Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115437
Revised: December 3, 2025
Accepted: January 21, 2026
Published online: April 15, 2026
Processing time: 180 Days and 2.5 Hours
Diabetic nephropathy (DN) is a major microvascular complication of diabetes, and progressive renal fibrosis is the central feature that drives long-term kidney function decline. Epigenetic regulation is increasingly recognized in metabolic diseases, but the contribution of the histone acetyltransferase EP300 to diabetic renal fibrosis remains unclear. Nephrocystin-4 (NPHP4), a cilia-related gene, has been implicated in tubular injury and fibrotic changes, yet its regulation and role in DN have not been defined.
To determine whether EP300 promotes renal fibrosis in DN by activating NPHP4 through H3K27 acetylation and to evaluate the functional consequences of tar
Transcriptomic analysis of human diabetic kidney samples was performed using RNA sequencing, with EP300 expression patterns validated by real-time quan
Our results revealed that EP300 was significantly upregulated in the kidneys of DN patients, and it transcriptionally activated NPHP4 by enriching H3K27ac at its promoter, as demonstrated by chromatin immunoprecipitation-RT-qPCR. In high glucose-stimulated HK-2 cells, knockdown of either EP300 or NPHP4 effectively attenuated fibrosis marker expression (alpha-smooth muscle actin and fibronectin) and restored the balance between apoptosis and proliferation. In a DN mouse model, pharmacological inhibition of EP300 (C646) or AAV-mediated knockdown of NPHP4 substantially improved renal function (reduced serum creatinine, blood urea nitrogen, and proteinuria) and attenuated renal fibrosis, with combination therapy showing the most pronounced benefits. Immunohistochemical and immunofluorescence analyses further validated that the EP300–NPHP4 axis critically regulates renal fibrotic progression.
In conclusion, our study identifies the EP300/H3K27ac/NPHP4 axis as a novel epigenetic pathway driving renal fibrosis in DN. These findings provide mechanistic insight into the diabetic renal fibrotic process and highlight the therapeutic potential of targeting EP300 or NPHP4 in the treatment of DN.
Core Tip: Renal fibrosis is a key pathological process driving diabetic nephropathy progression, yet its epigenetic regulation remains unclear. This study identifies EP300 as a histone acetyltransferase that activates nephrocystin-4 transcription through Histone H3 lysine 27 acetylation, promoting renal fibrosis. Inhibition of the EP300-nephrocystin-4 axis alleviates fibrotic injury and improves renal function in diabetic mice, providing new mechanistic insight and a potential therapeutic target for diabetic nephropathy.
- Citation: Si W, Dai Y, Hu GP, Zhang Q, Lv F, Zhang Q. EP300 drives renal fibrosis in diabetic nephropathy via histone acetyltransferase-mediated nephrocystin-4 expression. World J Diabetes 2026; 17(4): 115437
- URL: https://www.wjgnet.com/1948-9358/full/v17/i4/115437.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i4.115437
Diabetic nephropathy (DN) is one of the most serious microvascular complications of diabetes mellitus and a leading cause of end-stage renal disease worldwide[1]. Epidemiologically, it affects approximately 20%-40% of diabetic patients, contributing significantly to the global healthcare burden and necessitating renal replacement therapy in a substantial proportion of cases[1]. The central feature and terminal pathological outcome is progressive renal fibrosis[2]. Despite the increasing clinical burden of DN, there is still a paucity of specific treatments for renal fibrosis, and existing therapies are ineffective in slowing disease progression.
The pathogenesis of DN is complex and multifactorial, initiated by persistent hyperglycemia and encompassing a web of interconnected pathways. These include metabolic disturbances (e.g., advanced glycation end product accumulation, polyol pathway flux), hemodynamic changes (e.g., activation of the renin-angiotensin-aldosterone system), oxidative stress, and chronic inflammation[2]. These initial insults converge to activate profibrotic signaling cascades, most notably the transforming growth factor-beta pathway, which promotes renal cell transdifferentiation and excessive extracellular matrix deposition, ultimately manifesting as glomerulosclerosis and tubulointerstitial fibrosis.
In recent years, the critical role of epigenetic modifications in metabolic disease pathogenesis has become a hot research topic[3-5]. As a reversible gene expression regulation, histone acetylation is involved in pathological processes, such as disorders of glucose and lipid metabolism, insulin resistance and target organ damage by dynamically altering chromatin structure and transcription factor activity[6-8].
EP300 is an important histone acetyltransferase (HAT) that activates the downstream gene transcription by catalyzing histone H3 lysine 27 lysine acetylation (H3K27ac)[9]. H3K27ac is an active chromatin mark involved in several biological processes, such as cell proliferation, differentiation, and fibrosis[10,11]. Aberrant EP300 expression has been found to promote organ damage by regulating inflammation- and fibrosis-related pathways in metabolic diseases[12,13]. While the role of EP300 in hepatic fibrosis is regulated by the AMPK signaling pathway[14] and EP300 inhibition can alleviate pulmonary fibrosis progression[15], few studies have reported on its role in renal fibrosis, and its specific regulatory mechanisms remain unclear.
The nephrocystin-4 (NPHP4) gene is a key regulator of cilia structure and function. Mutations in NPHP4 have been associated with various renal tubular diseases and fibrotic lesions[16,17]. Recently, it was discovered that NPHP4 is involved not only in cilia assembly and signaling but also in fibrosis progression by affecting apoptosis and proliferation[18]. However, NPHP4 expression in DN and its association with epigenetic modifications remain unclear. Given EP300’s acetyltransferase activity, investigating whether it drives DN fibrosis by regulating NPHP4 expression could provide new insights into the molecular mechanisms of DN.
Therefore, we aimed to reveal the epigenetic regulatory mechanism of the EP300-NPHP4 axis in DN and provide a theoretical basis for the development of targeted intervention strategies. By integrating transcriptomic analyses with functional experiments, elucidate how EP300 promotes fibrosis marker expression and renal function impairment by regulating NPHP4 expression, providing a new target for the precision treatment of DN.
This study includes peripheral blood samples from 15 patients with DN from the Inpatient Department at Hefei First People’s Hospital and 12 healthy controls for RNA-seq data. Data quality was assessed using FastQC (v0.11.9). After removing low-quality sequences with Trimmomatic (v0.39), the RNA-seq data were compared with the GRCh38 reference genome using HISAT2 (v2.2.1). Then, featureCounts (v2.0.3) was used to calculate gene expression. DESeq2 (1.44.0) screened for differentially expressed genes (DEGs) (|log2foldchange| > 1 and false discovery rate < 0.05), and candidate genes were obtained by intersecting with 40 histone acetylation-related genes reported in the literature[19]. The characterized genes were screened further using least absolute shrinkage and selection operator regression (glmnet package) with the optimal penalty parameter (λ) determined by 10-fold cross-validation based on the minimum mean cross-validated error. The support vector machine-recursive feature elimination algorithm (caret package) was implemented with a linear kernel, and the optimal feature subset was selected based on the highest cross-validated accuracy. GSEA analysis (MSigDB database) revealed functional pathways of differentially expressed genes. The regulatory mechanism of EP300 was investigated using chromatin immunoprecipitation (ChIP)-seq data obtained from the ENCODE database. The binding site was identified using MACS2 (v2.2.7.1) with a q-value of < 0.01. The peak was associated with the transcription start site (TSS) region using the ChIPseeker package (v1.36.0). Intersection with the set of fibrosis-related genes identified putative target genes. HOMER (v4.11) predicted the binding motif. All analyses were performed in R (v4.2.2). Statistical tests were performed using the Wilcoxon rank-sum test (P < 0.05). Visualization was performed using several tools to ensure reproducible results, including ggplot2, pheatmap, and IGV (v2.12.3).
To model the direct metabolic insult of diabetes on kidney cells in a controlled setting, a high glucose (HG) stimulation model was employed using the HK-2 human renal proximal tubular epithelial cell line. This in vitro approach isolates the effect of hyperglycemia, a central driver of DN, allowing for the investigation of specific signaling pathways and epigenetic changes in renal epithelial cells, which are key players in the initiation of tubulointerstitial fibrosis.
HK-2 cells were purchased from Cellverse Co., Ltd. (Shanghai, China) and cultured in DMEM (Gibco, United States) supplemented with 10% fetal bovine serum (FBS) (Gibco, United States) and 1% penicillin-streptomycin (Gibco, United States) at 5% CO2 and 37 °C. To induce an in vitro model of DN, cells were treated with 30 mmol/L D-glucose for 48 hours. HK-2 cells cultured in DMEM containing 10% FBS under normal glucose conditions were used as controls.
Transfection experiments for small interfering RNA targeting EP300 (siEP300), small interfering RNA targeting NPHP4 (siNPHP4), short hairpin RNA targeting NPHP4 (shNPHP4), and overexpression of NPHP4 (OE-NPHP4) were performed using Lipofectamine 3000 Transfection Reagent (Invitrogen, United States) according to the manufacturer’s instructions. HK-2 were seeded at a density of 2 × 105 cells per well in 6-well plates and cultured to 70%-80% density in a 37 °C, 5% CO2 incubator using high glucose Dulbecco’s modified Eagle’s medium (DMEM, Gibco, United States) supplemented with 10% FBS (Gibco, United States) and 1% penicillin-streptomycin (Gibco, United States). For siEP300 and siNPHP4 transfection, 100 pmol siRNA targeting the gene of interest (SignaGen, United States) or scrambled siRNA (negative control, Sigma-Aldrich, United States) was diluted in 250 µL serum-free DMEM, mixed with 3 µL Lipo
Sequences are as follows: (1) The siEP300: GUACCAGAGCUACUGCUAATT; (2) The siNPHP4: GGAUCAAGU
After the transfection complex was added to cells, they were incubated at 37 °C for 8 hours, followed by replacement with fresh complete DMEM medium. After 48 hours, cells were collected for downstream analysis including real-time quantitative polymerase chain reaction (RT-qPCR), western blot and functional assays. EP300 and NPHP4 expression levels were detected by RT-qPCR to confirm transfection efficiency.
Cells were cross-linked with 1% formaldehyde for 10 minutes at room temperature, and then the reaction was terminated with glycine and washed with PBS. Cells were lysed using SDS lysate containing protease inhibitors, and chromatin was fragmented by sonication (Bioruptor sonicator, 30 seconds on/30 seconds off, 10 cycles) to obtain DNA fragments of 200-500 bp. The fragment size was verified by agarose gel electrophoresis. For ChIP experiments, 100 μg of chromatin fra
Primer sequences: (1) NPHP4-F: 5’-ACCTACCGGCACTTCTTTGG-3’, NPHP4-R: 5’-CACAGCCACGATATGAGGG
To recapitulate the systemic metabolic disturbances and progressive renal pathology of type 2 DN in vivo, a well-established mouse model combining high-fat diet (HFD) feeding and low-dose streptozotocin (STZ) injection was utilized. HFD induces insulin resistance and metabolic syndrome, while STZ causes partial pancreatic β-cell damage, together generating stable hyperglycemia, dyslipidemia, and the gradual development of renal functional and structural abnormalities characteristic of human DN. This model bridges our in vitro findings to a pathophysiologically relevant whole-organism context.
The 4-6-week-old male C57BL/6 mice were purchased from Jiangsu GemPharmatech Co., Ltd (Jiangsu Province, China). After 1 week of acclimatization, they were randomly divided into six groups: (1) Control group; (2) HFD/STZ + vehicle group; (3) HFD/STZ + C646 group; (4) HFD/STZ + AAV-NC group; (5) HFD/STZ + AAV-shNPHP4 group; and (6) HFD/STZ + AAV-shNPHP4 + C646 group. All mice were sacrificed at the end of week 8.
Control group: Fed with normal diet.
HFD/STZ + vehicle group: Mice were fed with 60% HFD (research diets, No. D12492) for 4 weeks, followed by STZ (50 mg/kg) injection for 5 consecutive days. Starting from day 1 of week 6, mice received daily intraperitoneal injections of equal volume solvent for 2 weeks.
HFD/STZ+C646 group: Same scheme as HFD/STZ + vehicle group. From day 1 of week 6, mice received daily intraperitoneal injections of 20 nmol/g C646 (equivalent to approximately 5.4 mg/kg/day) for 2 weeks. This dosage was selected based on its established efficacy in inhibiting EP300 and ameliorating fibrosis in metabolic disease models. Specifically, a comparable dose of C646 (10 mg/kg) has been shown to effectively attenuate inflammation and fibrosis in a mouse model of non-alcoholic steatohepatitis, supporting the translational relevance of this dosage range for targeting EP300 in DN.
HFD/STZ + AAV-NC group: Same scheme as HFD/STZ + vehicle group. On the last day of week 5, tail vein injection of empty AAV (5 × 10¹¹ vg/mouse) was performed. Mice were sacrificed at the end of week 8.
HFD/STZ + AAV-shNPHP4 group: Same modeling as HFD/STZ + vehicle group. On the last day of week 5, mice received a tail vein injection of target gene-knockdown AAV (5 × 1011 vg/mouse).
HFD/STZ + AAV-shNPHP4 + C646 group: Same modeling as HFD/STZ + vehicle group. On the last day of week 5, mice received a tail vein injection of target gene-knockdown AAV (5 × 1011 vg/mouse). Starting at day 1 of week 6, mice received daily intraperitoneal injections of 20 nmol/g C646 for 2 weeks. All animal experiments were approved by the Institutional Animal Care and Use Committee of Hefei First People’s Hospital and conducted in accordance with the National Institutes of Health guidelines for the care and use of laboratory animals.
HK-2 cell proliferation was measured using a cell counting kit-8 (CCK-8) (Biomedicals, United States). Cells were seeded in 96-well plates at a density of 2 × 104 cells/mL per well. Prior to detection, 10 µL CCK-8 reagent was added to each well, followed by incubation for 2 hours in a cell culture incubator. The optical density was measured at 450 nm using a microplate reader. All experiments were performed with six replicates to ensure reproducibility.
Cells were grown on coverslips and fixed with 4% paraformaldehyde for 15 minutes (room temperature), then washed 3 times with PBS. For DN mouse kidney tissues, paraffin sections were deparaffinized to water treatment. Subsequently, permeabilization was performed with 0.1% Triton X-100 (ebiogo, China). Cell samples were processed for 10 minutes at room temperature and tissue samples were processed for 20 minutes, followed by three washes with PBS. The TUNEL reaction was performed, the TUNEL mix (enzyme marker solution: Fluorescent marker solution = 1:9) was prepared according to the manufacturer’s instructions (ebiogo, China), 50 μL of covered samples was added dropwise, incubated at 37 °C in the dark (60 minutes for cells and 90 minutes for tissues), and washed 3 times with PBS. Finally, nuclei were stained with DAPI (ebiogo, China) for 5 minutes, rinsed with PBS and sealed with an antifade mounting medium (Beyotime, China), and observed by fluorescence microscope (Thermo Fisher Scientific, United States).
Kidney tissue sections were washed in PBS and processed using the Dual-Labeled Triple-Color Multiplex Immunofluorescence Kit Plus (China Antibiotics, China). Sections were permeabilized with 0.3% Triton X-100 (ebiogo, China), subjected to antigen retrieval (RuiBu Biologicals, China), and blocked with 5% bovine serum albumin (BSA). Primary antibodies against alpha-smooth muscle actin (α-SMA; Proteintech, 80008-1-RR, 1:500) and fibronectin (Proteintech, 15613-1-AP, 1:300) antibodies were incubated overnight at 4 °C. Alexa Fluor-conjugated secondary antibodies (Zhongshan Jinqiao, ZF-0311) were incubated for 1 hour in the dark, followed by nuclear counterstaining with DAPI. Sections were mounted using an antifade mounting medium (Beyotime, China) and imaged with a confocal micrcoscope (Zeiss LSM 900).
Total RNA was extracted from renal tissue and cells using TRIzol reagent (Invitrogen, United States) and reverse transcribed to cDNA using SweScript RT kit (Servicebio, United States). RT-qPCR was performed using ArtiCanATM SYBR qPCR Mix (Servicebio) on a QuantReady K9600 real-time PCR system (Hangzhou Suizhen Biotechnology Co., Ltd.). Gene expression levels were normalized using GAPDH as an internal reference and calculated using the 2-ΔΔCt method.
Primer sequences were as follows: (1) EP300-F: 5’-AGCCAAGCGGCCTAAACTC-3’, EP300-R: 5’-TCACCACCATTGGTTAGTCCC-3’; (2) NPHP4-F: 5’-ACCTACCGGCACTTCTTTGG-3’, NPHP4-R: 5’-CACAGCCACGATATGAGGGTG-3’; and (3) GAPDH-F: 5’-GGAGCGAGATCCCTCCAAAAT-3’, GAPDH-R: 5’-GGCTGTTGTCATACTTCTCATGG-3’.
Protein extracts were prepared from HG-stimulated HK-2 cells for in vitro analyses or from homogenized mouse kidney tissues for in vivo analyses using radioimmunoprecipitation assay (RIPA) lysis buffer supplemented with a protease inhibitor cocktail (Solarbio, China). Proteins were separated by SDS-PAGE and transferred to polyvinyl difluoride (PVDF) membranes (Merck Millipore). Membranes were blocked with 5% non-fat milk and incubated overnight at 4 °C with primary antibodies against EP300, H3K27ac, NPHP4, α-SMA, fibronectin, and GAPDH. After washing, membranes were incubated with horseradish peroxidase-conjugated secondary antibodies at room temperature for 1 hour. Protein bands were visualized using the SH-Advance523 Multi-functional Ultrasensitive Imaging System and quantified by grayscale analysis with ImageJ software.
Primary antibodies: (1) EP300 (Cell Signaling Technology, 54062S, 1:1000, validated for use in renal cells[1]); (2) H3K27ac (Proteintech, 39085, 1:1000, validated in models of diabetic kidney disease[2]); (3) NPHP4 (Proteintech, 13812-1-AP, 1: 600); (4) α-SMA (Proteintech, 80008-1-RR, 1:10000); (5) fibronectin (Proteintech, 15613-1-AP, 1:2000); and (6) GAPDH (Proteintech, 60004-1-Ig, 1:10000).
Twenty-four-hour urine samples were collected for measurement of urinary protein levels using an enzyme-linked immunosorbent assay (ELISA; Cusabio Biotech, China). Serum samples were analyzed for blood urea nitrogen (BUN) using the urease method (Nanjing Jiancheng, C013-2-1, China) and for serum creatinine using the sarcosine oxidase method (Biosharp, BL890B, China). All assays were performed according to the manufacturers’ instructions.
Mouse kidney tissues were collected, fixed and embedded for paraffin sectioning. Tissue sections were dried on a slide warmer, deparaffinized in xylene, rehydrated through graded ethanol solutions, and rinsed with distilled water. Sections were then stained using a hematoxylin and eosin (H&E) staining kit (ebiogo, China), mounted with neutral balsam, and examined and imaged using a light microscope (OLYMPUS, Japan).
Mouse kidney tissues were paraffin-embedded and sectioned. Sections were baked on a slide warmer, deparaffinized in xylene, rehydrated through graded ethanol solutions, and rinsed with distilled water. Nuclei were stained with Weigert’s iron hematoxylin for 5-10 minutes, differentiated in acid alcohol, and blued in running water. Cytoplasm was stained with ponceau red for 5-10 minutes, followed by differentiation with phosphomolybdic acid. Collagen fibers were then stained with aniline blue for 3-5 minutes. Sections were then dehydrated through graded ethanol, cleared in xylene, mounted with neutral balsam, and examined using a light microscope (OLYMPUS, Japan) for image acquisition.
Kidney tissue samples were paraffin-embedded and cut into 4 μm sections on slides. After deparaffinizing with xylene, samples were rehydrated through graded alcohols. Antigen retrieval was performed using citrate buffer. Sections were treated with 3% hydrogen peroxide to block endogenous peroxidase activity. After serum blocking, anti-Ki67 antibody (1:300; Fuzhou Maixin Biotech Co., Ltd.) and NPHP4 (Proteintech, 1:50) were added and incubated overnight at 4 °C. Horseradish peroxidase-conjugated secondary antibody was incubated at room temperature for 1 hour, followed by DAB development for 1-3 minutes. After counterstaining with hematoxylin, sections were mounted and observed under a light microscope (OLYMPUS, Japan) for image acquisition.
All experiments were performed in triplicate unless otherwise specified. Data are presented as mean ± SD. Statistical significance was determined using a Student’s t-test or one-way analysis of variance (ANOVA) with Tukey’s post hoc test, as appropriate, in GraphPad Prism 9 software (GraphPad Software, United States). P < 0.05 was considered statistically significant.
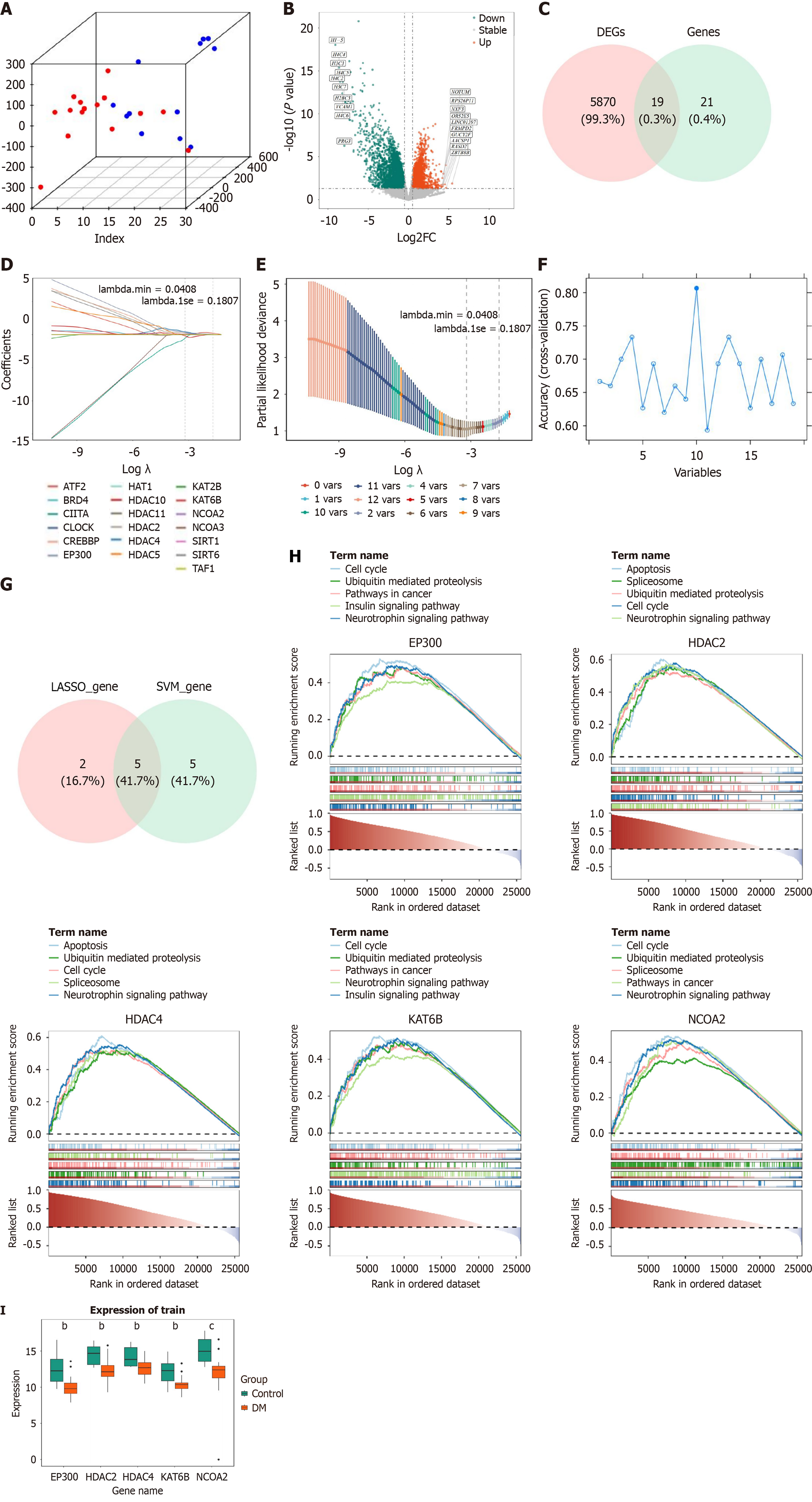
In this study, mRNA sequencing data from blood samples of 15 patients with DN and 12 healthy controls were analyzed. Principal component analysis demonstrated clear separation between control and DN samples, indicating high data quality and good reproducibility (Figure 1A). Differential expression analysis identified 5889 differentially expressed genes (DEGs), including 1977 upregulated and 3912 downregulated genes in the DN group compared with controls. To identify histone acetylation-related candidates, these DEGs were intersected with 40 previously reported histone acetylation-related genes, yielding 19 candidate genes (Figure 1B and C).
Subsequently, least absolute shrinkage and selection operator (LASSO) logistic regression identified seven feature genes, including CIITA, EP300, HDAC10, and HDAC2 (Figure 1D and E). Integration of these results with the support vector machine-recursive feature elimination (SVM-RFE) algorithm (Figure 1F) yielded 10 overlapping candidate genes (Figure 1G), from which five key genes were ultimately defined: EP300, HDAC2, HDAC4, KAT6B, and NCOA2. Gene set enrichment analysis (GSEA) revealed that EP300 waqs significantly enriched in 105 pathways, including insulin signaling and cell cycle regulation (Figure 1G), suggesting a potential role in disease progression via epigenetic mechanisms. Wilcoxon rank-sum testing further confirmed that these five key genes, including EP300, were differentially expressed in the DN group compared with controls (Figure 1H and I). Notably, dysregulation of EP300 – a histone acetyltransferase (HAT) – may contribute to renal fibrogenesis by modulating profibrotic gene expression, highlighting its potential as a therapeutic target.
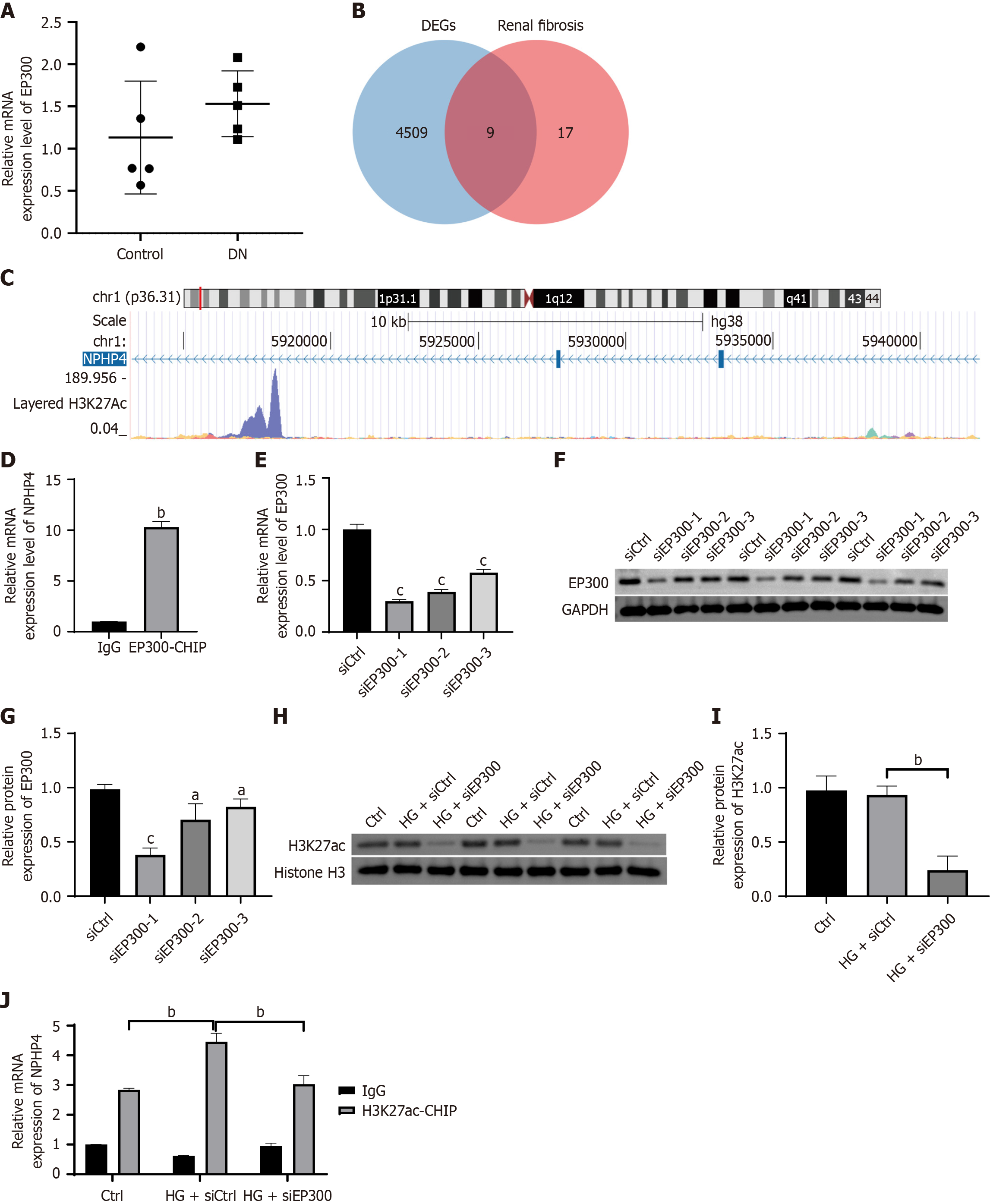
EP300 expression was elevated in the DN group compared with controls (Figure 2A). Genome-wide ChIP-seq analysis identified 11716 potential EP300 target genes. Intersection of these targets with previously reported fibrosis-related gene sets yielded nine candidate genes (Figure 2B). Notably, enrichment of H3K27ac was detected near the TSS of NPHP4 (Figure 2C), suggesting that EP300 may regulate NPHP4 expression through histone acetylation. This finding was further supported at the protein level by western blot analysis (Figure 2D).
To further evaluate the regulatory role of EP300 in NPHP4 expression, we transfected HK-2 cells with three EP300-specific siRNAs (siEP300-1, siEP300-2, siEP300-3) or a negative control siRNA (siCtrl) using lentiviral vectors. RT-qPCR and western blot analysis identified siEP300-1 as the most efficient knockdown construct (Figure 2E-G), and this siRNA was used in subsequent experiments. Western blot analysis (Figure 2H and I) showed that although global H3K27ac levels remained unchanged under HG conditions, EP300 silencing significantly reduced HG-induced NPHP4 upregulation (Figure 2J). These results demonstrate that EP300 promotes NPHP4 expression through H3K27ac-dependent regulation. Collectively, our results suggest that EP300 enhances NPHP4 transcriptional activation by mediating H3K27ac enrichment at the NPHP4 TSS.
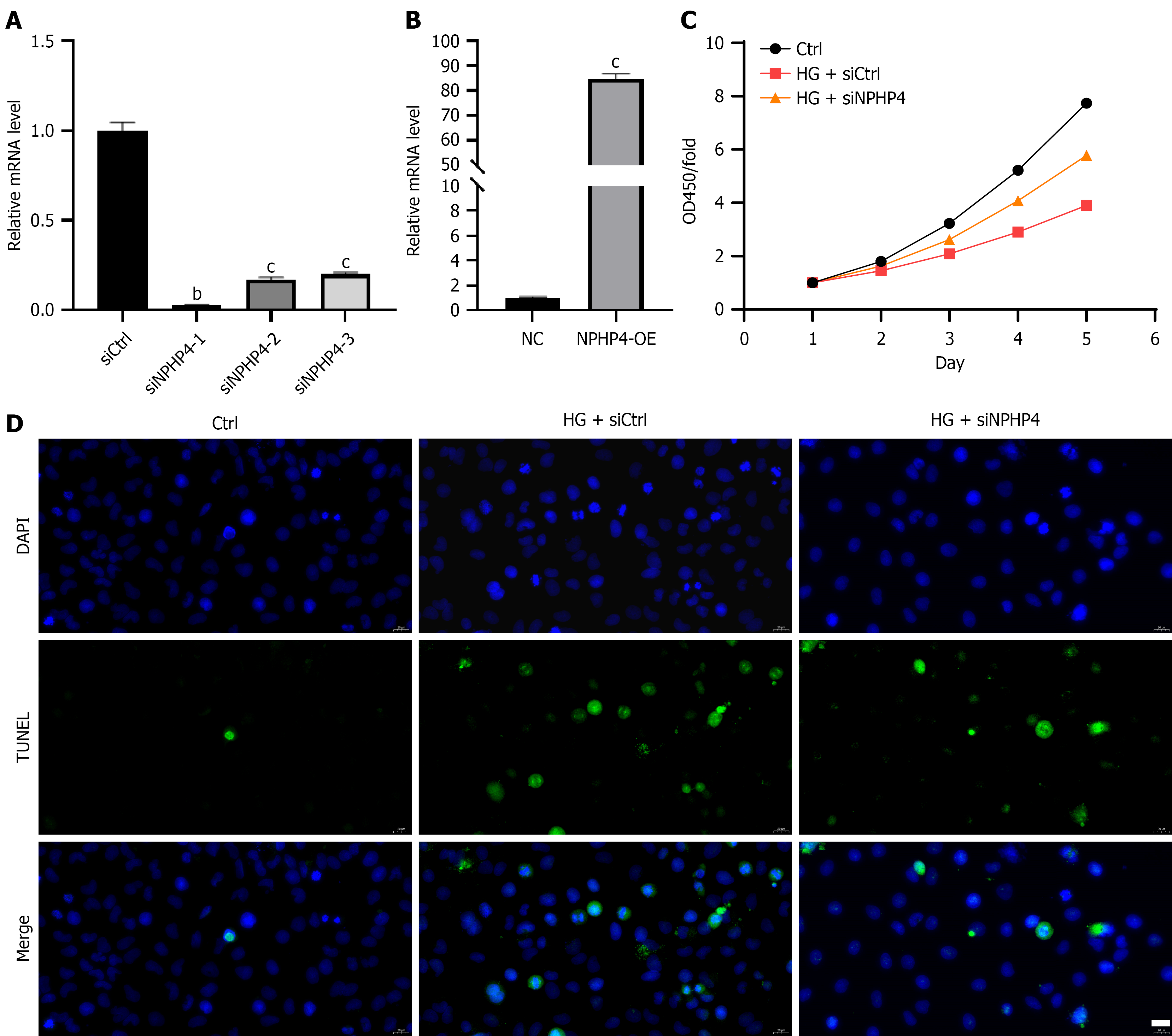
To investigate the functional mechanism of NPHP4 in DN, we established an in vitro hyperglycemia model by treating HK-2 cells with 30 mmol/L D-glucose. RT-qPCR analysis confirmed efficient knockdown of NPHP4 by three independent siRNAs (siNPHP4-1/2/3), resulting in reductions of NPHP4 mRNA levels by 97%, 83%, and 80%, respectively (Figure 3A). In contrast, transfection with an NPHP4 overexpression plasmids significantly increased NPHP4 mRNA levels compared with the negative control group (Figure 3B). Functional assays demonstrated that, under HG conditions, NPHP4 silencing significantly enhanced cell viability and reduced apoptosis, as assessed by Cell Counting Kit-8 (CCK-8) assays and TUNEL staining (Figure 3C and D). These results confirm that NPHP4 contributes to HG-induced renal tubular epithelial cell dysfunction by regulating cell viability and apoptosis, implicating NPHP4 in DN progression.
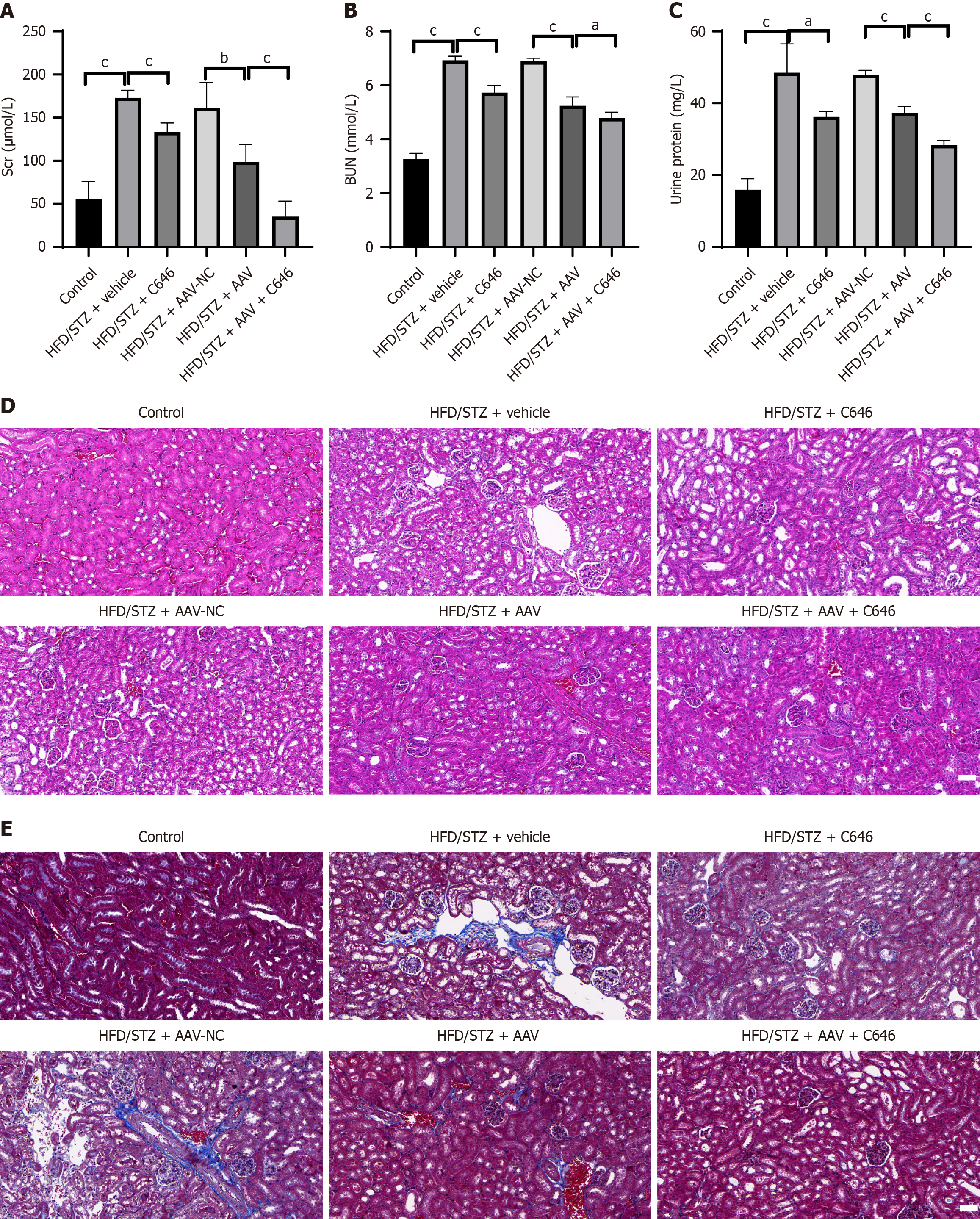
To evaluate the effects of EP300 inhibition and NPHP4 silencing on renal function and histopathology, we established a DN animal model using a high-fat diet (HFD) combined with streptozotocin (STZ) administration. Biochemical analysis (Figure 4A-C) revealed significantly elevated serum creatinine, blood urea nitrogen, and urinary protein levels in the HFD/STZ group compared with the control group and were markedly ameliorated following treatment with the EP300 inhibitor C646 or AAV-mediated NPHP4 silencing. Notably, combined treatment produced a great reduction in these parameters than either monotherapy. Histopathological examination (Figure 4D and E) revealed characteristic renal lesions in the HFD/STZ group, including glomerular hypertrophy, structural disorganization, and increased collagen deposition. All treatment groups exhibited histological improvement, with the combined intervention group conferring the most pronounced renal protection.
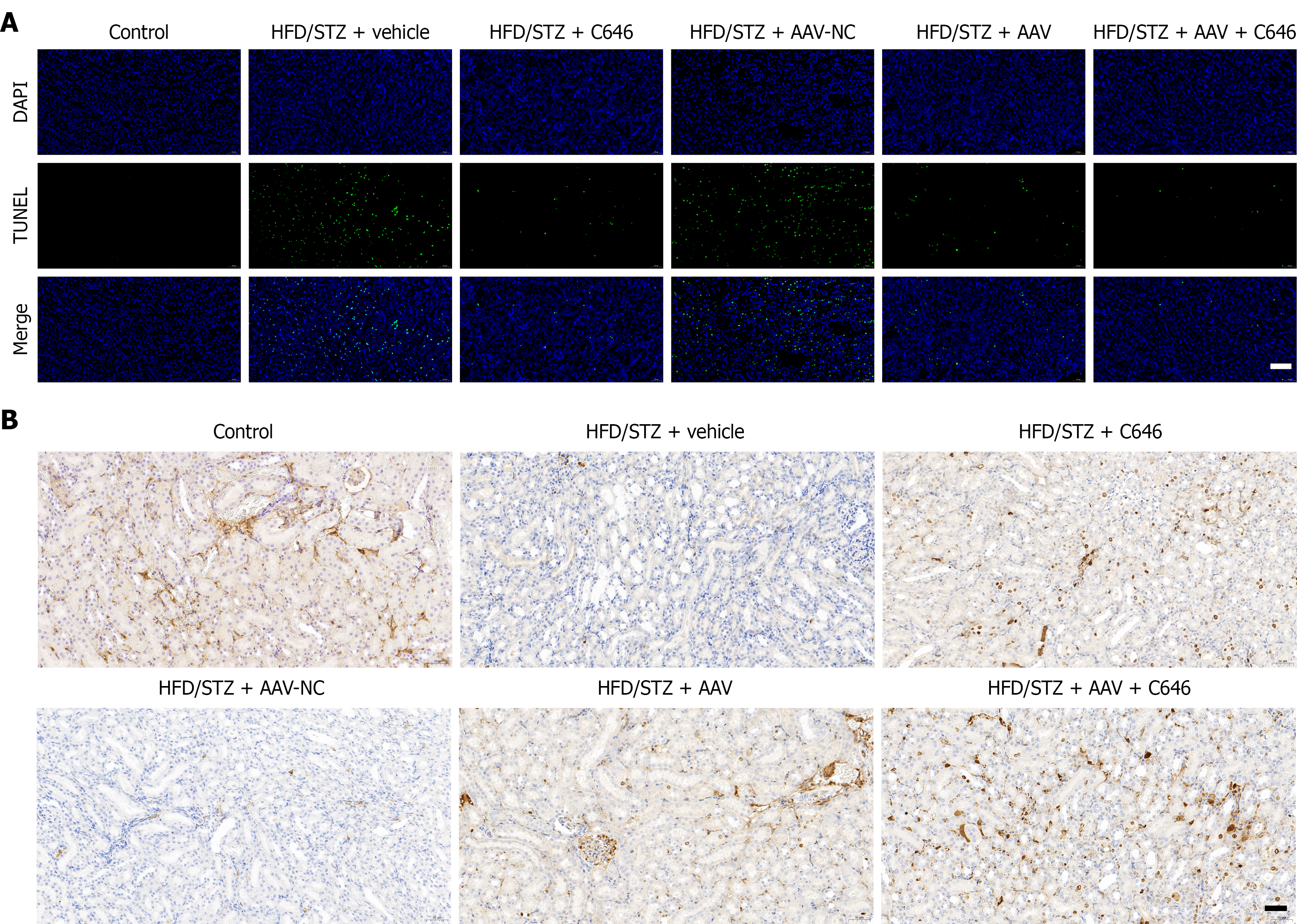
Furthermore, to assess the role of EP300 in regulating the balance between apoptosis and proliferation in DN, TUNEL staining and immunohistochemistry analyses were performed (Figure 5). Hyperglycemic and hyperlipidemic conditions significantly increased renal cell apoptosis while suppressing expression of the proliferation marker Ki67, consistent with enhanced EP300 activity and impaired renal tissue repair. Treatment with the EP300 inhibitor C646 or AAV-mediated NPHP4 silencing markedly reduced apoptosis and restored Ki67 expression. These findings indicate that EP300 contributes to apoptosis-proliferation imbalance in DN, at least in part by upregulating NPHP4. Targeting EP300 activity or NPHP4 expression effectively corrects this pathological imbalance, with combined inhibition of upstream EP300 and downstream NPHP4 pathways producing the most pronounced protective effects, suggesting potential therapeutic synergy.
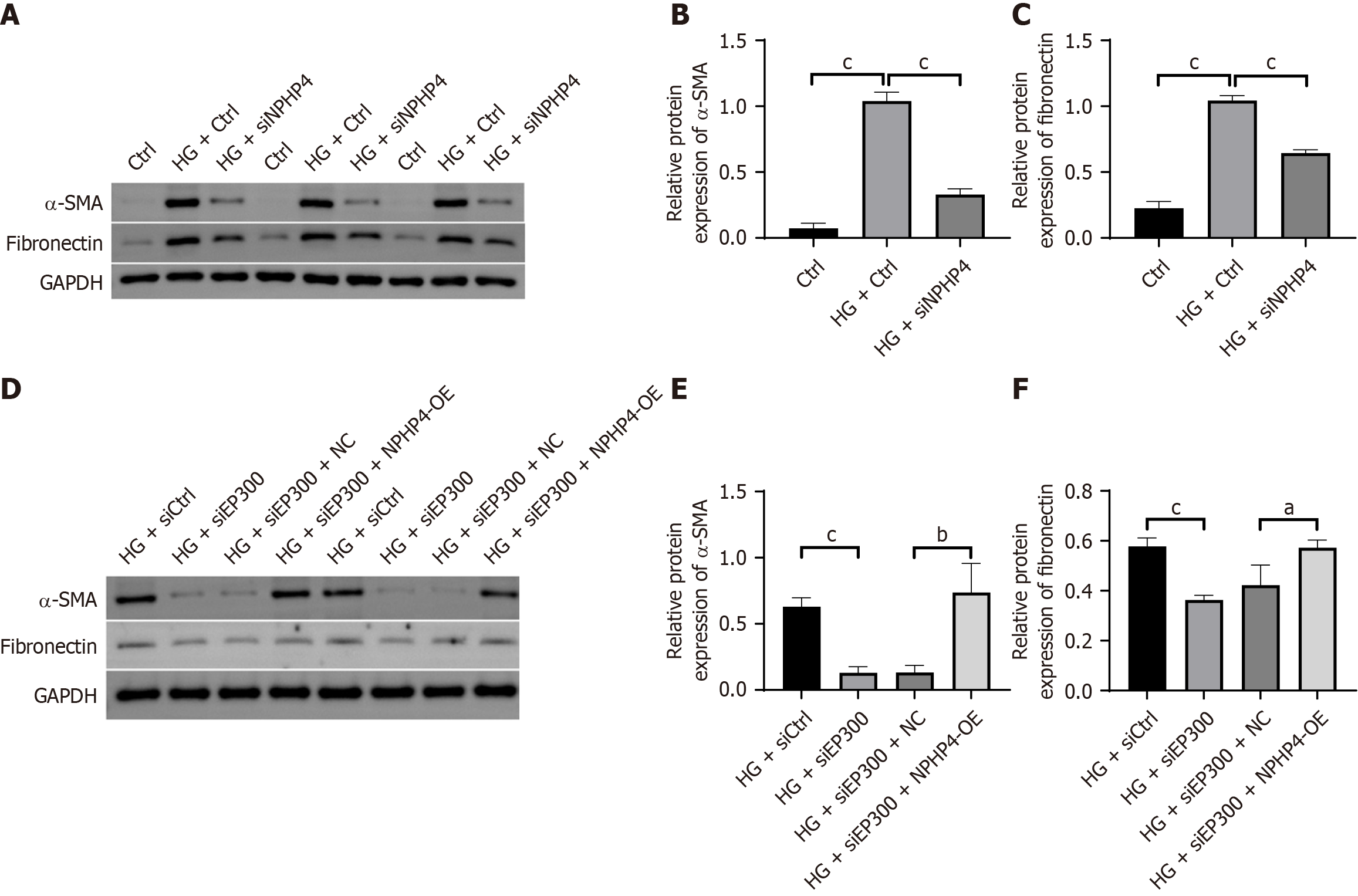
To elucidate the mechanistic role of the EP300-NPHP4 axis in DN-associated fibrosis, we conducted systematic in vivo and in vitro analyses. In in vitro experiments, HG treatment significantly increased protein expression of fibrosis markers α-SMA and fibronectin. Compared with the HG + siCtrl group, silencing of either EP300 or NPHP4 markedly attenuated HG-induced upregulation of α-SMA and fibronectin (Figure 6A-C). Moreover, NPHP4 overexpression in EP300 knockdown partially restored α-SMA and fibronectin expression compared to EP300 silencing alone (Figure 6D-F), suggesting that NPHP4 acts downstream of EP300 to mediate fibrotic signaling.
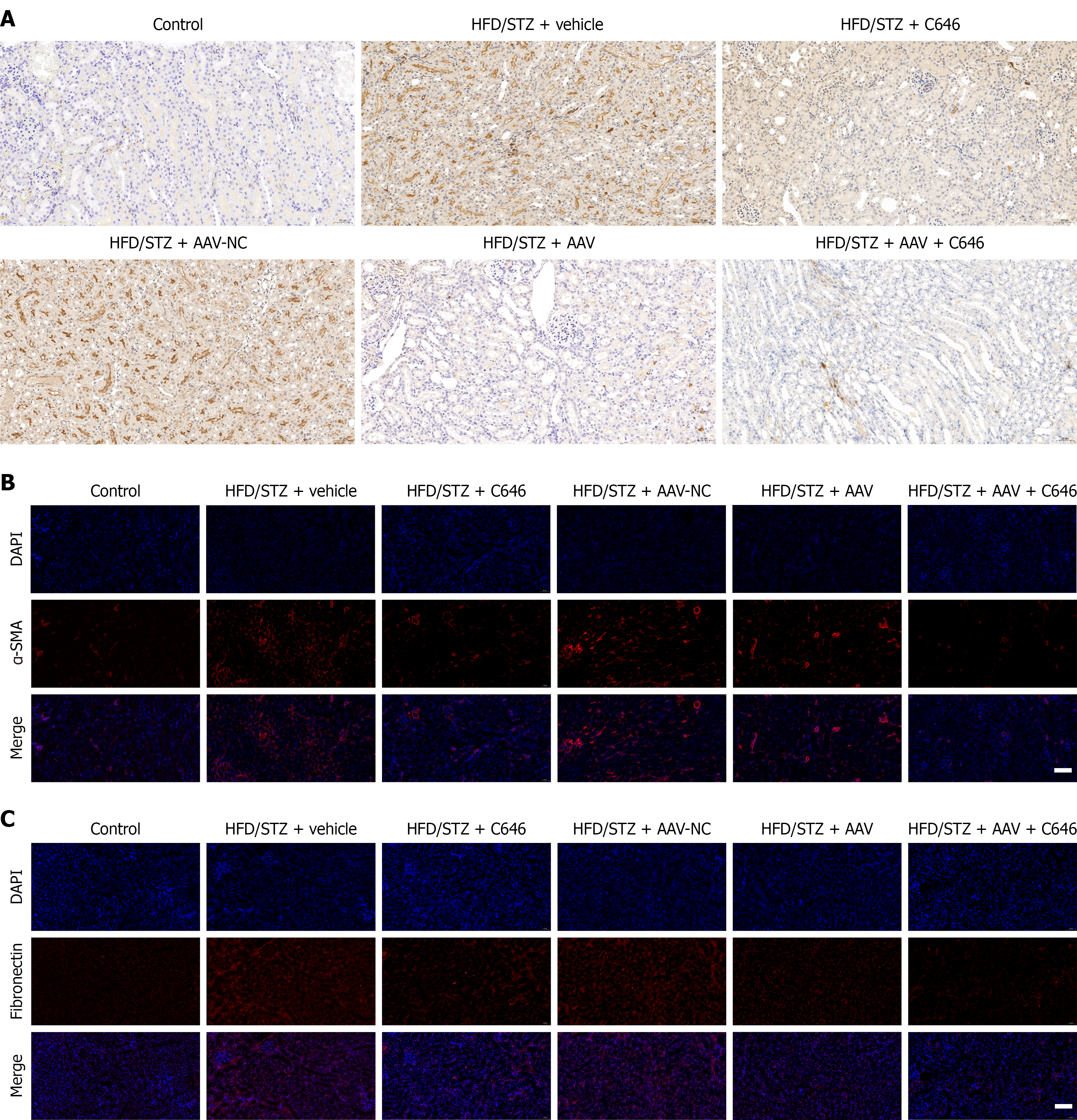
In vivo results strongly corroborated the in vitro findings. Renal tissue immunohistochemistry (Figure 7A) and immunofluorescence (Figure 7B and C) revealed significantly increased NPHP4 expression in the DN model compared with controls, accompanied by marked accumulation of the fibrotic markers α-SMA and fibronectin. Treatment with the EP300 inhibitor C646 or AAV-mediated NPHP4 silencing reduced NPHP4 expression and concomitantly decreased fibrosis marker levels. Notably, combined treatment produced a greater reduction in NPHP4 expression and fibrosis severity than either intervention alone. Collectively, these molecular, cellular, and whole-animal data support a model in which EP300 promotes renal fibrosis, at least in part, through regulation of NPHP4 expression in DN.
In the pathological progression of DN, renal fibrosis serves as the central pathological alteration that directly drives progressive renal function deterioration. Recent studies suggest that aberrant epigenetic modifications may contribute to this process[20], yet the precise regulatory mechanisms remain unclear. While multiple investigations have confirmed that imbalances between HATs and deacetylases can influence DN progression by modulating insulin signaling pathways and inflammatory responses[21,22], direct evidence remains limited regarding how histone acetylation modifications specifically regulate the expression of fibrosis-related key genes.
Indeed, emerging studies have begun to map the epigenetic landscape in DN, revealing alterations in global histone acetylation patterns and highlighting the dysregulation of various histone modifiers, including certain HATs and dea
Building upon this research, our study is the first to establish a functional link between the HAT EP300 and the cilia-associated gene NPHP4. As a crucial epigenetic regulator, the mechanism by which EP300 activates downstream gene expression through H3K27ac modification has been documented in various diseases[23,24], yet remains understudied in DN. We discovered that EP300 specifically binds to the NPHP4 promoter region and enhances its transcriptional acti
Notably, our finding that NPHP4 upregulation promotes renal fibrosis is in contrast with earlier reports where loss-of-function mutations in NPHP4 have been associated with cystic kidney diseases and fibrotic lesions[16,17]. We propose that this discrepancy highlights the context-dependent functionality of NPHP4. In genetic ciliopathies, NPHP4 deficiency impairs primary cilium assembly and integrity, leading to developmental defects and subsequent injury-induced fibrosis. In contrast, in the acquired metabolic milieu of DN, sustained hyperglycemic and inflammatory stimuli may pathologically induce NPHP4 overexpression. Under such conditions, NPHP4 may transcend its classical structural role and acquire novel, gain-of-function properties that actively drive profibrotic signaling, apoptosis-proliferation imbalance, and ECM deposition. This paradigm is reminiscent of other molecules, such as p53, which can exert either tumor-suppressive or senescence-promoting effects depending on cellular context and stress duration. Therefore, our data do not contradict but rather expand the understanding of NPHP4, revealing its potential as a pathological driver in acquired metabolic fibrosis, distinct from its role as a structural guardian in genetic developmental disorders. Future studies are needed to elucidate the precise molecular switch that converts NPHP4 from a ciliary protector to a fibrotic accelerator.
The specificity of EP300-mediated H3K27ac enrichment at the NPHP4 promoter despite unchanged global levels aligns with its role as a recruited transcriptional co-activator. Our ENCODE-based analysis confirms EP300-accessible chromatin at this locus. We propose that diabetic signals activate specific transcription factors that recruit EP300 to the NPHP4 promoter, enabling local H3K27ac deposition and transcriptional activation without altering the genome-wide acetylation landscape.
Fibrosis, as a common terminal pathological alteration in various chronic diseases, may be influenced by epigenetic regulatory mechanisms. This study reveals that in the context of DN, EP300 significantly upregulates NPHP4 expression by specifically acetylating H3K27 at the NPHP4 gene promoter region, thereby promoting renal fibrotic progression. This finding aligns with recent reports on the role of histone acetylation in renal, cardiac, and pulmonary fibrosis[27-29], suggesting that EP300-mediated epigenetic regulation may represent a shared mechanism underlying multi-organ fibrosis. Notably, in vitro gene silencing experiments confirmed that either EP300 or NPHP4 knockdown significantly suppressed HG -induced fibrosis marker expression (Figure 6) and ameliorated the apoptosis-proliferation imbalance (Figure 3). Furthermore, NPHP4 overexpression partially reversed the anti-fibrotic effects of EP300 silencing, solidifying the upstream-downstream regulatory relationship within the EP300-NPHP4 axis. In vivo studies similarly demonstrated that both EP300 inhibitor C646 and AAV-mediated NPHP4 silencing alleviated renal fibrosis, with combined intervention showing synergistic effects (Figure 7). These functional experiments, through both gain-function and loss-of-function approaches, definitively establish that EP300 drives fibrotic progression by regulating NPHP4 expression through H3K27ac-dependent transcriptional activation. The discovery of acetylation-mediated regulation of NPHP4, a cilia-associated gene, provides novel molecular insights into the epigenetic basis of organ fibrosis.
While this study identifies a novel role for the EP300-NPHP4 axis in DN fibrosis, several limitations should be acknowledged. First, our in vivo experiments utilized the HFD/STZ model in C57BL/6 mice. Although this model recapitulates key features of human DN, including hyperglycemia, insulin resistance, and progressive renal fibrosis, these mice are less prone to developing the advanced glomerulosclerosis seen in other models (e.g., db/db mice) or in late-stage human DN. This may affect the generalizability of our findings to the most severe fibrotic phenotypes. Second, our initial tran
In summary, this study reveals a critical role for the EP300/H3K27ac/NPHP4 regulatory axis in DN-associated fibrosis, but also provides a theoretical foundation for developing targeted intervention strategies. These findings expand our understanding of DN pathogenesis and offer insights for investigating fibrosis in other organs. Future studies should further elucidate the detailed molecular mechanisms of this axis, optimize intervention approaches, and evaluate its clinical potential.
| 1. | Haneda M. Diabetic nephropathy. Nihon Rinsho. 2015;73:489-494. |
| 2. | Liu H, Wang X, Gao H, Yang C, Xie C. Physiological and pathological characteristics of vascular endothelial injury in diabetes and the regulatory mechanism of autophagy. Front Endocrinol (Lausanne). 2023;14:1191426. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 19] [Article Influence: 6.3] [Reference Citation Analysis (1)] |
| 3. | Wu YL, Lin ZJ, Li CC, Lin X, Shan SK, Guo B, Zheng MH, Li F, Yuan LQ, Li ZH. Epigenetic regulation in metabolic diseases: mechanisms and advances in clinical study. Signal Transduct Target Ther. 2023;8:98. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 357] [Cited by in RCA: 292] [Article Influence: 97.3] [Reference Citation Analysis (1)] |
| 4. | Xie Z, Zhang D, Chung D, Tang Z, Huang H, Dai L, Qi S, Li J, Colak G, Chen Y, Xia C, Peng C, Ruan H, Kirkey M, Wang D, Jensen LM, Kwon OK, Lee S, Pletcher SD, Tan M, Lombard DB, White KP, Zhao H, Li J, Roeder RG, Yang X, Zhao Y. Metabolic Regulation of Gene Expression by Histone Lysine β-Hydroxybutyrylation. Mol Cell. 2016;62:194-206. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 284] [Cited by in RCA: 575] [Article Influence: 63.9] [Reference Citation Analysis (1)] |
| 5. | Kwak SH, Park KS. Recent progress in genetic and epigenetic research on type 2 diabetes. Exp Mol Med. 2016;48:e220. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 98] [Cited by in RCA: 150] [Article Influence: 15.0] [Reference Citation Analysis (9)] |
| 6. | He W, Li Q, Li X. Acetyl-CoA regulates lipid metabolism and histone acetylation modification in cancer. Biochim Biophys Acta Rev Cancer. 2023;1878:188837. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 116] [Reference Citation Analysis (1)] |
| 7. | Castellano-Castillo D, Ramos-Molina B, Oliva-Olivera W, Ocaña-Wilhelmi L, Queipo-Ortuño MI, Cardona F. Genome Profiling of H3k4me3 Histone Modification in Human Adipose Tissue during Obesity and Insulin Resistance. Biomedicines. 2021;9:1363. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 10] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 8. | Vidal CM, Alva-Ornelas JA, Chen NZ, Senapati P, Tomsic J, Robles VM, Resto C, Sanchez N, Sanchez A, Hyslop T, Emwas N, Aljaber D, Bachelder N, Martinez E, Ann D, Jones V, Winn RA, Miele L, Ochoa AC, Dietze EC, Natarajan R, Schones D, Seewaldt VL. Insulin Resistance in Women Correlates with Chromatin Histone Lysine Acetylation, Inflammatory Signaling, and Accelerated Aging. Cancers (Basel). 2024;16:2735. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 9. | Zhang B, Day DS, Ho JW, Song L, Cao J, Christodoulou D, Seidman JG, Crawford GE, Park PJ, Pu WT. A dynamic H3K27ac signature identifies VEGFA-stimulated endothelial enhancers and requires EP300 activity. Genome Res. 2013;23:917-927. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 85] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 10. | Huang L, Li L, Cheng B, Xing T. SLC38A6, regulated by EP300-mediated modifications of H3K27ac, promotes cell proliferation, glutamine metabolism and mitochondrial respiration in hepatocellular carcinoma. Carcinogenesis. 2022;43:885-894. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 11. | Kang Y, Kim YW, Kang J, Kim A. Histone H3K4me1 and H3K27ac play roles in nucleosome eviction and eRNA transcription, respectively, at enhancers. FASEB J. 2021;35:e21781. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 61] [Article Influence: 12.2] [Reference Citation Analysis (1)] |
| 12. | Rafehi H, Kaspi A, Ziemann M, Okabe J, Karagiannis TC, El-Osta A. Systems approach to the pharmacological actions of HDAC inhibitors reveals EP300 activities and convergent mechanisms of regulation in diabetes. Epigenetics. 2017;12:991-1003. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 19] [Article Influence: 2.4] [Reference Citation Analysis (1)] |
| 13. | Rubio K, Molina-Herrera A, Pérez-González A, Hernández-Galdámez HV, Piña-Vázquez C, Araujo-Ramos T, Singh I. EP300 as a Molecular Integrator of Fibrotic Transcriptional Programs. Int J Mol Sci. 2023;24:12302. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 28] [Reference Citation Analysis (1)] |
| 14. | Zhang M, Barroso E, Peña L, Rada P, Valverde ÁM, Wahli W, Palomer X, Vázquez-Carrera M. PPARβ/δ attenuates hepatic fibrosis by reducing SMAD3 phosphorylation and p300 levels via AMPK in hepatic stellate cells. Biomed Pharmacother. 2024;179:117303. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 17] [Reference Citation Analysis (1)] |
| 15. | Tao J, Zhang M, Wen Z, Wang B, Zhang L, Ou Y, Tang X, Yu X, Jiang Q. Inhibition of EP300 and DDR1 synergistically alleviates pulmonary fibrosis in vitro and in vivo. Biomed Pharmacother. 2018;106:1727-1733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 16. | Miri Karam Z, Gohari AK, Khabaz MJR, Yari A, Meybodi SME, Attari R, Torabi M, Vafaeie F, Moraddahande FM, Amiri S, Saeidi K. Identification of a Novel Deletion Variant (c.2999_3005delTGTGTGT/p.Asn1000SerfsTer4) in NPHP4 Associated With Nephronophthisis-4. J Clin Lab Anal. 2024;38:e25077. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 17. | Otto E, Hoefele J, Ruf R, Mueller AM, Hiller KS, Wolf MT, Schuermann MJ, Becker A, Birkenhäger R, Sudbrak R, Hennies HC, Nürnberg P, Hildebrandt F. A gene mutated in nephronophthisis and retinitis pigmentosa encodes a novel protein, nephroretinin, conserved in evolution. Am J Hum Genet. 2002;71:1161-1167. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 158] [Article Influence: 6.6] [Reference Citation Analysis (1)] |
| 18. | Zhang YC, Du WQ, Zhang RY, Zheng JN, Pei DS. Jade-1: its structure, regulation and functions in the renal cancer. Curr Mol Med. 2016;16:63-69. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 19. | Dai Q, Ye Y. Development and Validation of a Novel Histone Acetylation-Related Gene Signature for Predicting the Prognosis of Ovarian Cancer. Front Cell Dev Biol. 2022;10:793425. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 21] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 20. | Sakshi, Ragini, Saini A, Verma C, Mani I. Epigenetics in renal diseases. Prog Mol Biol Transl Sci. 2023;198:61-71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (1)] |
| 21. | van den Bosch T, Boichenko A, Leus NGJ, Ourailidou ME, Wapenaar H, Rotili D, Mai A, Imhof A, Bischoff R, Haisma HJ, Dekker FJ. The histone acetyltransferase p300 inhibitor C646 reduces pro-inflammatory gene expression and inhibits histone deacetylases. Biochem Pharmacol. 2016;102:130-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 51] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 22. | Lazar AG, Vlad ML, Manea A, Simionescu M, Manea SA. Activated Histone Acetyltransferase p300/CBP-Related Signalling Pathways Mediate Up-Regulation of NADPH Oxidase, Inflammation, and Fibrosis in Diabetic Kidney. Antioxidants (Basel). 2021;10:1356. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 32] [Article Influence: 6.4] [Reference Citation Analysis (1)] |
| 23. | Wang M, Huang Z, Li X, He P, Sun H, Peng Y, Fan Q. Apabetalone, a BET protein inhibitor, inhibits kidney damage in diabetes by preventing pyroptosis via modulating the P300/H3K27ac/PLK1 axis. Pharmacol Res. 2024;207:107306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 15] [Article Influence: 7.5] [Reference Citation Analysis (1)] |
| 24. | Xiang H, Tang H, He Q, Sun J, Yang Y, Kong L, Wang Y. NDUFA8 is transcriptionally regulated by EP300/H3K27ac and promotes mitochondrial respiration to support proliferation and inhibit apoptosis in cervical cancer. Biochem Biophys Res Commun. 2024;693:149374. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 11] [Article Influence: 5.5] [Reference Citation Analysis (1)] |
| 25. | Wang L, Wen X, Wang Z, Lin Z, Li C, Zhou H, Yu H, Li Y, Cheng Y, Chen Y, Lou G, Pan J, Cao M. Ciliary transition zone proteins coordinate ciliary protein composition and ectosome shedding. Nat Commun. 2022;13:3997. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 39] [Article Influence: 9.8] [Reference Citation Analysis (1)] |
| 26. | Mc Fie M, Koneva L, Collins I, Coveney CR, Clube AM, Chanalaris A, Vincent TL, Bezbradica JS, Sansom SN, Wann AKT. Ciliary proteins specify the cell inflammatory response by tuning NFκB signalling, independently of primary cilia. J Cell Sci. 2020;133:jcs239871. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 25] [Article Influence: 4.2] [Reference Citation Analysis (1)] |
| 27. | Shen F, Zhuang S. Histone Acetylation and Modifiers in Renal Fibrosis. Front Pharmacol. 2022;13:760308. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 24] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 28. | Zhan W, Liao X, Xie RJ, Tian T, Yu L, Liu X, Liu J, Li P, Han B, Yang T, Zhang B, Cai LJ, Li R, Yang Q. The effects of blueberry anthocyanins on histone acetylation in rat liver fibrosis. Oncotarget. 2017;8:96761-96773. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 25] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 29. | Li M, Zheng Y, Yuan H, Liu Y, Wen X. Effects of dynamic changes in histone acetylation and deacetylase activity on pulmonary fibrosis. Int Immunopharmacol. 2017;52:272-280. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 35] [Article Influence: 3.9] [Reference Citation Analysis (1)] |