Copyright: ©Author(s) 2026.
World J Orthop. Mar 18, 2026; 17(3): 115770
Published online Mar 18, 2026. doi: 10.5312/wjo.v17.i3.115770
Published online Mar 18, 2026. doi: 10.5312/wjo.v17.i3.115770
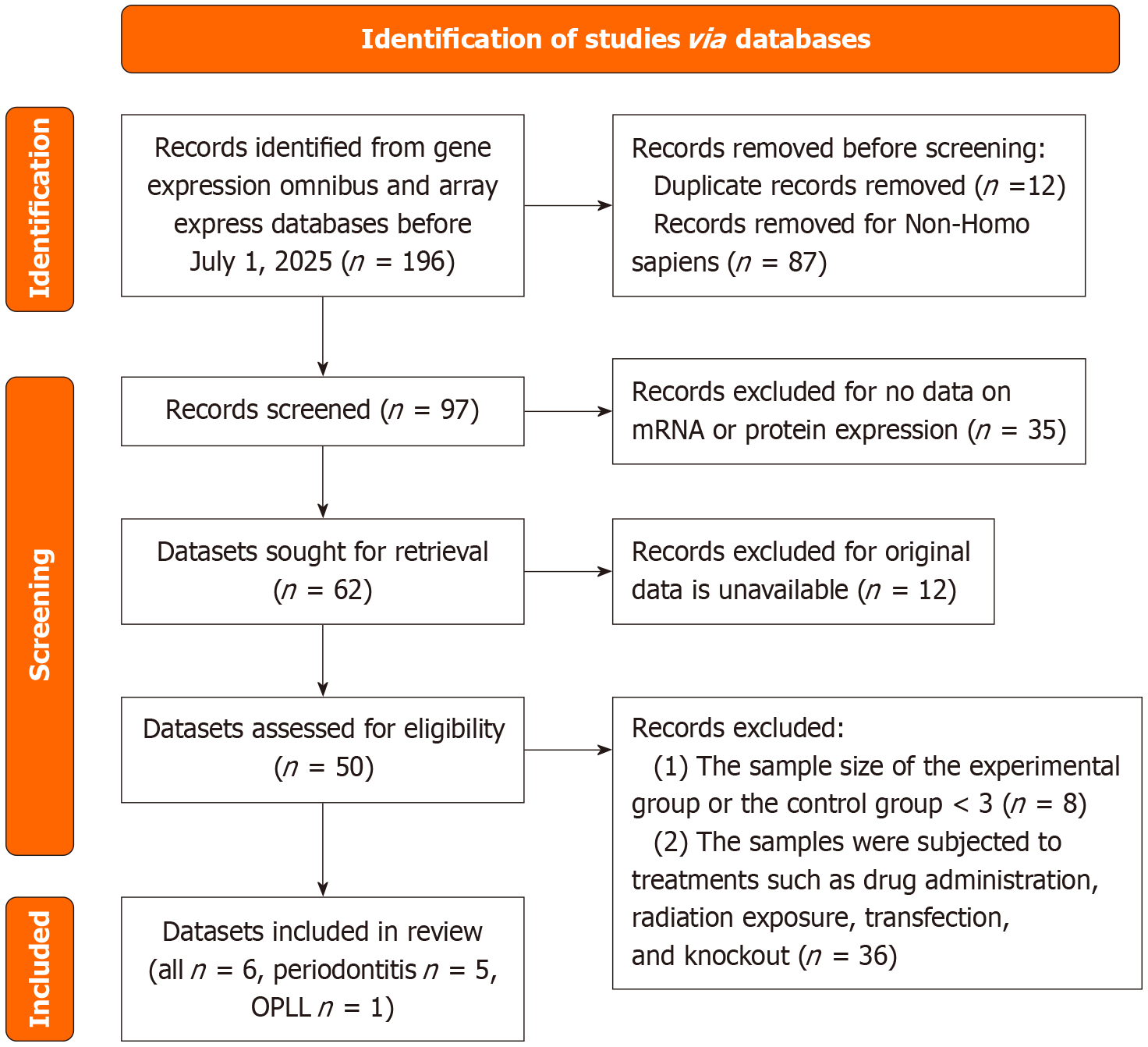
Figure 1 PRISMA flow diagram for dataset identification and selection.
The diagram illustrates the systematic process of dataset identification, screening, eligibility assessment, and final inclusion in the integrated analysis, following PRISMA guidelines. OPLL: Ossification of the posterior longitudinal ligament.
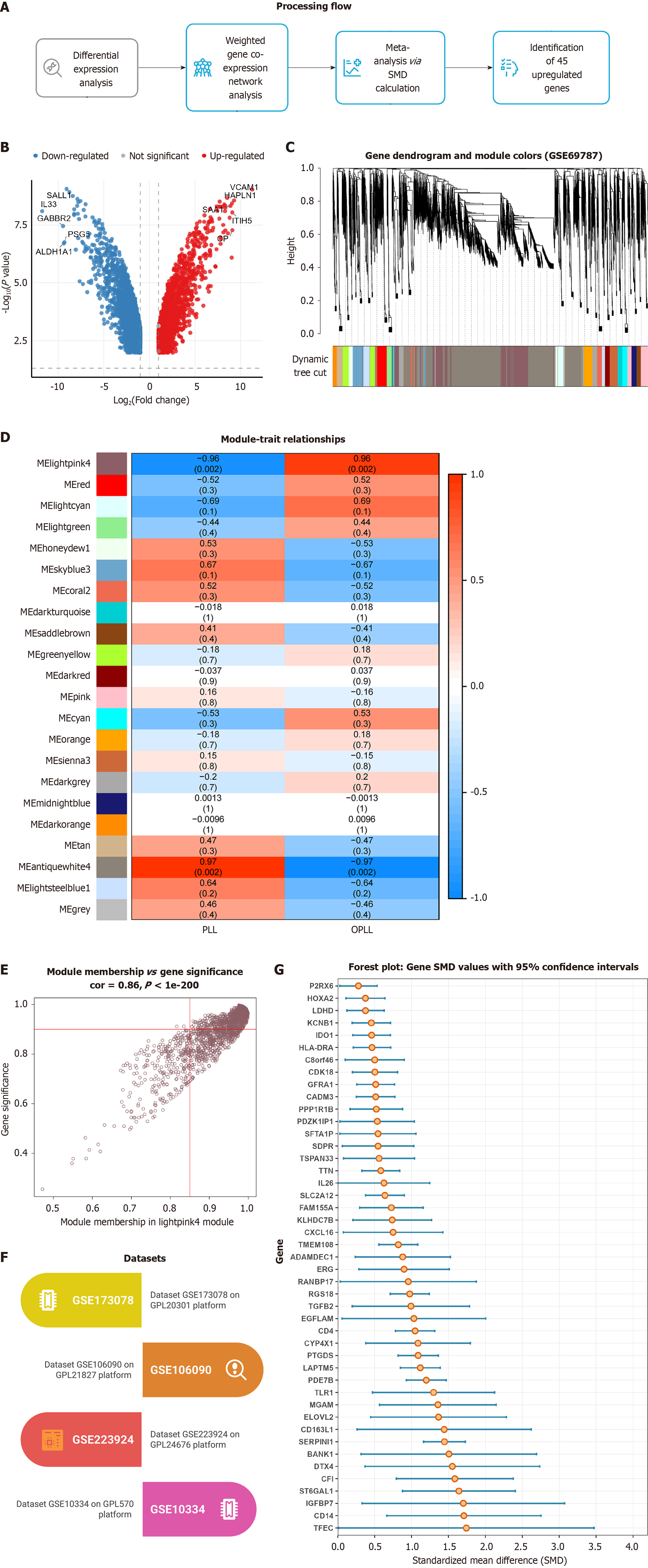
Figure 2 Identification of shared core genes between ossification of the posterior longitudinal ligament and periodontitis via weighted gene co-expression network analysis and meta-analysis.
A: Schematic workflow of the study; B: Differential gene expression analysis in ossification of the posterior longitudinal ligament (OPLL); C-E: Identification of key modules by weighted gene co-expression network analysis; Gene clustering dendrogram (C); Module-trait relationships showing the “lightpink4” module with the strongest association to OPLL (D); Characteristic scatter plot of the “lightpink4” module (E); F: Information of the periodontitis transcriptomic datasets; G: Standardized mean difference analysis of 45 co-expressed genes demonstrating consistent differential expression in periodontitis. SMD: Standardized mean difference.
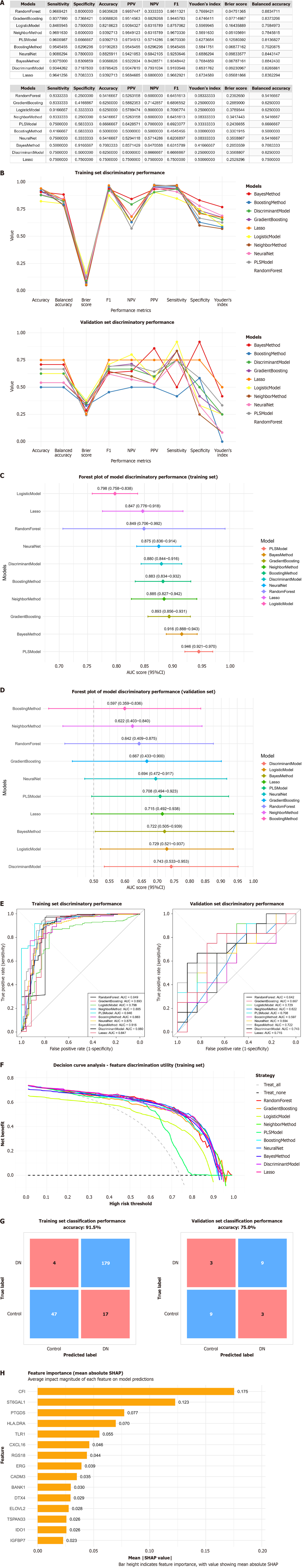
Figure 3 Feature variable selection via integrated machine learning algorithms and SHapley Additive exPlanations explainability analysis for disease discrimination.
A: Summary of performance metrics (e.g., accuracy, sensitivity, specificity) for the 10 machine learning models (the training set is on the top and the validation set is on the bottom); B: Line charts depicting model performance across different evaluation dimensions; C and D: Forest plot displaying area under the curve values and their 95% confidence intervals for all models; E: Receiver operating characteristic curves illustrating the predictive performance of the models; F: Decision curve analysis evaluating the clinical utility of the optimal model; G: Confusion matrix visualization for the selected model; H: SHapley Additive exPlanations analysis identifying complement factor I as the top contributing feature in the optimal model.
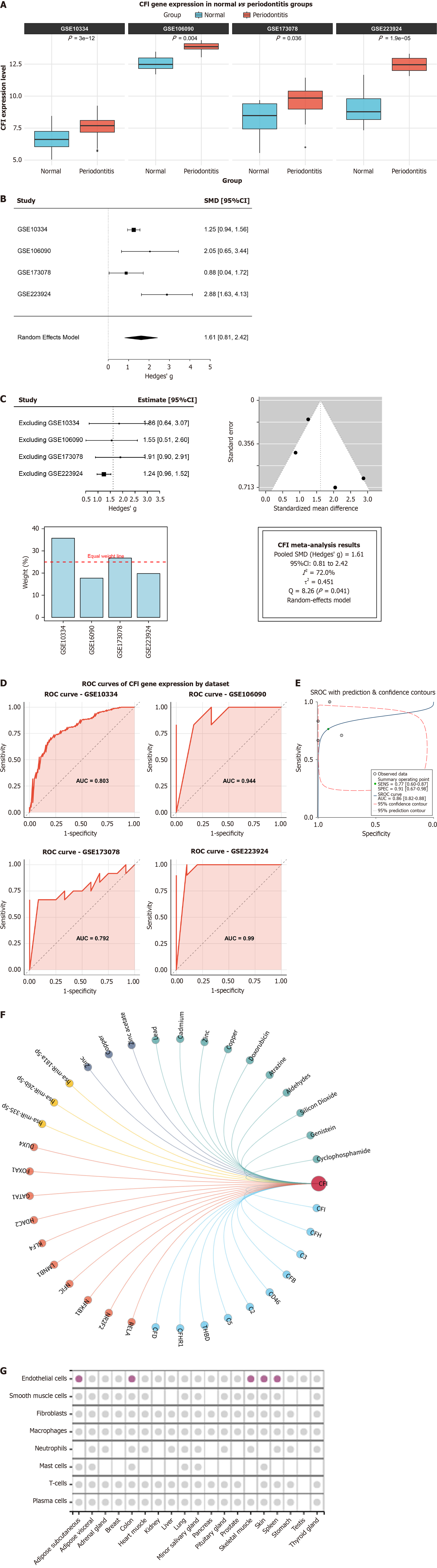
Figure 4 Exploration of clinical significance and biological mechanisms of complement factor I.
A: Validation of complement factor I (CFI) upregulation in four independent periodontitis datasets; B and C: The analysis results indicate a large pooled effect size, with a Hedges’ g value of 1.61 (95%CI: 0.81-2.42). Heterogeneity tests indicated substantial heterogeneity among the studies (I2 = 72.0%, τ2 = 0.451). The leave-one-out sensitivity analysis confirmed the robustness of this pooled result. No significant publication bias was detected by Egger’s linear regression test (P = 0.217); D: Receiver operating characteristic (ROC) curve evaluating the discriminatory capability of CFI for periodontitis; E: Summary ROC (sROC) curve with a composite area under the curve of 0.86; F: CFI-centered multi-level regulatory network showing interactions with miRNAs, transcription factors, lncRNAs, and compounds; G: Human protein atlas validation of CFI protein expression localized primarily in endothelial cells.
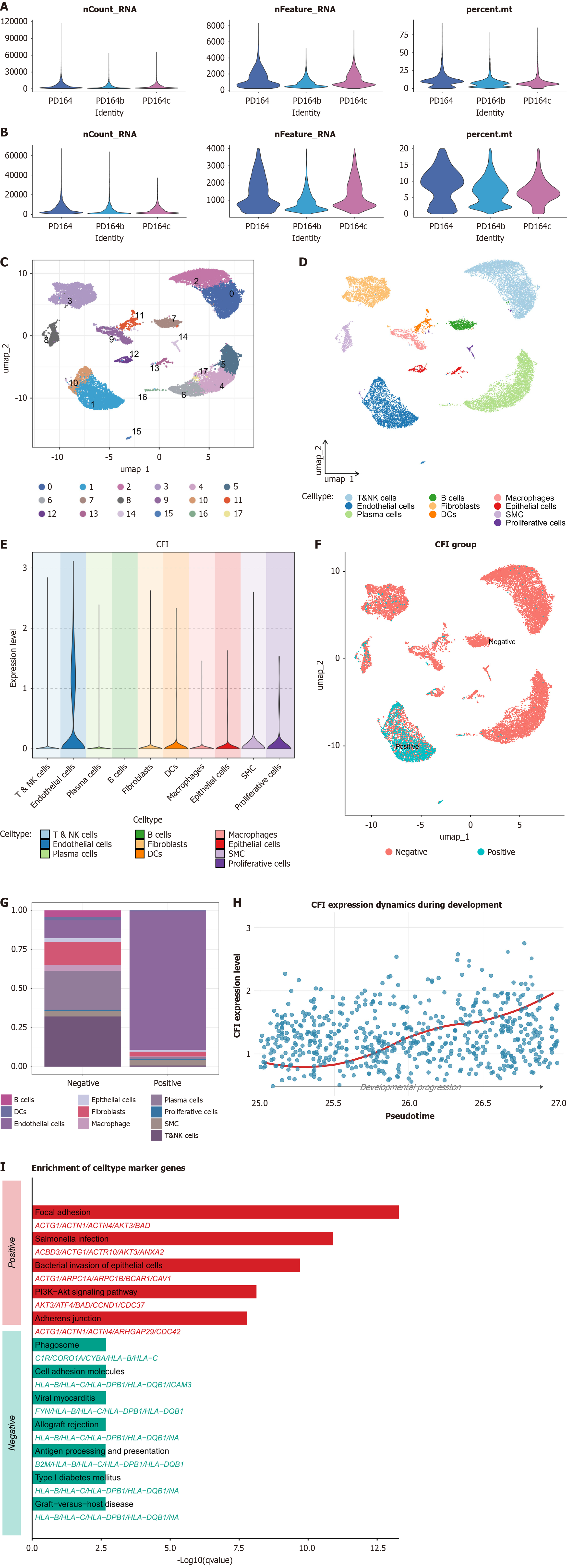
Figure 5 Single-cell characterization of complement factor I expression and functional enrichment in periodontitis.
A and B: Quality control of single cell data; C-E: Expression distribution of complement factor I (CFI) across different cell types; F and G: Identification of the primary cellular source of CFI; F: Proportion of each cell subset in CFI-positive and CFI-negative groups; G: Comparative analysis demonstrating endothelial cells as the dominant source of CFI expression; H: Cell trajectory analysis showing progressive increase of CFI expression during endothelial cell differentiation; I: Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis of differentially expressed genes in CFI-positive vs CFI-negative groups, highlighting significant enrichment in pathways related to cell adhesion, bacterial infection, and intracellular signaling.
- Citation: Man YN, Zhong LY, Wen YL, He ML. Machine learning identifies complement factor I as a shared mediator of periodontitis and ossification of posterior longitudinal ligament. World J Orthop 2026; 17(3): 115770
- URL: https://www.wjgnet.com/2218-5836/full/v17/i3/115770.htm
- DOI: https://dx.doi.org/10.5312/wjo.v17.i3.115770