©Author(s) (or their employer(s)) 2026.
World J Gastrointest Surg. Feb 27, 2026; 18(2): 114951
Published online Feb 27, 2026. doi: 10.4240/wjgs.v18.i2.114951
Published online Feb 27, 2026. doi: 10.4240/wjgs.v18.i2.114951
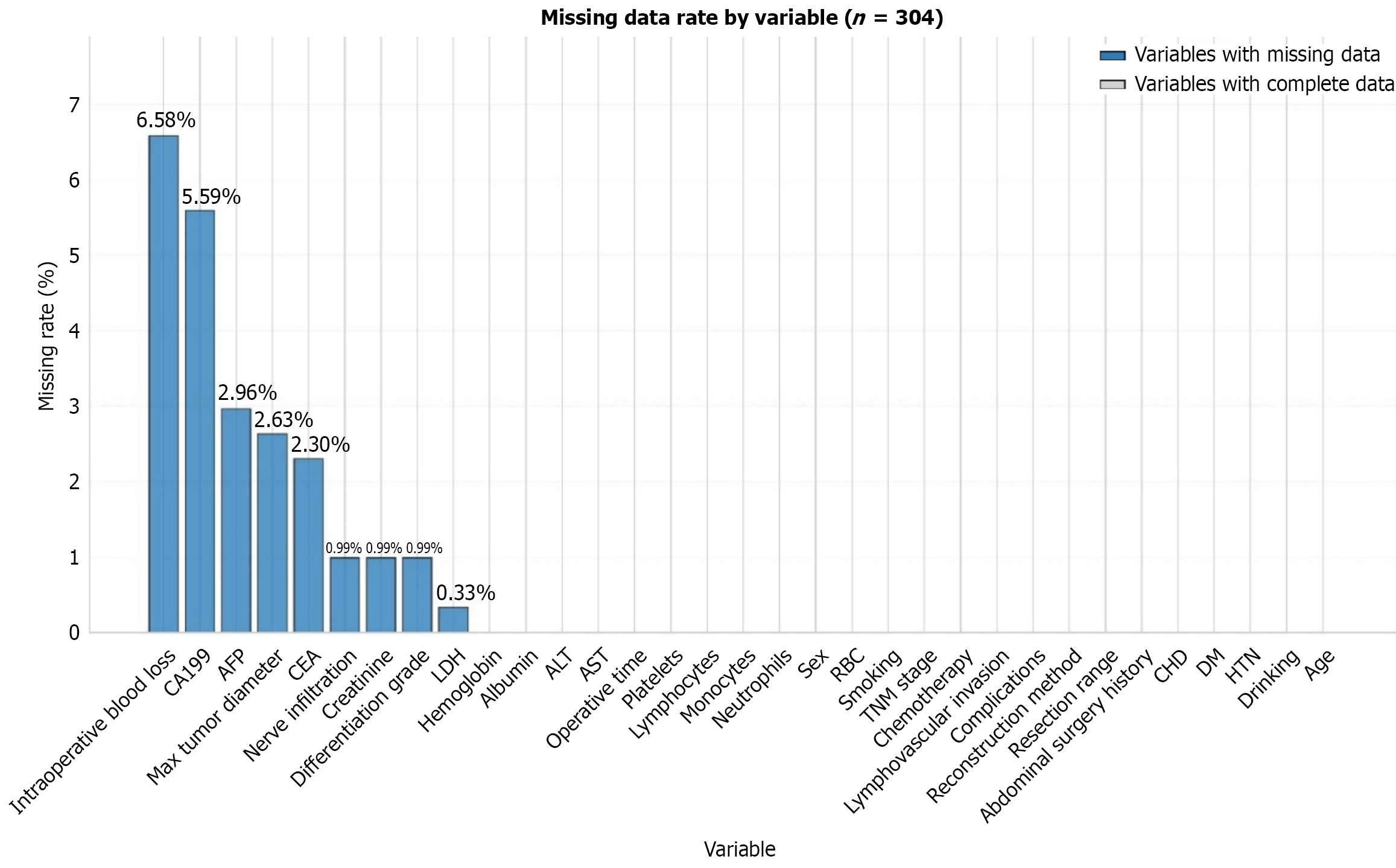
Figure 1 Schematic diagram of the missing dataset situation.
CA199: Carbohydrate antigen 19-9; AFP: Alpha-fetoprotein; CEA: Carcinoembryonic antigen; LDH: Lactate dehydrogenase; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; RBC: Red blood cell count; CHD: Coronary heart disease; DM: Diabetes mellitus; HTN Hypertension.
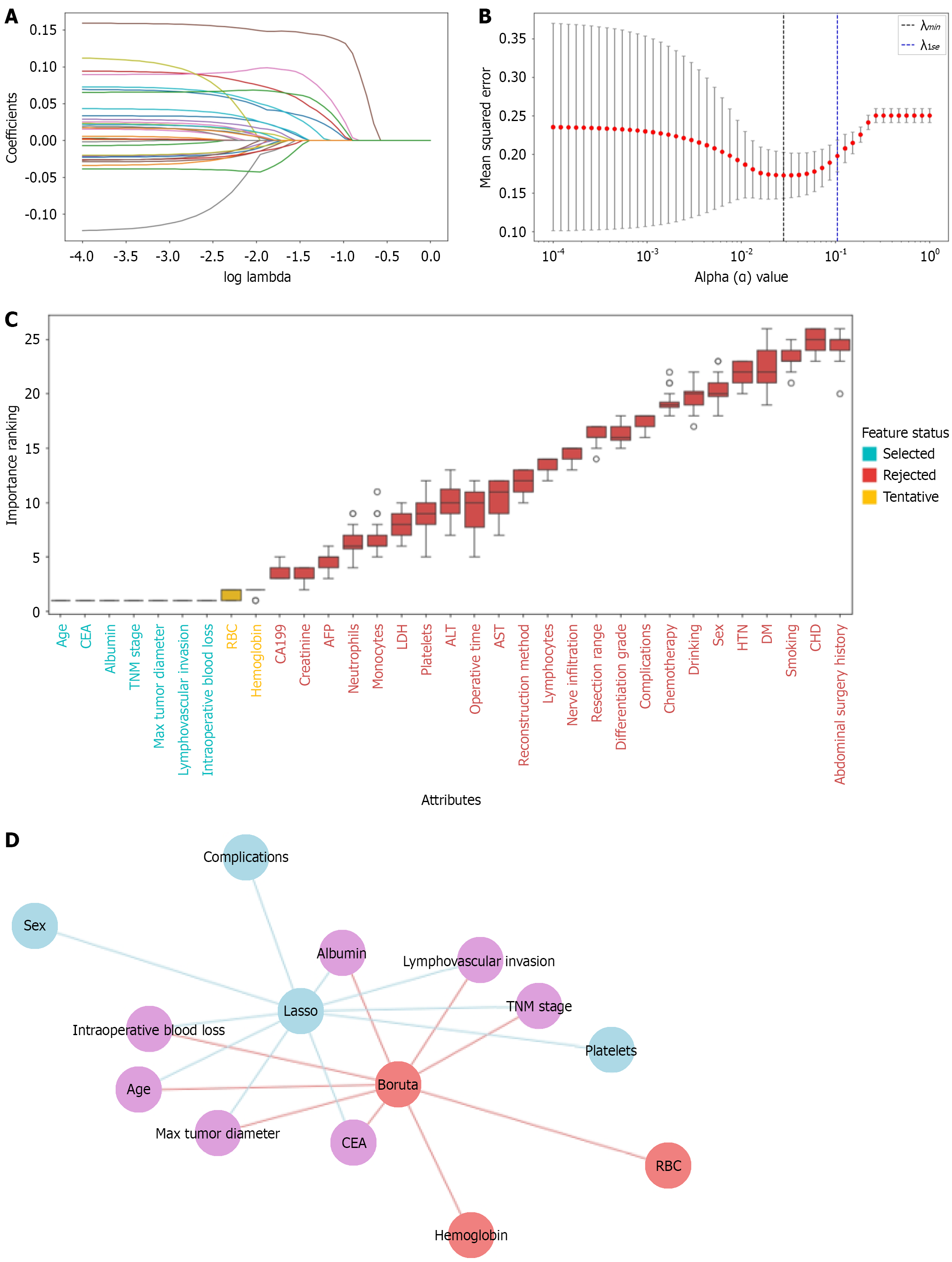
Figure 2 Feature selection.
A: Least absolute shrinkage and selection operator coefficient path; B: Least absolute shrinkage and selection operator regularization path diagram; C: Boruta algorithm; D: Final feature subset. CEA: Carcinoembryonic antigen; TNM: Tumor-node-metastasis; CA199: Carbohydrate antigen 19-9; AFP: Alpha-fetoprotein; LDH: Lactate dehydrogenase; ALT: Alanine aminotransferase; AST: Aspartate aminotransferase; HTN: Hypertension; DM: Diabetes mellitus; CHD: Coronary heart disease; RBC: Red blood cell count.
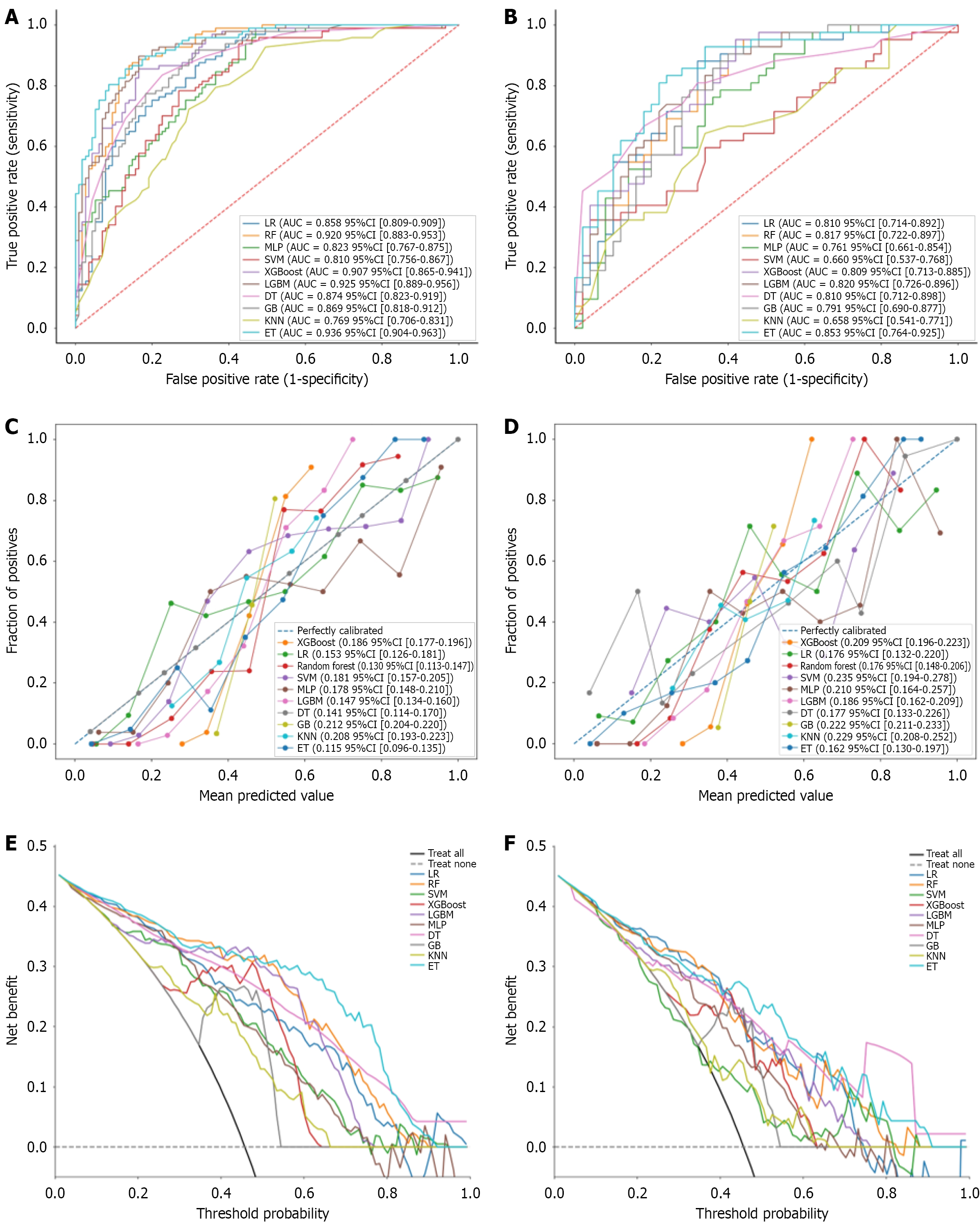
Figure 3 Comparative analysis of the performance of the ten machine learning models.
A: Receiver operating characteristic curves for the training set; B: Receiver operating characteristic curves for the validation set; C: Calibration curves for the training set; D: Calibration curves for the validation set; E: Decision curve analysis for the training set; F: Decision curve analysis for the validation set. LR: Logistic regression; AUC: Area under the curve; CI: Confidence interval; RF: Random forest; MLP: Multi-layer perceptron; SVM: Support vector machine; XGBoost: Extreme gradient boosting; LGBM: Light gradient boosting machine; DT: Decision tree; GB: Gradient boosting; KNN: K-nearest neighbors; ET: Extremely randomized trees.
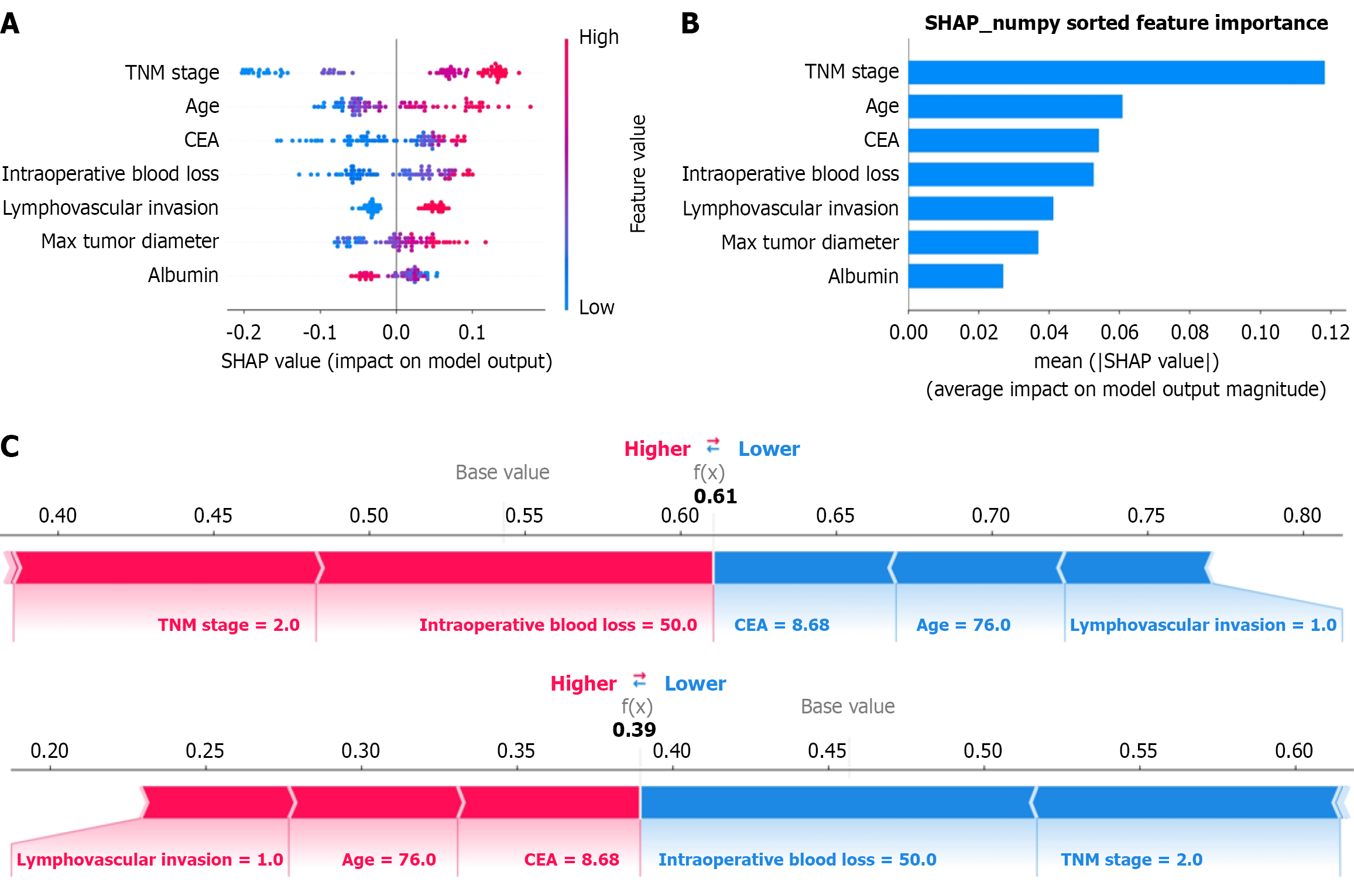
Figure 4 SHapley Additive exPlanations analysis for the optimal extra trees model.
A: Beeswarm plot summarizing feature impacts across the dataset; B: Ranking of feature importance based on mean absolute SHapley Additive exPlanations values for the extremely randomized trees model; C: Force plot illustrating the explanation for an individual prediction. TNM: Tumor-node-metastasis; CEA: Carcinoembryonic antigen; SHAP: SHapley Additive exPlanations.
- Citation: Lü YN, Liu D, Tao S, Wu J, Yu SJ, Yuan HL. Development of a machine learning-based model for predicting postoperative survival in gastric cancer. World J Gastrointest Surg 2026; 18(2): 114951
- URL: https://www.wjgnet.com/1948-9366/full/v18/i2/114951.htm
- DOI: https://dx.doi.org/10.4240/wjgs.v18.i2.114951