published online: March 8, 2026
Processing time: 165 Days and 20.1 Hours
Upper gastrointestinal bleeding is a medical emergency requiring prompt triage and management. Although traditionally classic risk scores such as the Rockall score, Glasgow-Blatchford score, and albumin, international normalized ratio, mental status, systolic blood pressure, age > 65 years have been employed to di
Core Tip: There are validated risk assessment scores which are being conventionally used to predict the need for endotherapy and complications in patients with acute upper gastrointestinal bleed (AUGIB). However, those have many limitations which restrict their use effectively and in a timely manner in the real world. In this era of artificial intelligence, these predictive scores can be complemented and/or replaced by machine learning analysis of clinical and laboratory data in conjunction with endoscopy images to diagnose and treat AUGIB patients accurately. In recent years, new evidence has been coming in support of this, and future research is still ongoing. In this mini-review, the use of artificial intelligence in AUGIB is briefly discussed with emphasis on current evidence, its clinical application and future directions.
- Citation: Kumar SR, Panigrahi MK, Sasmal PK. Artificial intelligence in upper gastrointestinal bleeding: Can machine learning predict endotherapy requirements? Artif Intell Gastrointest Endosc 2026; 7(1): 114426
- URL: https://www.wjgnet.com/2689-7164/full/v7/i1/114426.htm
- DOI: https://dx.doi.org/10.37126/aige.v7.i1.114426
Acute upper gastrointestinal bleeding (AUGIB) is a major cause of hospital admission and death worldwide, imposing a huge burden on healthcare services. It is based upon early and accurate risk stratification to identify those needing urgent treatment, such as endoscopic therapy. So far, clinical decision-making has relied on conventional risk stratification using scores including the Glasgow-Blatchford score (GBS), the Rockall score, and the albumin, international normalized ratio, mental status, systolic blood pressure, age > 65 years (AIMS65). AUGIB patients comprise a diverse age, gender, bleeding site, and amount of blood loss, thus accurate outcome prediction is difficult. While helpful for predicting mortality or rebleeding, their performance in anticipating the need for endoscopic hemostasis and blood transfusion, as well as mortality, is suboptimal. The need for more accurate and rapid diagnostic methods and personalised treatment strategies for specific patient populations must be explored.
Artificial intelligence (AI) and machine learning (ML) have created a paradigm shift in numerous fields of medicine. Integrated ML algorithms will improve clinicians’ ability to assess and decide on gastrointestinal bleeding soon. Recent multicentric studies show many interests in evaluating and validating AI model technologies. These technologies rely on large data sets to recognise complicated patterns and associations that typical statistical models might not see. In upper gastrointestinal bleeding (UGIB), the prospect of AI to direct triage and decide on endoscopic need in real time is es
This mini-review is a structured but non-systematic narrative description. Our focus was to apply a search strategy to find any recent (last 5 years) articles of original nature, meta-analysis and systematic review that provide quantitative or qualitative evaluation of AI and/or ML models to predict the outcome of AUGIB. We searched PubMed, Scopus and Google scholar from January 1, 2020 to July 31, 2025 using following keywords/Boolean operators: (“AI” OR “ML” OR “deep learning”) AND (“UGIB” OR “GI Bleeding” OR “peptic ulcer bleeding” OR “Non- variceal GI bleed” OR “variceal Bleed”) AND (“Risk prediction” OR “Endotherapy” OR “Rebleeding” OR “mortality” OR “triage”).
Reference Citation Analysis and backwards snowballing from relevant articles were also performed to find additional studies. Only those studies were included that evaluated AI/ML models for diagnosis, risk stratification, prediction of endoscopic therapy, rebleeding, transfusion requirement, or mortality in AUGIB. Care was taken to include those which were originally in English and human studies. Non-original papers without any additional relevance, or which did not include the UGIB part and non-English papers were excluded. Study selection and data extraction were performed independently by two reviewers, with disagreements resolved by discussion with the third author.
A qualitative appraisal was done rather than a formal, systematic scoring approach, as it was a narrative review. Studies were assessed for their heterogeneity with respect to study populations, single-centre nature and data modality. Risk of overfitting and limitations in generalizability were kept in mind while dealing with a study based on limited data and a single-centre study.
GBS focuses on vital signs, haemoglobin levels, blood urea nitrogen, and comorbidities to identify patients who may be safely discharged[2]. The Rockall score includes pre- and post-endoscopy components and predicts mortality[3]. AIMS65 evaluates albumin, international normalized ratio, mental status, systolic blood pressure, and age. While each score serves a distinct purpose, none specifically predicts the need for endoscopic hemostasis with high sensitivity.
Several studies have shown that GBS performs well in identifying low-risk patients but over-triages many who do not require urgent endoscopy. Rockall and AIMS65 have shown only moderate accuracy in stratifying patients for endoscopic intervention. Additionally, these tools are static, based on linear assumptions, and limited by the data from which they were derived - often not reflective of diverse or contemporary patient populations. The interdependency and complex interaction among clinical variables have hindered the individual application of these methods. Many clinical variables are interdependent and may interact with each other with reciprocal enhancement. These drawbacks underscore the need for more dynamic, personalised, and data-driven risk prediction strategies.
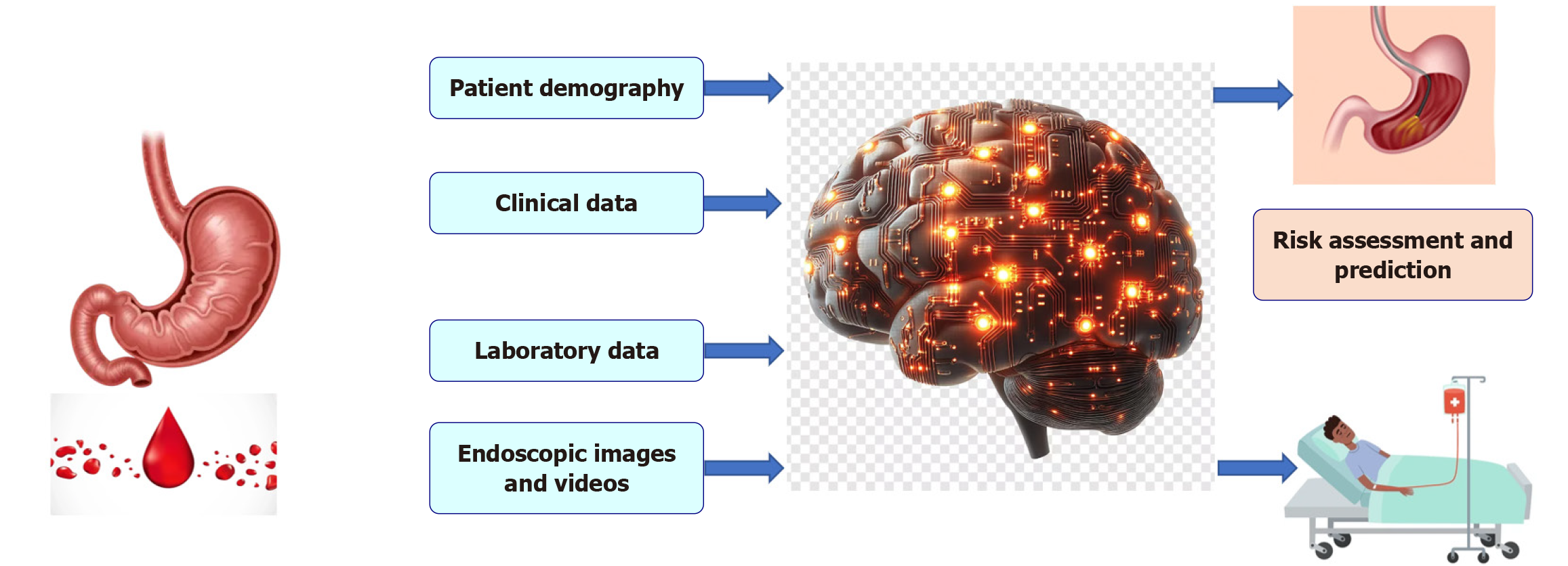
AI represents a machine-simulated form of human intelligence that enhances its capabilities through experience. It primarily operates via ML algorithms and deep learning, a specialised subset of ML that utilises complex multilayered neural networks to analyse and learn from data[4]. ML encompasses various techniques for analysing complex, non-linear relationships in large datasets. Supervised learning algorithms like logistic regression, random forests, support vector machine and gradient boosting machines have been frequently applied in UGIB predictive models (Table 1). More recently, computational models inspired by the structure and function of biological neural networks, particularly the human brain working as deep learning algorithms, including neural networks like artificial neural networks, convolutional neural networks (CNNs) and recurrent neural networks, have shown promise, particularly in image interpretation and temporal data analysis[5]. An important ML technique is linear discriminant analysis, which reduces dimensionality and classifies objects. In supervised learning, a linear combination of features is used to separate two or more data classes. These models can incorporate diverse variables, including laboratory data, clinical features, hemodynamic trends, medication history, imaging, and endoscopic findings (Figure 1). In addition, ML models learn and evolve with the appearance of fresh information, hence allowing a dynamic model that is different from the fixed rigidity of permanent scores.
| Supervised machine learning | |||
| Classification task | Comment | Regression task | Comment |
| K-nearest neighbours | Data classification as per k-nearest neighbours, non-parametric | Gradient boosting model | A combination of weaker models (e.g., a decision tree) to create a stronger prediction model |
| XGBoost | |||
| LightGBM | |||
| CatBoost | |||
| Neural network | Deep learning method composed of interconnected layers of artificial neurons | Support vector machine | Technique to categorise data points by finding an optimal hyperplane |
| ANN | |||
| CNN | |||
| Decision tree | An arranged tree in which internal nodes are attributes, branches are decisions, and leaves are outcomes or labels | Regression analysis | Useful for predicting time-to-event outcomes, including covariates and event times, such as bleeding recurrence |
The application of AI in UGIB can be conveniently discussed in three distinct phases. Pre-endoscopic prediction uses the available clinical and laboratory data to predict the urgency of endotherapy, optimising the triage and resource allocation. Endoscopic (real-time) prediction is based on an algorithm-based image and video feed analysis that detects bleeding sources and characterises stigmata (e.g., forrest classification) to predict the need for hemostasis during the procedure objectively. Lastly, post-endoscopic prediction combines procedure results with clinical information to estimate rebleeding or mortality risk, as well as direct discharge timing, follow-up, and aggressiveness of care.
AUGIB management depends on accurate risk stratification. Traditional scoring systems, such as GBS and the pre-endoscopic Rockall score, are initial estimates but lack sensitivity for predicting endoscopic intervention since they rely on static variables and pre-specified thresholds. ML algorithms have evolved with sophisticated, non-linear patient in
These AI algorithms integrate various features, including dynamic vital signs, serial laboratory values (like trending haemoglobin and lactate), comorbidities, and unstructured clinical notes, resulting in more individualised risk profiles. For example, gradient boosting is superior to GBS in predicting the need for endoscopic intervention [area under the curve (AUC) 0.94 vs 0.81], transfusion, and mortality, thereby identifying high-risk patients who would benefit most from urgent endoscopy[6]. In addition, ML can forecast post-endoscopic outcomes based on endoscopic appearances, like Forrest classification and stigmata of recent haemorrhage, along with pre-endoscopic clinical information. This fun
Despite these advances, integrating these models into clinical practice remains tricky. The “black box” quality of some advanced ML models risks eroding clinician confidence, so recourse is taken to explainable AI (xAI) methods for explaining factors in prediction[9]. Apart from that, the majority of the models available currently are developed and tested using single-centre, retrospective data with questionable generalizability. Thus, prospective multicenter validation trials are crucial to determine the role of ML in the prediction of risk and outcome analysis for AUGIB, thereby paving the way for its integration into real-time clinical decision support systems.
The most important transformation AI has brought to UGIB is its real-time application during endotherapy in predicting therapeutic needs. Neural networks like CNN and artificial neural network analyse live video feeds and images during endoscopy to provide an instantaneous second assessment to the endoscopist[10]. These deep learning models are trained on large annotated datasets to accomplish at the same time two most important tasks, i.e., firstly, appropriate detection and localisation of sources of localisation (e.g., ulcers, varices, Mallory-Weiss tears), and secondly, fine characterisation of stigmata of recent haemorrhage, such as the Forrest system. Recent studies have validated AI models, such as a random forest classifier, to accurately identify high-risk bleeding stigmata requiring hemostasis and the need for further endotherapy[11]. A study by Yen et al[12] analysed and trained AI models (MobileNetV2) on 2738 endoscopic images (analysed 2289 and tested 449) of peptic ulcer bleed and effectively achieved an AUC of 0.91 for the prediction of the need for therapy, which outperformed a junior endoscopist.
The implication of AI-aided endoscopy is profound. It can eliminate operator bias in identification and provide a well-planned endotherapy by objectively standardising it. The role in detecting subtle high-risk lesions and standardising in a bloody field cannot be overemphasised; this ultimately standardises care by ensuring that all lesions with a high probability of standardisation are flagged for the endoscopist’s attention. It significantly shortens the time to therapy and improves haemostasis rates. It limits the overall cost of patient management by decreasing hospital stay, using appropriate endotherapy modalities and avoiding unnecessary intervention.
In cirrhotic patients after an acute gastrointestinal bleed event, AI models with high accuracy can predict the predictive performance for in-hospital death. A Chinese multicentric trial developed and validated a logistic regression analysis-based ML score for cirrhosis and acute gastrointestinal bleeding, which showed good predictive performance for in-hospital mortality for cirrhosis with acute UGIB by retrospectively analysing a dataset of 1682 patients[13]. Recently, the score was prospectively analysed with even larger data of 2467 patients, and the least square SVM regression model in that study had the best predictive performance (AUC = 0.986)[14].
Randomised controlled trials have influenced clinical practices, especially in developing a prudent red blood cell transfusion algorithm in acute gastrointestinal bleeding[15]. The evidence for restrictive strategies of blood transfusion for patients with acute gastrointestinal bleeding is limited, as conventional risk scores do not accurately predict transfusion requirements. There has been considerable enhancement in predicting the requirement for blood transfusion after acute gastrointestinal bleeding in intensive care units due to advancements in ML. Using big data such as Medical Information Mart for Intensive Care and eICU Collaborative Research Database v2.0, investigators built models incorporating demographic factors, laboratory values, and clinical factors. A long short-term memory model performed more optimally than baseline regression models, with an AUC of 0.80, considering data from the first five hours of intensive care unit admission[16]. Dynamic risk prediction models provide information, promoting improved individualised patient care and clinical outcomes in the critical environment.
A multicentre study developed the Bleeding after Endoscopic Submucosal Dissection Trend from Japan score using advanced statistical modelling to predict post-endoscopic submucosal dissection bleeding in early gastric cancer, demonstrating moderate discrimination with a c-statistic of 0.70 in the external validation cohort[17]. Similarly, in a retrospective multicentre study of 1084 patients, Maruyama et al[18] demonstrated that an ML model predicted post-endoscopic submucosal dissection bleeding within 28 days for early gastric carcinoma with superior performance to a non-ML model (AUC 0.80 vs 0.71, P = 0.03) and higher sensitivity for bleed detection.
As with variceal UGIB, various studies used ML to assess rebleed, complications, and mortality prediction, comparing them with multiple risk assessment scores. A study employing multiple ML models concluded that integrating five independent risk assessment scores - GBS, AIMS65, Rockall, T-score, and Baylor Bleeding score - significantly improved the accuracy of mortality prediction in patients with non-variceal UGIB (NVUGIB)[19]. According to the training and testing datasets, the hyperparameter-tuned k-nearest neighbor classifier had the highest precision and recall, which can accurately predict mortality in patients with NVUGIB. However, the retrospective and single-centre nature of the data was found to be insufficient to validate and implement this tool in the real world yet.
A landmark 2020 study by Shung et al[6] utilised a gradient boosting ML model trained on over 1958 patients. The model was better than GBS at predicting the requirement for endoscopic intervention (area under the receiver operating characteristic curves 0.90 vs 0.87 in the external validation cohort). Discrimination and calibration remained good in a prospective validation cohort that met composite endpoints. Remarkably, incorporating temporal trends in lab results and vital signs improved predictive accuracy, illustrating the value of longitudinal data analysis. Boros et al[8] used the electronic health records (EHR) data of the Hungarian gastrointestinal bleed registry and compared ML with risk assessment scores utilising the XGBoost and categorical boosting models, attaining AUC up to 0.84 (95% confidence interval: 0.76-0.90 and 95% confidence interval: 0.77-0.90, respectively) with five-fold cross-validation. The XGBoost model had better specificity (96%) for mortality prediction compared to GBS and Rockall. Another study with a large retrospective training data set (n = 3868) with internal validation, as well as four other external validation data sets, aimed to assess Forrest classification in peptic ulcer bleed. The system achieved superior diagnostic performance with an accuracy of 91.2% and an AUC of 0.80 in validation data[20]. A few of these important and recent evidences are presented in Table 2. Other detailed analyses of various studies are shown in Table 3.
| Ref. | Primary focus | Data type | Study type | Study metrics | Key AI/ML method(s) | Comment |
| Raghareutai et al[31], 2025 | Pre-endoscopy risk stratification | Demograph, clinical and lab values, n = 1389 | Retrospective review of prospectively collected data, internal validation | AUROC 0.74 to predict endoscopic intervention (0.81 in validation test set) | linear discriminant analysis | Identify patients of AUGIB who need endoscopic intervention. Dynamic changes need multiple inputs of data |
| Shung et al[6], 2020 | Pre-endoscopy risk stratification | Clinical and lab data, n = 1958 | Retrospective, multicentre, external validation | ML vs GBS; AUROC 0.90 vs 0.87 (external validation) sensitivity 100%, specificity 26% | Gradient boosting machines (XGBoost) | The ML model could identify low-risk patients who can be safely discharged |
| He et al[20], 2024 | Real-time endoscopic prediction | Endoscopic images training data n = 3868, internal; validation data, n = 834; external validation data, n = 521 | Multicenter prospective; external validation | AUC 0.80 in the validation data set, accuracy of 91.2% | DCNN (image-based Forrest) | The DCNN system showed more accurate and stable diagnostic performance than endoscopists in the prospective clinical comparison test |
| Bai et al[14], 2025 | Post-GI bleed mortality in cirrhosis | Clinical and lab data, n = 2467 cirrhotics | Multicenter prospective, international internal validation, no external validation | AUC of 0.789 (up to 0.986 in LS-SVMR model) | LS-SVMR (least squares support vector machine regression) | CAGIB score is similar to CTP, MELD and MELD-Na for the prediction of mortality. No comparison with AIMS65, Rockall and GBS |
| Boros et al[8], 2025 | Post-GI bleed mortality | EHR registry (Hungarian GI bleed registry), n = 1021 | Retrospective, fivefold cross-validation | XGBoost and CatBoost AUC 0.79 vs 0.62 of GBS in GIB | XGBoost and CatBoost | CatBoost reached a sensitivity of 78% and a specificity of 74% |
| Ref. | Design/setting | Dataset size | Population/data type | Predictors/inputs | Algorithm/model | Comparator (if any) | Performance metrics (AUROC/sensitivity/specificity /PPV/NPV) | Validation type | Major limitations |
| Shung et al[6], 2020 | Retrospective, multicentre, external validation | 1958 patients (training + validation) | Acute UGIB (clinical + lab data) | Vital signs, Hb, BUN, comorbidities, transfusion need | Gradient boosting machine (XGBoost) | GBS, AIMS65 | AUROC 0.90 vs 0.87 (GBS); sensitivity 100%, specificity 26% | External (prospective cohort) | Limited ethnic diversity; no imaging variables |
| Raghareutai et al[31], 2025 | Retrospective review of a prospectively collected dataset | 1389 cases | Non-variceal UGIB; demographic, clinical + lab data | Age, vitals, Hb, BUN, SRH | Linear discriminant analysis | Rockall, GBS | AUROC 0.74 (0.81 in validation test set) | Internal + temporal validation | Small sample; limited external testing; static variables |
| Nazarian et al[11], 2024 | Multicentre retrospective cohort | 970 patients | AUGIB: Need for hemostatic therapy | Clinical + lab + endoscopic features | Random forest classifier | GBS, Rockall | AUROC 0.84 vs 0.72 in GBS | Five-fold cross-validation | Retrospective bias; heterogeneous image quality |
| He et al[20], 2024 | Multicenter prospective; external validation | 3868 images from 1200 patients | Peptic ulcer bleed; Forrest classification | Endoscopic images | Deep CNN | Endoscopist | Accuracy 91.2%; AUC 0.80 in validation data set | External validation | Limited to image data; no clinical integration |
| Yen et al[12], 2021 | Retrospective single-centre image analysis | 2738 images (2289 train, 449 test) | Peptic ulcer bleed | Endoscopic image features | MobileNetV2 (CNN) | Human endoscopist | AUROC 0.91 vs 0.80 (human); sensitivity 94%; specificity 92% (3 class category) | Internal (hold-out) | Retrospective design; limited generalisability |
| Boros et al[8], 2025 | Retrospective EHR registry (Hungarian GI bleed registry) | 1021 records | GI bleed (mixed aetiology) | Demographics, labs, vitals, comorbidities | XGBoost/CatBoost models | GBS, Rockall | AUROC 0.79 in GIBleed, AUC 0.84 in mortality | 5-fold cross-validation | Retrospective; national registry bias |
| Bai et al[14], 2025 | Multicenter prospective; international; internal | 2467 cirrhotic patients with UGIB | Cirrhosis with acute UGIB | Clinical + lab parameters | LS-SVMR (least-squares SVM regression) | Logistic regression model | AUROC 0.986 | External (prospective) | Limited to the cirrhotic population |
| Levi et al[16], 2021 | Retrospective ICU database (MIMIC-III + eICU-CRD) | Approximately 6000 ICU admissions | GI bleeding patients in the ICU | Demographic + vital + lab series | LSTM neural network | Logistic regression | AUROC > 0.80 in the MIMIC-III data set | Cross-dataset validation | Limited prospective validation; EHR data noise |
Included studies had methodological heterogeneity in design, validation method and AI used for analysis. Sample sizes were varied (n = 449 to 3868), with most of the data extracted from the EHR or the data registry. Endoscopic image analysis was also done by curated still images, which differ from real-world video feeds, limiting its generalisability. These create an inherent problem in the qualitative appraisal of the AI system due to selection bias and information bias. With this limitation, model overfitting was a concern mostly in studies which used high-dimensional but limited frameworks, such as CNN and gradient boosting ensembles. Importantly, most of the studies lacked external or pro
One important criticism of AI is the “black-box” nature of complex algorithms involving layered and variable data sets. Clinicians are often reluctant to adopt models they cannot intuitively understand and/or control effectively. Research on xAI has become increasingly important in recent years, focusing on sophisticated and difficult-to-understand ML algorithms. Attempts at filling this gap have been through interpretability tools such as SHapley Additive exPlanations (SHAP), which applies an additive model in approximating how input features relate to their respective coefficients and assigning relevant values to each feature based on their contribution to model predictions[21]. An xAI tool called local interpretable model-agnostic explanations describes individual ML model predictions by approximating them with a simple, local model. The feature and area analysis provides insight into the significance of features and areas within the image, which is useful when analysing endoscopic images[22]. Ustaalioğlu and Ak[23] used SHAP to visualise the output attributes in assessing the role of ML in predicting mortality in NVUGIB, helping clinicians to determine the important variables for gastrointestinal bleed, such as haemoglobin, blood urea nitrogen, systolic blood pressure and age. Likewise, Boros et al[8] employed explainable risk assessment models using the SHAP method, where each feature’s SHAP value, positive indicating increased and negative indicating decreased predicted mortality risk, is visualised on a summary plot. However, there were only a few studies which addressed the clinical usability and acceptability of the interpretable AI. None formally measured the impact of xAI visualisations on diagnostic confidence or workflow time. It highlights an important research gap, as their practical integration into endoscopic decision-making has not yet matched the increasing use of various explainability methods. These resources boost physicians’ self-assurance and internal decision-making transparency, which is a crucial precondition for AI’s moral and responsible application in healthcare.
Effective ML model deployment in real-world settings needs seamless integration with EHR systems. It includes real-time data extraction and model input automation. The common application of electronic patient records within the healthcare sector has facilitated the integration of AI models into such systems. Effective use of scores for risk stratification depends on accurately identifying the patient groups for which these models are applicable, a process known as phenotyping. Traditionally, this has involved manual chart and/or clinical codes reviews. However, AI and ML can automate phe
For clinical translation, an operational framework constitutes three functionalities; first is data pipeline requirements, which ensures automated input of structured (vitals, labs) and unstructured (clinical notes, endoscopy reports) data with standardised nomenclatures such as SNOMED. Secondly, latency performance is an important part which assesses and minimises the delay between real-time observation and digital interpretation. It targets sub-second prediction rendering for real-time endoscopy assistance to avoid workflow latency. Lastly, validation and regulatory compliance mandate multicentre prospective validation and adherence to regulatory classifications such as Food and Drug Administration “software as a medical device” or Conformite Europeenne-marked clinical decision support tools[25].
From a technical perspective, local data pre-processing, on-device model inference, and edge computing may reduce dependency on network connectivity during live endoscopy. Such architecture ensures model latency below 500 milliseconds - crucial for real-time AI overlays in endoscopic video feeds[26,27]. However, these advantages should be proven by large-scale, multicentre prospective trials. Also, systematic data entry with AI analysis in mind, using SNOMED or natural language processing, will facilitate prediction, faster with easy-to-use interfaces with risk scores, confidence intervals and suggestions.
AI in healthcare must address algorithmic bias, data privacy, and equity issues. ML models trained on data from specific populations may not generalise well, leading to disparate outcomes. Having diverse training datasets to generalise and validate AI models is essential before coming to standard practices. Data feed should undergo periodic audit for performance across subgroups (e.g., age, sex, race). Nonetheless, the revolutionary potential of AI comes with challenges of data privacy, consent and governance[28]. National and international frameworks are necessary for a deeper understanding of its ethical implications and unintended consequences. These ethical dimensions are technical challenges and crucial determinants of patient dependability and standardisation.
Research must strive for multicenter verification on a large scale in many health systems in order to improve AI adoption for UGIB therapy with adequate proof. The objective should be to develop hybrid models that incorporate appropriate ML and clinical judgment. Thorough history, clinical examinations, laboratory data, and AI can better predict endoscopy requirements, transfusion needs, post-endoscopy complication management, and clinical care. When AI is used in managing these patients compared to standard care, unnecessary endoscopic interventions can be avoided, optimising overall hospital stay and cost effectiveness. Clinicians must receive optimising educational initiatives to use AI tools effectively and safely to improve patient outcomes[29]. Regulatory guidance from medical societies and health authorities is essential to standardise the development and evaluation of AI-based tools. Subsequent studies should standardise the integration of AI systems into today’s endoscopy platforms and electronic medical records, creating training modules in the use of AI tools for clinicians and supporting new AI tools[30].
ML is now a revolutionary force in stratifying and managing UGIB. Across multiple independent studies, ML-based models have demonstrated higher discriminative accuracy than conventional risk scores such as GBS, Rockall, or AIMS65, particularly for predicting endoscopic therapy, transfusion requirement, and short-term outcomes. However, these advantages should be interpreted with caution. The majority of available models derive from single-centre or retrospective cohorts and are prone to overfitting and limited generalisability. Few have undergone multicentre or real-time validation, and standardised reporting of calibration, interpretability, and clinical utility remains inconsistent. The future of UGIB care lies in a synergistic model where AI augments but does not replace clinical acumen, ensuring safe and personalised patient care.
| 1. | Nigam GB, Murphy MF, Travis SPL, Stanley AJ. Machine learning in the assessment and management of acute gastrointestinal bleeding. BMJ Med. 2024;3:e000699. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 2. | Blatchford O, Murray WR, Blatchford M. A risk score to predict need for treatment for upper-gastrointestinal haemorrhage. Lancet. 2000;356:1318-1321. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 907] [Cited by in RCA: 716] [Article Influence: 27.5] [Reference Citation Analysis (8)] |
| 3. | Rockall TA, Logan RF, Devlin HB, Northfield TC. Risk assessment after acute upper gastrointestinal haemorrhage. Gut. 1996;38:316-321. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1109] [Cited by in RCA: 929] [Article Influence: 31.0] [Reference Citation Analysis (7)] |
| 4. | Taye MM. Understanding of Machine Learning with Deep Learning: Architectures, Workflow, Applications and Future Directions. Computers. 2023;12:91. [DOI] [Full Text] |
| 5. | Kufel J, Bargieł-Łączek K, Kocot S, Koźlik M, Bartnikowska W, Janik M, Czogalik Ł, Dudek P, Magiera M, Lis A, Paszkiewicz I, Nawrat Z, Cebula M, Gruszczyńska K. What Is Machine Learning, Artificial Neural Networks and Deep Learning?-Examples of Practical Applications in Medicine. Diagnostics (Basel). 2023;13:2582. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 306] [Cited by in RCA: 159] [Article Influence: 53.0] [Reference Citation Analysis (3)] |
| 6. | Shung DL, Au B, Taylor RA, Tay JK, Laursen SB, Stanley AJ, Dalton HR, Ngu J, Schultz M, Laine L. Validation of a Machine Learning Model That Outperforms Clinical Risk Scoring Systems for Upper Gastrointestinal Bleeding. Gastroenterology. 2020;158:160-167. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 188] [Cited by in RCA: 160] [Article Influence: 26.7] [Reference Citation Analysis (7)] |
| 7. | Zheng NS, Keloth VK, You K, Kats D, Li DK, Deshpande O, Sachar H, Xu H, Laine L, Shung DL. Detection of Gastrointestinal Bleeding With Large Language Models to Aid Quality Improvement and Appropriate Reimbursement. Gastroenterology. 2025;168:111-120.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 12] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 8. | Boros E, Pintér J, Molontay R, Prószéky KG, Vörhendi N, Simon OA, Teutsch B, Pálinkás D, Frim L, Tari E, Gagyi EB, Szabó I, Hágendorn R, Vincze Á, Izbéki F, Abonyi-Tóth Z, Szentesi A, Vass V, Hegyi P, Erőss B. New machine-learning models outperform conventional risk assessment tools in Gastrointestinal bleeding. Sci Rep. 2025;15:6371. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (6)] |
| 9. | Basubrin O. Current Status and Future of Artificial Intelligence in Medicine. Cureus. 2025;17:e77561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (1)] |
| 10. | Choi J, Shin K, Jung J, Bae HJ, Kim DH, Byeon JS, Kim N. Convolutional Neural Network Technology in Endoscopic Imaging: Artificial Intelligence for Endoscopy. Clin Endosc. 2020;53:117-126. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 39] [Article Influence: 6.5] [Reference Citation Analysis (5)] |
| 11. | Nazarian S, Lo FPW, Qiu J, Patel N, Lo B, Ayaru L. Development and validation of machine learning models to predict the need for haemostatic therapy in acute upper gastrointestinal bleeding. Ther Adv Gastrointest Endosc. 2024;17:26317745241246899. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 5] [Reference Citation Analysis (1)] |
| 12. | Yen HH, Wu PY, Su PY, Yang CW, Chen YY, Chen MF, Lin WC, Tsai CL, Lin KP. Performance Comparison of the Deep Learning and the Human Endoscopist for Bleeding Peptic Ulcer Disease. J Med Biol Eng. 2021;41:504-513. [RCA] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (2)] |
| 13. | Bai Z, Li B, Lin S, Liu B, Li Y, Zhu Q, Wu Y, Yang Y, Tang S, Meng F, Chen Y, Yuan S, Shao L, Qi X. Development and Validation of CAGIB Score for Evaluating the Prognosis of Cirrhosis with Acute Gastrointestinal Bleeding: A Retrospective Multicenter Study. Adv Ther. 2019;36:3211-3220. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 26] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 14. | Bai Z, Lin S, Sun M, Yuan S, Marcondes MB, Ma D, Zhu Q, Li Y, He Y, Philips CA, Liu X, Pinyopornpanish K, Shao L, Méndez-Sánchez N, Basaranoglu M, Wu Y, Chen Y, Yang L, Mancuso A, Tacke F, Li B, Liu L, Ji F, Qi X. Machine learning based CAGIB score predicts in-hospital mortality of cirrhotic patients with acute gastrointestinal bleeding. NPJ Digit Med. 2025;8:489. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (2)] |
| 15. | Villanueva C, Colomo A, Bosch A, Concepción M, Hernandez-Gea V, Aracil C, Graupera I, Poca M, Alvarez-Urturi C, Gordillo J, Guarner-Argente C, Santaló M, Muñiz E, Guarner C. Transfusion strategies for acute upper gastrointestinal bleeding. N Engl J Med. 2013;368:11-21. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1449] [Cited by in RCA: 1089] [Article Influence: 83.8] [Reference Citation Analysis (5)] |
| 16. | Levi R, Carli F, Arévalo AR, Altinel Y, Stein DJ, Naldini MM, Grassi F, Zanoni A, Finkelstein S, Vieira SM, Sousa J, Barbieri R, Celi LA. Artificial intelligence-based prediction of transfusion in the intensive care unit in patients with gastrointestinal bleeding. BMJ Health Care Inform. 2021;28:e100245. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 34] [Article Influence: 6.8] [Reference Citation Analysis (1)] |
| 17. | Hatta W, Tsuji Y, Yoshio T, Kakushima N, Hoteya S, Doyama H, Nagami Y, Hikichi T, Kobayashi M, Morita Y, Sumiyoshi T, Iguchi M, Tomida H, Inoue T, Koike T, Mikami T, Hasatani K, Nishikawa J, Matsumura T, Nebiki H, Nakamatsu D, Ohnita K, Suzuki H, Ueyama H, Hayashi Y, Sugimoto M, Yamaguchi S, Michida T, Yada T, Asahina Y, Narasaka T, Kuribasyashi S, Kiyotoki S, Mabe K, Nakamura T, Nakaya N, Fujishiro M, Masamune A. Prediction model of bleeding after endoscopic submucosal dissection for early gastric cancer: BEST-J score. Gut. 2021;70:476-484. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 134] [Cited by in RCA: 123] [Article Influence: 24.6] [Reference Citation Analysis (0)] |
| 18. | Maruyama H, Takahashi K, Kojima K, Nakajima N, Sato H, Mizuno KI, Sugitani S, Terai S. Machine-Learning Prediction of Bleeding After Endoscopic Submucosal Dissection for Early Gastric Cancer: A Multicenter Study. JGH Open. 2025;9:e70203. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 19. | Ungureanu BS, Gheonea DI, Florescu DN, Iordache S, Cazacu SM, Iovanescu VF, Rogoveanu I, Turcu-Stiolica A. Predicting mortality in patients with nonvariceal upper gastrointestinal bleeding using machine-learning. Front Med (Lausanne). 2023;10:1134835. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 9] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 20. | He XJ, Wang XL, Su TK, Yao LJ, Zheng J, Wen XD, Xu QW, Huang QR, Chen LB, Chen CX, Lin HF, Chen YQ, Hu YX, Zhang KH, Jiang CS, Liu G, Li DZ, Li DL, Wen W. Artificial intelligence-assisted system for the assessment of Forrest classification of peptic ulcer bleeding: a multicenter diagnostic study. Endoscopy. 2024;56:334-342. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 10] [Article Influence: 5.0] [Reference Citation Analysis (2)] |
| 21. | Ponce-Bobadilla AV, Schmitt V, Maier CS, Mensing S, Stodtmann S. Practical guide to SHAP analysis: Explaining supervised machine learning model predictions in drug development. Clin Transl Sci. 2024;17:e70056. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 496] [Cited by in RCA: 281] [Article Influence: 140.5] [Reference Citation Analysis (1)] |
| 22. | Hassan SU, Abdulkadir SJ, Zahid MSM, Al-Selwi SM. Local interpretable model-agnostic explanation approach for medical imaging analysis: A systematic literature review. Comput Biol Med. 2025;185:109569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 38] [Article Influence: 38.0] [Reference Citation Analysis (2)] |
| 23. | Ustaalioğlu İ, Ak R. Predicting Mortality in Non-Variceal Upper Gastrointestinal Bleeding: Machine Learning Models Versus Conventional Clinical Risk Scores. J Clin Med. 2025;14:7425. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 24. | Shung D, Tsay C, Laine L, Chang D, Li F, Thomas P, Partridge C, Simonov M, Hsiao A, Tay JK, Taylor A. Early identification of patients with acute gastrointestinal bleeding using natural language processing and decision rules. J Gastroenterol Hepatol. 2021;36:1590-1597. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 18] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 25. | Joshi G, Jain A, Araveeti SR, Adhikari S, Garg H, Bhandari M. FDA-Approved Artificial Intelligence and Machine Learning (AI/ML)-Enabled Medical Devices: An Updated Landscape. Electronics. 2024;13:498. [DOI] [Full Text] |
| 26. | Gong EJ, Bang CS. Edge Artificial Intelligence Device in Real-Time Endoscopy for the Classification of Colonic Neoplasms. Diagnostics (Basel). 2025;15:1478. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 4] [Reference Citation Analysis (1)] |
| 27. | Gong EJ, Woo J, Lee JJ, Bang CS. Role of artificial intelligence in gastric diseases. World J Gastroenterol. 2025;31:111327. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 4] [Reference Citation Analysis (1)] |
| 28. | Kiseleva A, Kotzinos D, De Hert P. Transparency of AI in Healthcare as a Multilayered System of Accountabilities: Between Legal Requirements and Technical Limitations. Front Artif Intell. 2022;5:879603. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 89] [Article Influence: 22.3] [Reference Citation Analysis (1)] |
| 29. | Schubert T, Oosterlinck T, Stevens RD, Maxwell PH, van der Schaar M. AI education for clinicians. EClinicalMedicine. 2025;79:102968. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 40] [Reference Citation Analysis (1)] |
| 30. | Alagappan M, Brown JRG, Mori Y, Berzin TM. Artificial intelligence in gastrointestinal endoscopy: The future is almost here. World J Gastrointest Endosc. 2018;10:239-249. [PubMed] [DOI] [Full Text] |
| 31. | Raghareutai K, Tanchotsrinon W, Sattayalertyanyong O, Kaosombatwattana U. Development and validation of a machine learning model to predict hemostatic intervention in patients with acute upper gastrointestinal bleeding. BMC Med Inform Decis Mak. 2025;25:145. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/