Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115275
Revised: December 15, 2025
Accepted: February 4, 2026
Published online: April 15, 2026
Processing time: 183 Days and 6.3 Hours
Gestational diabetes mellitus (GDM) affects up to 14% of pregnancies globally and remains a major threat to maternal and fetal health, contributing to complications such as preeclampsia, macrosomia, and long-term metabolic disorders. As GDM involves dynamic immune-metabolic changes at the maternal-fetal interface, characterizing its gene expression landscape across relevant tissues is essential to identify biomarkers, therapeutic targets, and disease mechanisms. Over the past several years, numerous transcriptomic studies and publicly available datasets have been generated to uncover molecular mechanisms of GDM across diverse tissues and experimental models. Despite their potential, these resources remain underutilized, with many datasets yet to be fully mined for their biological and clinical insights. This review provides an overview of methodological advances in gene expression profiling relevant to GDM and highlights key tissues and models used. We discuss limitations of current datasets such as the scarcity of single-cell data and metadata inconsistencies and propose future directions, including in
Core Tip: Gestational diabetes mellitus (GDM) affects up to 14% of pregnancies and poses significant risks to both mothers and offspring. This mini-review summarizes publicly available transcriptomic datasets related to GDM across tissues, data types, and platforms, highlighting critical gaps such as the underrepresentation of pancreatic islets and adipose tissue. It also outlines opportunities for integrative, multi-omics, and cell-type resolved approaches. By consolidating current resources, this review provides a roadmap for advancing biomarker discovery, therapeutic development, and mechanistic understanding of GDM pathogenesis.
- Citation: Xie LL, Li SW, Zou Y, Qin D, Sun J, Xiao YX, Li T, Hao YJ, Li B. Mining transcriptomic data for gestational diabetes mellitus: What public datasets reveal. World J Diabetes 2026; 17(4): 115275
- URL: https://www.wjgnet.com/1948-9358/full/v17/i4/115275.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i4.115275
Gestational diabetes mellitus (GDM) is a metabolic disorder defined by glucose intolerance with onset or first recognition during pregnancy[1-3]. Consistent with international guidelines, GDM is commonly diagnosed using the International Association of Diabetes and Pregnancy Study Groups/World Health Organization 75-g oral glucose tolerance test thresholds fasting plasma glucose ≥ 5.1 mmol/L, 1-hour ≥ 10.0 mmol/L, or 2-hour ≥ 8.5 mmol/L[4-6]. Affecting 14% of pregnancies globally[7], GDM is associated with serious short- and long-term complications for both mother and child[8], including preeclampsia[9], macrosomia[10] and an increased risk of metabolic syndrome later in life[11]. Understanding the underlying molecular mechanisms is therefore of great clinical and public health significance.
Over the past decade, transcriptomic profiling techniques such as DNA microarrays, bulk RNA sequencing (bulk RNA-seq), and more recently, single-cell and epitranscriptomic approaches have enabled comprehensive interrogation of gene expression changes in GDM[12]. However, public transcriptomic data on GDM are fragmented, lack standardization and often do not cover key biological tissues such as pancreatic islets or adipose tissue[13]. While several studies focus on placental dysfunction or circulating biomarkers, a comprehensive summary of available datasets and their research scope is lacking[13,14].
To address this gap, we identified and analyzed gene expression datasets related to GDM. We analyzed dataset characteristics, including organism, tissue source, data type, platform, and the presence of control groups, and classified them according to research themes. This mapping not only provides researchers with a practical reference for selecting data suitable for secondary analyses or cross-cohort comparisons, but also highlights tissues, data types, and methodological approaches that remain underrepresented, thereby drawing attention to research gaps with high potential to advance understanding of GDM pathophysiology. Building on these observations, we also outline opportunities for integrative, multi-omics, and cell type resolved analyses that can maximize the value of current resources and help guide future data-generation strategies. Collectively, these contributions transform a scattered and underutilized body of data into a coherent framework that can accelerate biomarker discovery, therapeutic target identification, and mechanistic insight into GDM.
As this article is a narrative minireview, we aimed to summarize representative and single-cell studies related to GDM, rather than to perform a formal systematic review. To ensure clarity in how the literature and datasets were identified, we briefly outline our search approach here.
Relevant publications and datasets were retrieved from PubMed, Gene Expression Omnibus (GEO), ArrayExpress, and Google Scholar. Search terms included various combinations of “gestational diabetes”, “GDM”, “placenta” or “islet” AND “single-cell RNA-seq” or “transcriptome”.
Datasets meeting the following inclusion criteria were considered: (1) Studies focusing on GDM; (2) Publication date between January 2005 and September 2025; (3) English-language publications with publicly accessible raw or processed data; and (4) Transcriptomic or single-cell RNA sequencing (scRNA-seq) datasets with clearly annotated tissue or cell type, species, and sufficient accompanying metadata. When available, ethical approval statements were cross-checked in the original publications.
Additionally, β-cell focused datasets were included for their critical mechanistic insights into β-cell adaptation, proliferation, and dysfunction during pregnancy or metabolic stress, even if not exclusively modeling GDM. The inclusion of these datasets is grounded in the established physiological paradigm that β-cell plasticity is central to gestational glucose homeostasis. Classic work by Kim et al[15] demonstrated that serotonin signaling downstream of placental lactogens drives β-cell proliferation during pregnancy, while more recent single-cell analyses by Chung et al[16] have revealed cell type specific transcriptional adaptations of pancreatic islets across pregnancy and postpartum. Thus, these datasets serve as essential references for understanding β-cell failure in GDM, directly informing the cellular and molecular context of gestational glucose regulation.
All datasets were subjected to a multi-step verification workflow that included: (1) Confirmation of dataset identity by cross-checking GEO/ArrayExpress entries with original publications; (2) Harmonization of sample annotations to ensure consistent naming of tissues, cell types, species, and experimental conditions; (3) Standardized categorization of datasets by organism (human vs mouse) and by tissue type (e.g., placenta, islet, peripheral blood); and (4) Removal of duplicate datasets or re-analyses derived from the same primary study. These procedures ensured accuracy, comparability, and transparency across all included datasets.
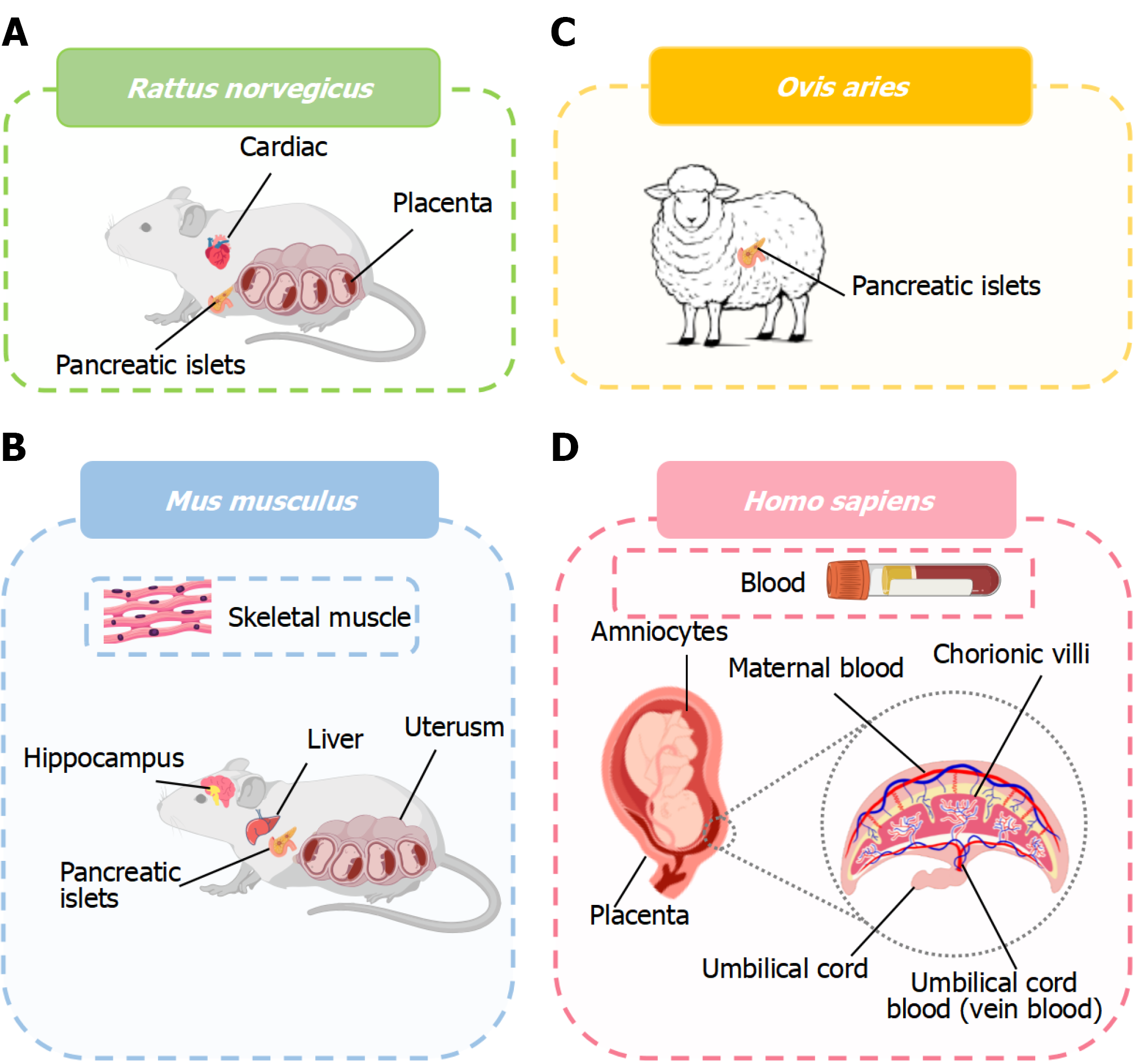
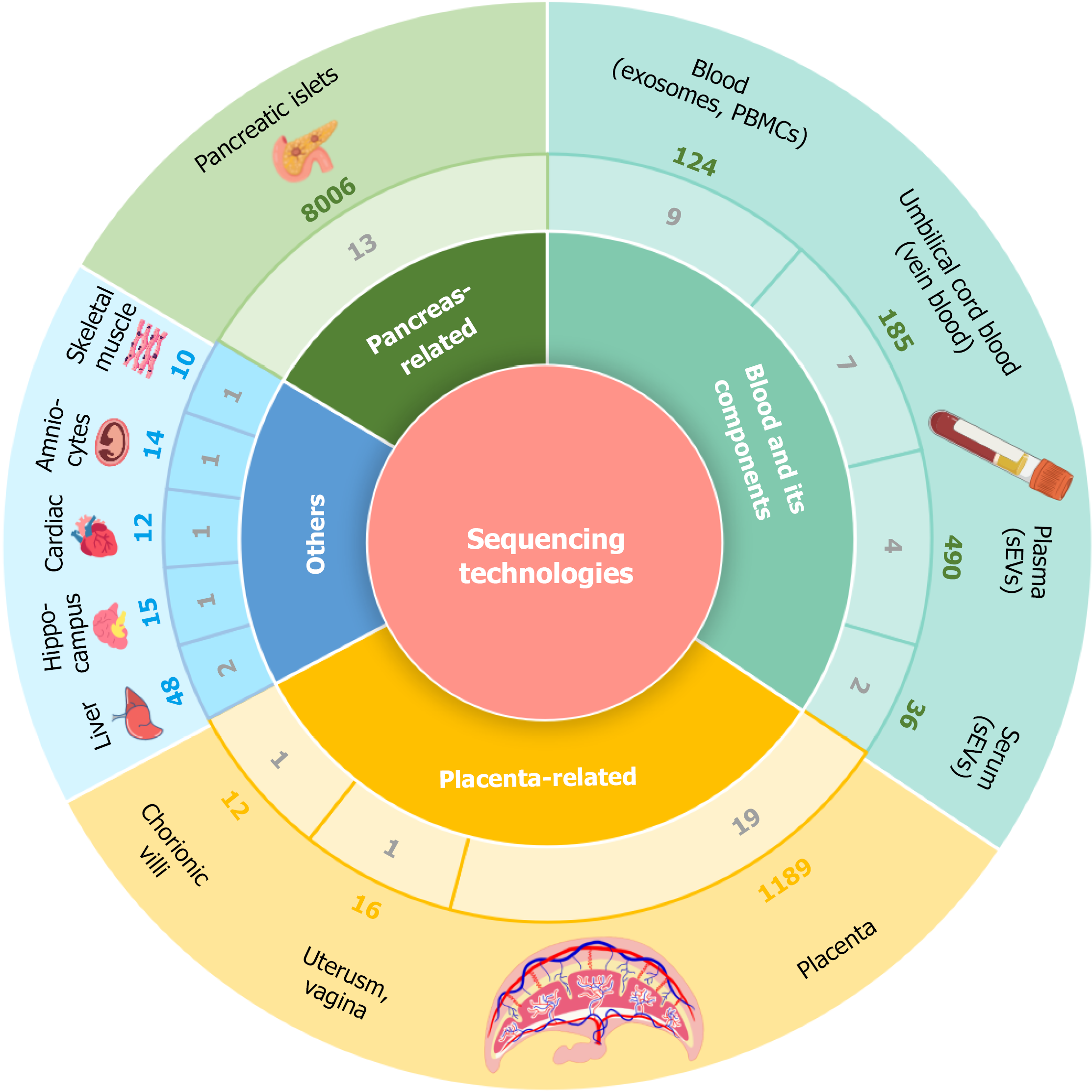
To better understand the biological focus of transcriptomic research in GDM, we surveyed the distribution of sample types, organisms, and corresponding sample sizes across publicly available datasets, as shown in Figures 1 and 2, Table 1 and Supplementary Table 1.
| Series | Organisms | Tissues | Sample source (M/O/B) | Sequencing type | Research theme | Sample count |
| GSE267340 | Homo sapiens | Placenta | M | scRNA-seq | Placenta-related | 4 |
| GSE173193 | Homo sapiens | Placenta | M | scRNA-seq | Placenta-related | 8 |
| GSE249311 | Homo sapiens | Placenta | M | RNA-seq | Placenta-related | 54 |
| GSE154414 | Homo sapiens | Placenta | M | RNA-seq | Placenta-related | 8 |
| GSE144276 | Rattus norvegicus | Placenta | M | RNA-seq | Placenta-related | 6 |
| E-MTAB-11439 | Mus musculus | Liver | M | RNA-seq | Placenta-related | 8 |
| E-MTAB-9203 | Homo sapiens | Placenta | M | RNA-seq | Placenta-related | 14 |
| GSE206042 | Homo sapiens | Placenta | M | ncRNA-seq | Placenta-related | 6 |
| GSE213799 | Homo sapiens | Placenta | O | ncRNA-seq | Placenta-related | 14 |
| GSE112168 | Homo sapiens | Chorionic villi | M | ncRNA-seq | Placenta-related | 12 |
| GSE154415 | Homo sapiens | Placenta | M | Multi-omics | Placenta-related | 24 |
| GSE206041 | Homo sapiens | Placenta | M | ncRNA-seq | Placenta-related | 6 |
| GSE2956 | Homo sapiens | Placenta | M | Microarray | Placenta-related | 1 |
| GSE19649 | Homo sapiens | Blood, placenta | M | Microarray | Placenta-related | 5 |
| GSE103552 | Homo sapiens | Placenta | O | Microarray | Placenta-related | 37 |
| GSE51546 | Homo sapiens | Umbilical cord | O | Microarray | Placenta-related | 12 |
| GSE154413 | Homo sapiens | Placenta | M | ncRNA-seq | Placenta-related | 16 |
| GSE89497 | Homo sapiens | Placenta | M | RNA-seq | Placenta-related | 764 |
| GSE200983 | Homo sapiens | Placenta | M | RNA-seq | Placenta-related | 6 |
| GSE49524 | Homo sapiens | Umbilical cord blood | O | Microarray | Placenta-related | 6 |
| GSE250374 | Homo sapiens | Placenta | M | ncRNA-seq | Placenta-related | 12 |
| GSE236335 | Mus musculus | Uterus, vagina | M | RNA-seq | Placenta-related | 16 |
| GSE87295 | Homo sapiens | Umbilical cord | O | Microarray | Placenta-related | 10 |
| GSE150621 | Homo sapiens | Amniocytes | O | RNA-seq | Circulating biomarkers | 14 |
| GSE70494 | Homo sapiens | Placenta | M | Multi-omics | Circulating biomarkers | 145 |
| GSE70493 | Homo sapiens | Placenta | M | Microarray | Circulating biomarkers | 63 |
| GSE267259 | Homo sapiens | Serum | M | RNA-seq | Circulating biomarkers | 8 |
| GSE284329 | Homo sapiens | PBMCs | M | RNA-seq | Circulating biomarkers | 17 |
| GSE212309 | Homo sapiens | Umbilical cord blood | O | scRNA-seq | Circulating biomarkers | 7 |
| GSE228990 | Homo sapiens | Umbilical cord blood | O | RNA-seq | Circulating biomarkers | 60 |
| GSE192813 | Homo sapiens | Plasma sEVs | M | ncRNA-seq | Circulating biomarkers | 24 |
| GSE216997 | Homo sapiens | Plasma | M | ncRNA-seq | Circulating biomarkers | 328 |
| GSE203346 | Homo sapiens | Umbilical cord blood | B | RNA-seq | Circulating biomarkers | 84 |
| GSE243374 | Homo sapiens | Serum maternal exosomes, serum placental exosomes | M | Microarray | Circulating biomarkers | 28 |
| GSE154377 | Homo sapiens | Plasma | M | RNA-seq | Circulating biomarkers | 134 |
| GSE92772 | Homo sapiens | Blood | M | Multi-omics | Circulating biomarkers | 32 |
| GSE98043 | Homo sapiens | Plasma | M | Microarray | Circulating biomarkers | 4 |
| GSE65737 | Homo sapiens | Umbilical cord vein blood | O | Microarray | Circulating biomarkers | 6 |
| E-MEXP-3382 | Homo sapiens | Blood | M | Microarray | Circulating biomarkers | 6 |
| E-MEXP-3966 | Homo sapiens | Blood | M | Microarray | Circulating biomarkers | 3 |
| E-MEXP-3349 | Homo sapiens | Blood | M | Microarray | Circulating biomarkers | 17 |
| GSE241770 | Mus musculus | Pancreatic islets | M | scRNA-seq | β-cell/islet function | 7899 |
| GSE278861 | Mus musculus | Pancreatic islets | O | scRNA-seq | β-cell/islet function | 9 |
| GSE234741 | Mus musculus | Pancreatic islets | M | scRNA-seq | β-cell/islet function | 2 |
| GSE289077 | Mus musculus | Pancreatic islets | M | RNA-seq | β-cell/islet function | 15 |
| GSE237149 | Mus musculus | Liver | B | RNA-seq | β-cell/islet function | 40 |
| GSE234740 | Mus musculus | Pancreatic islets | M | RNA-seq | β-cell/islet function | 11 |
| GSE116663 | Rattus norvegicus | Pancreatic islets | O | RNA-seq | β-cell/islet function | 12 |
| GSE118323 | Rattus norvegicus | Pancreatic islets | O | RNA-seq | β-cell/islet function | 6 |
| GSE104017 | Rattus norvegicus | Pancreatic islets | O | RNA-seq | β-cell/islet function | 12 |
| GSE90022 | Ovis aries | Pancreatic islets | O | RNA-seq | β-cell/islet function | 8 |
| GSE21860 | Mus musculus | Pancreatic islets | M | RNA-seq | β-cell/islet function | 6 |
| GSE130997 | Mus musculus | Pancreatic islets | M | RNA-seq | β-cell/islet function | 6 |
| GSE36067 | Rattus norvegicus | Pancreatic islets | M | Microarray | β-cell/islet function | 14 |
| GSE100645 | Mus musculus | Pancreatic islets | M | Microarray | β-cell/islet function | 6 |
| GSE255246 | Mus musculus | Skeletal muscle | O | RNA-Seq | β-cell/islet function | 10 |
| GSE136737 | Rattus norvegicus | Cardiac | O | RNA-seq | β-cell/islet function | 12 |
| GSE147039 | Mus musculus | Hippocampus | O | Microarray | β-cell/islet function | 15 |
| GSE194119 | Homo sapiens | Blood exosomes | M | Multi-omics | Predictive models | 6 |
| GSE182737 | Homo sapiens | PBMCs | M | Microarray | Predictive models | 12 |
| GSE114860 | Homo sapiens | Exosomes | M | ncRNA-seq | Predictive models | 28 |
Understanding the biological compartments represented in transcriptomic datasets is essential for interpreting disease-associated molecular signatures in GDM. The available datasets span a broad range of tissues including placenta, blood and circulating components, pancreatic islets, and multiple metabolic organs each offering a distinct window into the pathophysiology of GDM. However, their distribution is highly uneven, with certain tissues heavily profiled and others critically underrepresented. The subsections below examine these tissue sources in detail and highlight their relevance, limitations, and implications for understanding GDM biology.
Placenta-related datasets: Among the surveyed datasets, the placenta emerged as the predominant focus, with more than one-third devoted to this tissue. This predominance underscores its pivotal role at the maternal-fetal interface, where it regulates nutrient exchange, immune tolerance, and endocrine signaling all of which are disrupted in GDM[17,18]. Placental tissue is also easily accessible at delivery, further contributing to its prominence in transcriptomic studies[19,20].
However, placenta-focused datasets capture only a portion of the biology complexity underlying GDM. The placenta is highly heterogeneous, with extravillous trophoblasts, syncytiotropho blasts, villous cytotrophoblasts, endothelial cells, and immune populations exhibiting distinct and sometimes divergent responses to metabolic stress[21]. GDM affects these cell types differently, meaning bulk placental profiles may mask subtype-specific alterations[14]. Moreover, placental transcriptional programs vary markedly with gestational age, and most datasets derive from term samples, limiting insight into early pathogenic events. Maternal glycemia has also shown weak correlations with placental transcriptomic changes in several cohorts, indicating that placental tissue does not fully reflect systemic metabolic dysfunction such as β-cell failure or hepatic insulin resistance[22]. These limitations highlight the need to complement placenta-based studies with data from metabolic tissues and longitudinal sampling.
Blood and circulation-related datasets: Other commonly utilized tissues included peripheral blood, plasma, and cord blood derived endothelial colony-forming cells. These accessible, fluid-based samples provide minimally invasive access to systemic and fetal circulation related transcriptomic changes, making them especially attractive for biomarker discovery[13]. Notably, datasets categorized under blood and its components showed an average sample size of 38, reflecting the growing emphasis on scalable, high-throughput, non-invasive transcriptomic profiling[23,24], particularly through circulating microRNAs (miRNAs) and exosomal RNA[25].
Pancreatic islet-related datasets: There exist thirteen datasets related to pancreatic islet, including 8006 samples, largely derived from rodent models. These datasets enable exploration of β-cell proliferation, insulin secretion, mitochondrial adaptation, and endocrine stress responses under gestational metabolic load. Despite their biological relevance, pancreas-related datasets remain fewer in number than placenta-related ones, indicating a gap between clinical disease me
Other tissues-focused datasets: In contrast, other solid metabolic tissues such as, cardiac, skeletal muscle and liver were underrepresented, despite their direct involvement in glucose metabolism and insulin resistance[26]. Moreover, no datasets were available for adipose tissue, despite its established role in insulin sensitivity, lipid storage and inflammation during pregnancy. Their scarcity represents a critical gap in the current dataset landscape and limits mechanistic insight into the systemic nature of GDM.
Maternal and offspring datasets: In addition, for clarity, animal-based datasets were annotated according to the biological source as maternal, offspring, or both in Table 1.
In reviewing the included datasets, we observed that most human studies focused exclusively on the maternal transcriptome, consistent with the clinical emphasis on placental tissue, maternal blood, and pregnancy-associated biomarkers. In contrast, several animal-based studies particularly those involving mouse models of intrauterine hy
Mechanistic implications of missing metabolic tissues: The scarcity of metabolic tissue datasets has important me
Similarly, hepatic insulin resistance is a key contributor to maternal hyperglycemia. The liver orchestrates gestational changes in gluconeogenesis, lipid mobilization, and ketone metabolism; lack of liver transcriptomes limits our ability to characterize these shifts, understand mother fetus metabolic allocation, and identify dysregulated pathways such as peroxisome proliferator-activated receptor signaling, interleukin-6 mediated inflammation, or oxidative stress[37].
Adipose and skeletal muscle also play foundational roles in maternal metabolic remodeling, including insulin sensitivity, lipid storage, adipokine secretion, and immune infiltration[38]. Without datasets from these tissues, cross-organ integration linking placenta-derived signals to systemic insulin resistance remains speculative.
To address these gaps, rodent models provide a powerful complementary approach. They permit controlled dietary or genetic induction of GDM, longitudinal sampling across gestation, isolation of metabolic tissues at defined time points, and application of single-cell multi-omics[39]. Such models can reveal mechanistic pathways in islets, hepatocytes, myocytes, and adipocytes that are inaccessible in human pregnancy, thereby guiding hypotheses for future human studies.
Regarding species distribution, the majority of datasets (67.2%) were derived from Homo sapiens, consistent with the clinical focus and accessibility of human tissues during pregnancy. Human studies primarily focused on placenta, blood, and cord blood, enabling ethically feasible investigation of GDM pathophysiology[17].
In contrast, Mus musculus (21.3%) and Rattus norvegicus (9.8%) were primarily employed in experimental models to study mechanistic aspects of β-cell dysfunction, immune dysregulation, and fetal programming under diabetic con
Additionally, Ovis aries (1.6%) offers a large-animal model with developmental and placental physiology more similar to humans than rodents[43]. Sheep-based studies primarily focusing on fetal pancreatic islets in the context of intrauterine growth restriction associated gestational metabolic stress provide a valuable intermediate platform for investigating endocrine development and metabolic programming in utero, thus bridging the translational gap between rodent models and human studies[44].
Regarding sample size, placental datasets averaged around 62 samples, offering moderate statistical power for differential expression analysis. Blood-based studies were typically smaller (mean approximately 14), limiting interpretability. Conversely, plasma studies showed both high sample sizes and scalability, making them well-suited for biomarker discovery, although less informative for tissue-specific mechanistic exploration[45].
Taken together, current GDM transcriptomic studies emphasize two dominant strategies: (1) Mechanistic dissection of placental dysfunction; and (2) Scalable biomarker discovery using fluid-based transcriptomes.
However, the limited inclusion of key metabolic tissues and small sample sizes in several studies highlight the need for future work to expand greater biological diversity and apply higher-resolution approaches, thereby capturing GDM’s complex, multi-organ etiology.
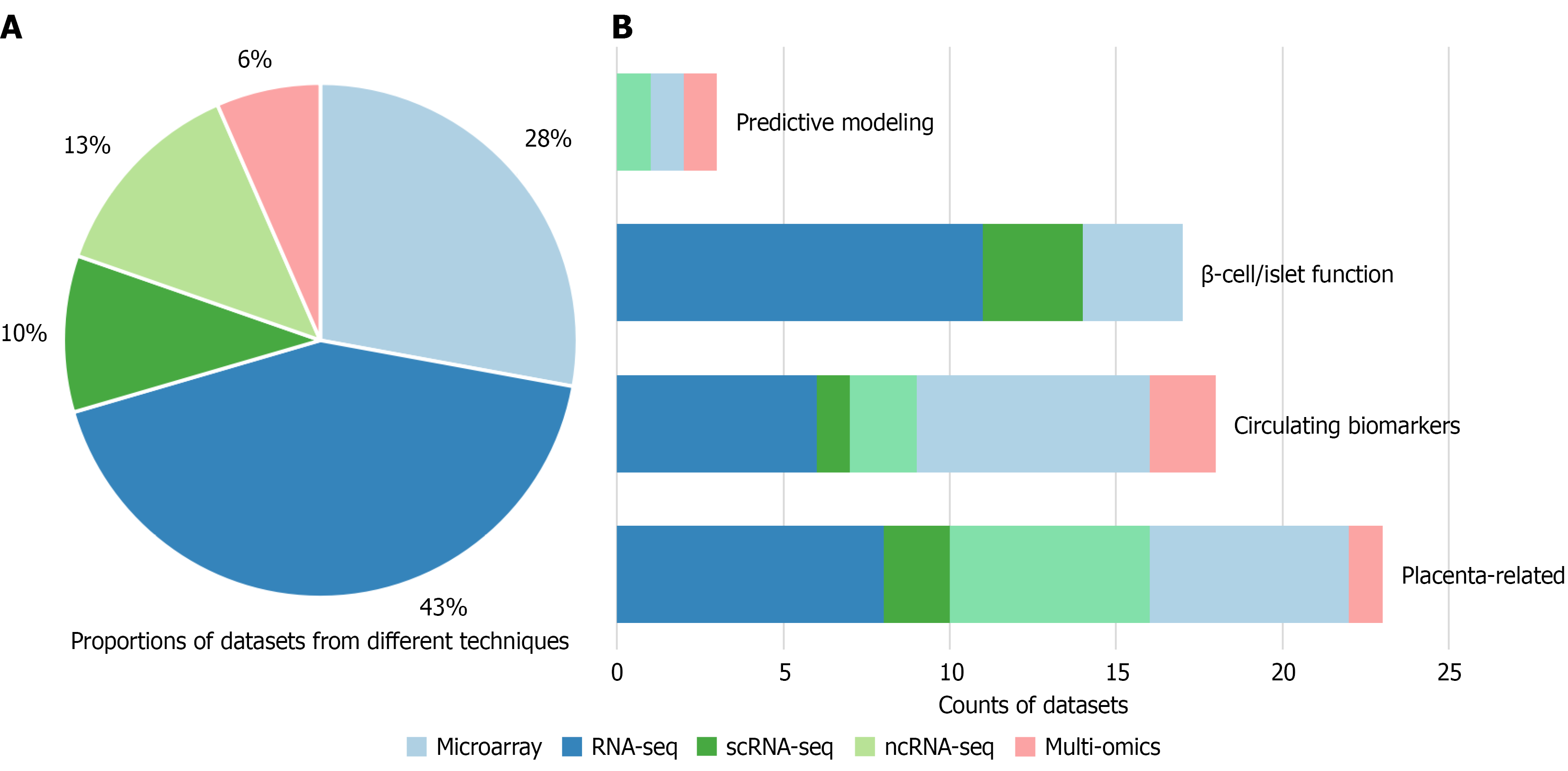
To better characterize the methodological evolution of transcriptomic studies in GDM, we also analyzed the usage of sequencing technologies, and the results are as shown in Figure 3A, Tables 1 and 2.
| Platform | Read configuration/probe architecture | Data output capacity | Advantages | Disadvantages | Frequency | Category |
| NovaSeq 6000 | 2 × 50 ~ 2 × 250 bp (2 × 150 bp common) | Ultra-high | High-throughput, good cost/data ratio | Expensive, best for large projects | 11 | NGS, short-read (Illumina) |
| HiSeq 4000 | 2 × 50 ~ 2 × 150 bp | High | Mainstream, reliable | Being replaced by NovaSeq | 10 | NGS, short-read (Illumina) |
| HiSeq 2000 | 2 × 100 bp | Moderate | Stable, classical platform | Older, limited throughput | 4 | NGS, short-read (Illumina) |
| HiSeq 2500 | 2 × 50 ~ 2 × 125 bp (rapid) | Moderate | Flexible (rapid/high modes) | Phasing out | 6 | NGS, short-read (Illumina) |
| NextSeq 500 | 2 × 75 ~ 2 × 150 bp | Middle | Fast run time, flexible | Limited throughput, costlier | 4 | NGS, short-read (Illumina) |
| NextSeq 2000 | 2 × 50 ~ 2 × 150 bp | Middle-high | Improved chemistry, higher quality | Still relatively costly | 1 | NGS, short-read (Illumina) |
| Illumina BeadChip | 48 k/24 k fixed probes | Ultra-high | Mature, low cost | Lower sensitivity, cannot detect novel transcripts | 5 | DNA microarray |
| Affymetrix | 3’-biased or whole-transcript probes | High | Mature database support | Lower resolution, narrow dynamic range | 4 | DNA microarray |
| Agilent | 8 × 60 k format, 20638 genes | High | Cost-effective, widely used | Limited depth of analysis | 4 | DNA microarray |
Early transcriptomic studies of GDM were dominated by microarray platforms [e.g., Gene 1.0 ST, Human Transcriptome Array 2.0 and specialized miRNA or long noncoding RNA (lncRNA) arrays], and Agilent whole-genome microarrays were frequently employed[46], alongside occasional use of NuGO human arrays and circular RNA (circRNA)-specific arrays[47,48] (as shown in Table 2). These platforms provided the earliest genome-wide expression profiles but were constrained by reduced sensitivity and limited dynamic range compared with next-generation sequencing technologies.
Since 2018, a clear methodological transition has occurred, with bulk RNA-seq rapidly becoming the dominant approach. The bulk RNA-seq offers higher sensitivity, broader transcriptome coverage and greater scalability, and it is now widely adopted as the standard high-throughput strategy for transcriptome interrogation in GDM research.
Notably, a temporal gap exists between the publication of the earliest GDM-related studies and the release of corresponding datasets. For example, microarray-based studies were first published in 2003, yet the earliest datasets did not appear until 2005; similarly, RNA-seq articles emerged in 2008, but the first datasets were only deposited in 2010.
Beyond conventional bulk messenger RNA-focused RNA-seq, recent years have seen a diversification of transcriptomic methods. Non-coding RNA profiling, particularly miRNA-sequencing (miRNA-seq), has shown steady growth from 2021, driven by increasing interest in circulating miRNAs as biomarkers of GDM[49]. Other modalities such as lncRNA-sequencing and circRNA-sequencing have also been applied, extending the molecular scope of GDM studies.
ScRNA-seq has markedly advanced our understanding of cellular heterogeneity at the maternal fetal interface; however, across-study discrepancies and limited dataset coverage remain evident, and single-cell investigations related to GDM are still emerging[50]. In addition, multi-omics approaches that integrate transcriptomics with DNA methylation or RNA modification profiling have begun to appear, underscoring the trend toward more integrative and system-level analyses. By contrast, epitranscriptomic assays [e.g., methylated RNA immunoprecipitation sequencing (MeRIP-seq)] and other specialized methods remain underrepresented, highlighting untapped opportunities for exploring RNA modifications in pregnancy-related disorders[51].
Here again, discrepancies in timing can be observed: Noncoding RNA (ncRNA)-sequencing (ncRNA-seq) articles first appeared in 2015 but the earliest datasets were not released until 2021, whereas both scRNA-seq and multi-omics studies showed near-synchronous trajectories, with publications and datasets emerging together in 2021 and 2015, respectively. These patterns suggest that earlier modalities often followed a “publication first, dataset later” model, while newer approaches increasingly embrace concurrent data sharing, reflecting broader cultural shifts toward open science and FAIR (findable, accessible, interoperable, reusable) data principles.
To improve accessibility for readers, we have added a concise glossary explaining the major transcriptomic technologies including microarray, bulk RNA-seq, ncRNA-seq, scRNA-seq, MeRIP-seq and spatial transcriptomics, as shown in Supplementary Table 2.
In parallel with evolving data types, the technological platforms employed have also shifted. The Illumina HiSeq series (2000, 2500 and 4000) remains the most frequently represented in GDM datasets (n = 24), reflecting its long-standing role as the mainstream instrument for large-scale transcriptome profiling[52,53]. More recently, the Illumina NovaSeq 6000 (n = 11) has gained prominence owing to its ultra-high throughput and cost-effectiveness, gradually displacing HiSeq in newly generated datasets. NextSeq instruments (e.g., NextSeq 500) were used only occasionally, often for smaller or targeted RNA-seq projects, while a single dataset employed the DNBSEQ-G400. Overall, these trends highlight the interplay between evolving transcriptomic methods and the platforms selected to generate GDM datasets.
The increasing diversity in transcriptomic methods particularly those capturing small RNAs or RNA modifications reflects the expanding scope of GDM research. At the same time, this methodological heterogeneity underscores the need for harmonized metadata and standardized reporting practices enable robust cross-study comparisons and large-scale integrative analyses[12,54].
To elucidate the research foci represented by current transcriptomic datasets in GDM, we conducted a title-based thematic analysis and categorized the studies into four major themes, as shown in Figure 3B and Supplementary Table 1. Specifically, datasets were grouped according to tissue origin and study purpose: (1) Placenta-related studies focusing on placental tissue or trophoblast-derived samples; (2) Blood-based circulating biomarker studies using whole blood, plasma, serum, or peripheral immune cells; (3) Pancreatic islet-related datasets derived from pancreatic islets, β-cell models, or other metabolic tissues; and (4) Predictive modeling studies aimed at identifying circulating molecular markers or developing diagnostic or predictive signatures.
The largest category comprised placental-focused studies (23 datasets), highlighting the central role of placental dysfunction in GDM pathophysiology. These investigations examined trophoblast gene regulation[55], vascular remo
Biomarker-oriented studies formed another prominent category (18 datasets), particularly those exploring circulating miRNAs and exosome-derived cargoes from plasma[24], serum[59], or placenta[56] as potential diagnostic tools or mechanistic mediators. This trend reflects a growing interest in developing non-invasive molecular markers for early risk prediction.
In contrast, studies on β-cell and islet biology were comparatively sparse (17 datasets) but provide valuable insights into maternal insulin secretion[60] and fetal endocrine development[44,61]. Although limited in number, these datasets contribute important mechanistic understanding and remain underrepresented relative to placenta- and biomarker-focused research[62].
A small number of studies applied predictive modeling (3 datasets), leveraging transcriptomic features to stratify clinical outcomes (e.g., macrosomia) or to derive diagnostic signatures[63].
Overall, this thematic distribution reveals a current dominance of placenta-focused and biomarker-oriented studies in the GDM transcriptomic field[14]. Meanwhile, mechanistic investigations into pancreatic β-cell function, fetal developmental programming and post-transcriptional regulatory layers remain limited but warrant further exploration in future research[13,22].
This mini-review offers a focused yet informative overview of transcriptomic datasets related to GDM available in public repositories. Our analysis of 61 datasets reveals that while the field has significantly expanded over the past decades particularly after 2020 there remains substantial heterogeneity in experimental design, tissue sources and annotation quality[64].
A dominant feature of GDM transcriptomic research is the reliance on placenta, which appeared in 23 datasets. This emphasis reflects the placenta’s central role in GDM pathophysiology and its accessibility at delivery[65]. Meanwhile, a smaller but growing number of studies have begun to explore circulating biomarkers (e.g., plasma-derived exosomal miRNAs), leveraging the potential for non-invasive diagnostics[66]. However, key metabolic tissues such as pancreatic islets, skeletal muscle and adipose remain underrepresented in publicly available transcriptomic data, despite their known involvement in GDM’s systemic effects.
From a methodological standpoint, DNA microarray and bulk RNA-seq are the most widely employed technique, but recent studies are beginning to adopt more advanced methods such as miRNA-seq, MeRIP-seq and scRNA-seq[67,68]. Still, single-cell datasets remain scarce and are often limited to small sample sizes or restricted tissue contexts. No spatial transcriptomic datasets were identified, suggesting an opportunity to apply high-resolution spatial methods to investigate placental architecture and fetal maternal interactions in GDM.
In terms of research themes, the field remains focused on biomarker discovery (especially miRNAs and extracellular vesicles) and placental dysfunction. While these are valuable areas, other critical processes such as maternal immune modulation, β-cell compensation and fetal programming of disease risk have received relatively limited transcriptomic exploration. Moreover, epitranscriptomic regulation, which emerged in two recent datasets (e.g., N6-methyladenosine RNA modification), remains an underdeveloped but promising frontier.
To advance beyond current limitations, several technically actionable future directions can be envisioned. First, applying single-cell multi-omics including scRNA-seq, single-cell assay for transposase-accessible chromatin using sequencing, and emerging spatial transcriptomic platforms to placental tissues would enable high-resolution mapping of trophoblast heterogeneity, immune stromal interactions, and functional differences between villous and decidual compartments[69]. Second, longitudinal sampling across trimesters, rather than single time points at delivery, would provide critical insight into the temporal evolution of GDM, allowing identification of early molecular predictors and dynamic immune metabolic adaptations[70]. Third, future studies should integrate transcriptomic data with comprehensive maternal metabolic phenotyping, encompassing insulin sensitivity indices, lipid metabolism, β-cell function and inflammatory biomarkers, to improve mechanistic interpretation. Fourth, targeted single-cell analysis of pancreatic β-cell adaptation during pregnancy represents a particularly promising direction, with potential to clarify pathways leading to insufficient insulin compensation in GDM. Finally, adopting standardized metadata and reporting frameworks such as minimum information about a microarray experiment and minimum information about sequencing experiments will improve dataset interoperability, reduce annotation inconsistency, and facilitate cross-cohort integration within the GDM transcriptomic field.
This minireview summarizes key trends, strengths and limitations in the current landscape of publicly available transcriptomic datasets on GDM. While the placental transcriptome and circulating biomarkers have been extensively studied, there is a lack of transcriptomic data covering essential metabolic tissues and advanced methodologies such as single-cell and spatial transcriptomics. To advance the field, we recommend expanding dataset diversity to include maternal metabolic organs (e.g., pancreas, adipose tissue), fetal tissues and longitudinal sampling across gestation. Future studies should adopt cutting-edge technologies such as scRNA-seq, spatial transcriptomics and multi-omics integration (e.g., proteomics, epigenomics) to capture the full biological complexity of GDM. Moreover, standardized metadata reporting including gestational age, diagnostic criteria, treatment exposure and maternal fetal outcomes is essential to ensure comparability and reusability. By integrating machine learning approaches and building harmonized trans
| 1. | Sweeting A, Wong J, Murphy HR, Ross GP. A Clinical Update on Gestational Diabetes Mellitus. Endocr Rev. 2022;43:763-793. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 717] [Cited by in RCA: 583] [Article Influence: 145.8] [Reference Citation Analysis (1)] |
| 2. | Moon JH, Jang HC. Gestational Diabetes Mellitus: Diagnostic Approaches and Maternal-Offspring Complications. Diabetes Metab J. 2022;46:3-14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 273] [Cited by in RCA: 234] [Article Influence: 58.5] [Reference Citation Analysis (1)] |
| 3. | McIntyre HD, Catalano P, Zhang C, Desoye G, Mathiesen ER, Damm P. Gestational diabetes mellitus. Nat Rev Dis Primers. 2019;5:47. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1412] [Cited by in RCA: 1190] [Article Influence: 170.0] [Reference Citation Analysis (4)] |
| 4. | International Association of Diabetes and Pregnancy Study Groups Consensus Panel; Metzger BE, Gabbe SG, Persson B, Buchanan TA, Catalano PA, Damm P, Dyer AR, Leiva Ad, Hod M, Kitzmiler JL, Lowe LP, McIntyre HD, Oats JJ, Omori Y, Schmidt MI. International association of diabetes and pregnancy study groups recommendations on the diagnosis and classification of hyperglycemia in pregnancy. Diabetes Care. 2010;33:676-682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2777] [Cited by in RCA: 3406] [Article Influence: 212.9] [Reference Citation Analysis (7)] |
| 5. | Diagnostic criteria and classification of hyperglycaemia first detected in pregnancy: a World Health Organization Guideline. Diabetes Res Clin Pract. 2014;103:341-363. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 704] [Cited by in RCA: 630] [Article Influence: 52.5] [Reference Citation Analysis (1)] |
| 6. | Simmons D, Immanuel J, Hague WM, Teede H, Nolan CJ, Peek MJ, Flack JR, McLean M, Wong V, Hibbert E, Kautzky-Willer A, Harreiter J, Backman H, Gianatti E, Sweeting A, Mohan V, Enticott J, Cheung NW; TOBOGM Research Group. Treatment of Gestational Diabetes Mellitus Diagnosed Early in Pregnancy. N Engl J Med. 2023;388:2132-2144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 285] [Cited by in RCA: 241] [Article Influence: 80.3] [Reference Citation Analysis (1)] |
| 7. | Xu H, Liu R. Comprehensive management of gestational diabetes mellitus: practical efficacy of exercise therapy and sustained intervention strategies. Front Endocrinol (Lausanne). 2024;15:1347754. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 8] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 8. | Damm P, Houshmand-Oeregaard A, Kelstrup L, Lauenborg J, Mathiesen ER, Clausen TD. Gestational diabetes mellitus and long-term consequences for mother and offspring: a view from Denmark. Diabetologia. 2016;59:1396-1399. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 526] [Cited by in RCA: 465] [Article Influence: 46.5] [Reference Citation Analysis (1)] |
| 9. | Weissgerber TL, Mudd LM. Preeclampsia and diabetes. Curr Diab Rep. 2015;15:9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 297] [Cited by in RCA: 242] [Article Influence: 22.0] [Reference Citation Analysis (9)] |
| 10. | Huang X, Zha B, Zhang M, Li Y, Wu Y, Zhang R, Sheng L, Xu J, Yu Z, Gao C, Chen Z, Ding H, Ma L, Zhang Y, Zang S, Zhang TN, Liu J. Decreased Monocyte Count Is Associated With Gestational Diabetes Mellitus Development, Macrosomia, and Inflammation. J Clin Endocrinol Metab. 2022;107:192-204. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 29] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 11. | Grieger JA, Bianco-Miotto T, Grzeskowiak LE, Leemaqz SY, Poston L, McCowan LM, Kenny LC, Myers JE, Walker JJ, Dekker GA, Roberts CT. Metabolic syndrome in pregnancy and risk for adverse pregnancy outcomes: A prospective cohort of nulliparous women. PLoS Med. 2018;15:e1002710. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 152] [Cited by in RCA: 125] [Article Influence: 15.6] [Reference Citation Analysis (1)] |
| 12. | Zhou Z, Yang X. An update review of the application of single-cell RNA sequencing in pregnancy-related diseases. Front Endocrinol (Lausanne). 2024;15:1415173. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 13. | Lewis KA, Chang L, Cheung J, Aouizerat BE, Jelliffe-Pawlowski LL, McLemore MR, Piening B, Rand L, Ryckman KK, Flowers E. Systematic review of transcriptome and microRNAome associations with gestational diabetes mellitus. Front Endocrinol (Lausanne). 2022;13:971354. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 14. | Yang Y, Guo F, Peng Y, Chen R, Zhou W, Wang H, OuYang J, Yu B, Xu Z. Transcriptomic Profiling of Human Placenta in Gestational Diabetes Mellitus at the Single-Cell Level. Front Endocrinol (Lausanne). 2021;12:679582. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 85] [Cited by in RCA: 68] [Article Influence: 13.6] [Reference Citation Analysis (1)] |
| 15. | Kim H, Toyofuku Y, Lynn FC, Chak E, Uchida T, Mizukami H, Fujitani Y, Kawamori R, Miyatsuka T, Kosaka Y, Yang K, Honig G, van der Hart M, Kishimoto N, Wang J, Yagihashi S, Tecott LH, Watada H, German MS. Serotonin regulates pancreatic beta cell mass during pregnancy. Nat Med. 2010;16:804-808. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 514] [Cited by in RCA: 456] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 16. | Chung JY, Ma Y, Zhang D, Bickerton HH, Stokes E, Patel SB, Tse HM, Feduska J, Welner RS, Banerjee RR. Pancreatic islet cell type-specific transcriptomic changes during pregnancy and postpartum. iScience. 2023;26:106439. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 17] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 17. | Lu S, Wang J, Kakongoma N, Hua W, Xu J, Wang Y, He S, Gu H, Shi J, Hu W. DNA methylation and expression profiles of placenta and umbilical cord blood reveal the characteristics of gestational diabetes mellitus patients and offspring. Clin Epigenetics. 2022;14:69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 31] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 18. | Bowman CE, Arany Z, Wolfgang MJ. Regulation of maternal-fetal metabolic communication. Cell Mol Life Sci. 2021;78:1455-1486. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 78] [Article Influence: 15.6] [Reference Citation Analysis (1)] |
| 19. | Yong HEJ, Chan SY. Current approaches and developments in transcript profiling of the human placenta. Hum Reprod Update. 2020;26:799-840. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 57] [Cited by in RCA: 48] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 20. | Pavličev M, Wagner GP, Chavan AR, Owens K, Maziarz J, Dunn-Fletcher C, Kallapur SG, Muglia L, Jones H. Single-cell transcriptomics of the human placenta: inferring the cell communication network of the maternal-fetal interface. Genome Res. 2017;27:349-361. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 290] [Cited by in RCA: 231] [Article Influence: 25.7] [Reference Citation Analysis (1)] |
| 21. | Li H, Peng H, Hong W, Wei Y, Tian H, Huang X, Jia L, Zheng J, Duan T, He Q, Wang K. Human Placental Endothelial Cell and Trophoblast Heterogeneity and Differentiation Revealed by Single-Cell RNA Sequencing. Cells. 2022;12:87. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 26] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 22. | Usman TO, Chhetri G, Yeh H, Dong HH. Beta-cell compensation and gestational diabetes. J Biol Chem. 2023;299:105405. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 35] [Article Influence: 11.7] [Reference Citation Analysis (1)] |
| 23. | Salomon C, Scholz-Romero K, Sarker S, Sweeney E, Kobayashi M, Correa P, Longo S, Duncombe G, Mitchell MD, Rice GE, Illanes SE. Gestational Diabetes Mellitus Is Associated With Changes in the Concentration and Bioactivity of Placenta-Derived Exosomes in Maternal Circulation Across Gestation. Diabetes. 2016;65:598-609. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 270] [Cited by in RCA: 238] [Article Influence: 23.8] [Reference Citation Analysis (1)] |
| 24. | Ye Z, Wang S, Huang X, Chen P, Deng L, Li S, Lin S, Wang Z, Liu B. Plasma Exosomal miRNAs Associated With Metabolism as Early Predictor of Gestational Diabetes Mellitus. Diabetes. 2022;71:2272-2283. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 48] [Article Influence: 12.0] [Reference Citation Analysis (1)] |
| 25. | Légaré C, Desgagné V, Thibeault K, White F, Clément AA, Poirier C, Luo ZC, Scott MS, Jacques PÉ, Perron P, Guérin R, Hivert MF, Bouchard L. First Trimester Plasma MicroRNA Levels Predict Risk of Developing Gestational Diabetes Mellitus. Front Endocrinol (Lausanne). 2022;13:928508. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 8] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 26. | Plows JF, Stanley JL, Baker PN, Reynolds CM, Vickers MH. The Pathophysiology of Gestational Diabetes Mellitus. Int J Mol Sci. 2018;19:3342. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1421] [Cited by in RCA: 1187] [Article Influence: 148.4] [Reference Citation Analysis (5)] |
| 27. | He L, Wang X, Chen X. Unveiling the role of microRNAs in metabolic dysregulation of Gestational Diabetes Mellitus. Reprod Biol. 2024;24:100924. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 2] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 28. | Money KM, Barke TL, Serezani A, Gannon M, Garbett KA, Aronoff DM, Mirnics K. Gestational diabetes exacerbates maternal immune activation effects in the developing brain. Mol Psychiatry. 2018;23:1920-1928. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 53] [Article Influence: 6.6] [Reference Citation Analysis (1)] |
| 29. | Barbosa MR, Marchi GF, Silva KMR, França DCH, Silva MABD, Ribeiro Barbosa J, Melo LVL, França EL, Honorio-França AC. Gestational Diabetes Mellitus Alters Cytokine Profiles and Macrophage Polarization in Human Placenta. Int J Mol Sci. 2025;26:10867. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 30. | Lorenzo PI, Martín-Montalvo A, Cobo Vuilleumier N, Gauthier BR. Molecular Modelling of Islet β-Cell Adaptation to Inflammation in Pregnancy and Gestational Diabetes Mellitus. Int J Mol Sci. 2019;20:6171. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 31. | Yan YS, Mo JY, Huang YT, Zhu H, Wu HY, Lin ZL, Liu R, Liu XQ, Lv PP, Feng C, Sheng JZ, Jin M, Huang HF. Intrauterine hyperglycaemia during late gestation caused mitochondrial dysfunction in skeletal muscle of male offspring through CREB/PGC1A signaling. Nutr Diabetes. 2024;14:56. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 32. | Muralimanoharan S, Maloyan A, Myatt L. Mitochondrial function and glucose metabolism in the placenta with gestational diabetes mellitus: role of miR-143. Clin Sci (Lond). 2016;130:931-941. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 112] [Article Influence: 11.2] [Reference Citation Analysis (1)] |
| 33. | Saucedo R, Magallón-Gayón E, Chavez-Santoscoy RA, Díaz-Velázquez MF, Ferreira-Hermosillo A, Ojeda-López D, Porras-Marcial W, López-Sánchez D, Valencia-Ortega J. Gene Expression Profile of Placenta and Adipose Tissue in Women with Gestational Diabetes Mellitus. Int J Mol Sci. 2025;26:9595. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 34. | Hill DJ, Hill TG. Maternal diet during pregnancy and adaptive changes in the maternal and fetal pancreas have implications for future metabolic health. Front Endocrinol (Lausanne). 2024;15:1456629. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 35. | Salazar-Petres ER, Sferruzzi-Perri AN. Pregnancy-induced changes in β-cell function: what are the key players? J Physiol. 2022;600:1089-1117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 60] [Article Influence: 15.0] [Reference Citation Analysis (1)] |
| 36. | Li J, Fan L, Nan Y, Wang M, Yang S. Focus on Cell Apoptosis, Pyroptosis and Ferroptosis to Explore Strategic Breakthrough for GDM. J Inflamm Res. 2025;18:10355-10373. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 37. | Fang H, Li Q, Wang H, Ren Y, Zhang L, Yang L. Maternal nutrient metabolism in the liver during pregnancy. Front Endocrinol (Lausanne). 2024;15:1295677. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 25] [Article Influence: 12.5] [Reference Citation Analysis (2)] |
| 38. | Lappas M. Effect of pre-existing maternal obesity, gestational diabetes and adipokines on the expression of genes involved in lipid metabolism in adipose tissue. Metabolism. 2014;63:250-262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 77] [Article Influence: 6.4] [Reference Citation Analysis (5)] |
| 39. | Li X, Jia T, Wu Y, Peng Y, Feng Y, Gong L, Dong S, Tian J, Sun L. Multi-omics integration analysis and association study reveal the potential of ADIPOQ function in gestational diabetes mellitus. Nutr Diabetes. 2025;15:9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 40. | Mishra A, Ruano SH, Saha PK, Pennington KA. A novel model of gestational diabetes: Acute high fat high sugar diet results in insulin resistance and beta cell dysfunction during pregnancy in mice. PLoS One. 2022;17:e0279041. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (1)] |
| 41. | Asfari M, Janjic D, Meda P, Li G, Halban PA, Wollheim CB. Establishment of 2-mercaptoethanol-dependent differentiated insulin-secreting cell lines. Endocrinology. 1992;130:167-178. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 949] [Cited by in RCA: 811] [Article Influence: 23.9] [Reference Citation Analysis (1)] |
| 42. | Gao X, He J, Zhu A, Xie K, Yan K, Jiang X, Xu Y, Li Q, Xu A, Ye D, Guo J. Modelling gestational diabetes mellitus: large animals hold great promise. Rev Endocr Metab Disord. 2021;22:407-420. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (1)] |
| 43. | Banstola A, Reynolds JNJ. The Sheep as a Large Animal Model for the Investigation and Treatment of Human Disorders. Biology (Basel). 2022;11:1251. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 78] [Article Influence: 19.5] [Reference Citation Analysis (1)] |
| 44. | Kelly AC, Bidwell CA, McCarthy FM, Taska DJ, Anderson MJ, Camacho LE, Limesand SW. RNA Sequencing Exposes Adaptive and Immune Responses to Intrauterine Growth Restriction in Fetal Sheep Islets. Endocrinology. 2017;158:743-755. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 32] [Article Influence: 3.6] [Reference Citation Analysis (1)] |
| 45. | Suhre K, McCarthy MI, Schwenk JM. Genetics meets proteomics: perspectives for large population-based studies. Nat Rev Genet. 2021;22:19-37. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 426] [Cited by in RCA: 310] [Article Influence: 62.0] [Reference Citation Analysis (1)] |
| 46. | Shashikadze B, Flenkenthaler F, Stöckl JB, Valla L, Renner S, Kemter E, Wolf E, Fröhlich T. Developmental Effects of (Pre-)Gestational Diabetes on Offspring: Systematic Screening Using Omics Approaches. Genes (Basel). 2021;12:1991. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 47. | Ilangovan H, Kothiyal P, Hoadley KA, Elgart R, Eley G, Eslami P. Harmonizing heterogeneous transcriptomics datasets for machine learning-based analysis to identify spaceflown murine liver-specific changes. NPJ Microgravity. 2024;10:61. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 48. | Wartmann H, Heins S, Kloiber K, Bonn S. Bias-invariant RNA-sequencing metadata annotation. Gigascience. 2021;10:giab064. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 49. | Guarino E, Delli Poggi C, Grieco GE, Cenci V, Ceccarelli E, Crisci I, Sebastiani G, Dotta F. Circulating MicroRNAs as Biomarkers of Gestational Diabetes Mellitus: Updates and Perspectives. Int J Endocrinol. 2018;2018:6380463. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 43] [Article Influence: 5.4] [Reference Citation Analysis (1)] |
| 50. | Derisoud E, Jiang H, Zhao A, Chavatte-Palmer P, Deng Q. Revealing the molecular landscape of human placenta: a systematic review and meta-analysis of single-cell RNA sequencing studies. Hum Reprod Update. 2024;30:410-441. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 33] [Article Influence: 16.5] [Reference Citation Analysis (1)] |
| 51. | Binder AM, LaRocca J, Lesseur C, Marsit CJ, Michels KB. Epigenome-wide and transcriptome-wide analyses reveal gestational diabetes is associated with alterations in the human leukocyte antigen complex. Clin Epigenetics. 2015;7:79. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 68] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 52. | Stark R, Grzelak M, Hadfield J. RNA sequencing: the teenage years. Nat Rev Genet. 2019;20:631-656. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1704] [Cited by in RCA: 1249] [Article Influence: 178.4] [Reference Citation Analysis (1)] |
| 53. | Metzger BE, Coustan DR, Trimble ER. Hyperglycemia and Adverse Pregnancy Outcomes. Clin Chem. 2019;65:937-938. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 67] [Article Influence: 9.6] [Reference Citation Analysis (1)] |
| 54. | Fang J, Wu X, He J, Zhang H, Chen X, Zhang H, Novakovic B, Qi H, Yu X. RBM15 suppresses hepatic insulin sensitivity of offspring of gestational diabetes mellitus mice via m6A-mediated regulation of CLDN4. Mol Med. 2023;29:23. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 31] [Article Influence: 10.3] [Reference Citation Analysis (1)] |
| 55. | Nair S, Jayabalan N, Guanzon D, Palma C, Scholz-Romero K, Elfeky O, Zuñiga F, Ormazabal V, Diaz E, Rice GE, Duncombe G, Jansson T, McIntyre HD, Lappas M, Salomon C. Human placental exosomes in gestational diabetes mellitus carry a specific set of miRNAs associated with skeletal muscle insulin sensitivity. Clin Sci (Lond). 2018;132:2451-2467. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 119] [Cited by in RCA: 104] [Article Influence: 13.0] [Reference Citation Analysis (1)] |
| 56. | Zhang L, Wu Q, Zhu S, Tang Y, Chen Y, Chen D, Liang Z. Chemerin-Induced Down-Regulation of Placenta-Derived Exosomal miR-140-3p and miR-574-3p Promotes Umbilical Vein Endothelial Cells Proliferation, Migration, and Tube Formation in Gestational Diabetes Mellitus. Cells. 2022;11:3457. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 17] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 57. | Evangelista AF, Collares CV, Xavier DJ, Macedo C, Manoel-Caetano FS, Rassi DM, Foss-Freitas MC, Foss MC, Sakamoto-Hojo ET, Nguyen C, Puthier D, Passos GA, Donadi EA. Integrative analysis of the transcriptome profiles observed in type 1, type 2 and gestational diabetes mellitus reveals the role of inflammation. BMC Med Genomics. 2014;7:28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 26] [Article Influence: 2.2] [Reference Citation Analysis (1)] |
| 58. | Mercado-Evans V, Mejia ME, Zulk JJ, Ottinger S, Hameed ZA, Serchejian C, Marunde MG, Robertson CM, Ballard MB, Ruano SH, Korotkova N, Flores AR, Pennington KA, Patras KA. Gestational diabetes augments group B Streptococcus infection by disrupting maternal immunity and the vaginal microbiota. Nat Commun. 2024;15:1035. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 33] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 59. | Razo-Azamar M, Nambo-Venegas R, Quevedo IR, Juárez-Luna G, Salomon C, Guevara-Cruz M, Palacios-González B. Early-Pregnancy Serum Maternal and Placenta-Derived Exosomes miRNAs Vary Based on Pancreatic β-Cell Function in GDM. J Clin Endocrinol Metab. 2024;109:1526-1539. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 60. | Zhang Y, He S, Wang X, Wang X, He MY, Yu XX, Xu CR. ACSS2 mediates an epigenetic pathway to regulate β-cell adaptation during gestation in mice. Nat Commun. 2025;16:4697. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 61. | Agarwal P, Brar N, Morriseau TS, Kereliuk SM, Fonseca MA, Cole LK, Jha A, Xiang B, Hunt KL, Seshadri N, Hatch GM, Doucette CA, Dolinsky VW. Gestational Diabetes Adversely Affects Pancreatic Islet Architecture and Function in the Male Rat Offspring. Endocrinology. 2019;160:1907-1925. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 27] [Article Influence: 3.9] [Reference Citation Analysis (1)] |
| 62. | Lowe WL Jr, Karban J. Genetics, genomics and metabolomics: new insights into maternal metabolism during pregnancy. Diabet Med. 2014;31:254-262. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 44] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 63. | Yuan Y, Li Y, Hu L, Wen J. Exosomal RNA Expression Profiles and Their Prediction Performance in Patients With Gestational Diabetes Mellitus and Macrosomia. Front Endocrinol (Lausanne). 2022;13:864971. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 19] [Article Influence: 4.8] [Reference Citation Analysis (1)] |
| 64. | Hawkins NT, Maldaver M, Yannakopoulos A, Guare LA, Krishnan A. Systematic tissue annotations of genomics samples by modeling unstructured metadata. Nat Commun. 2022;13:6736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 65. | Carrasco-Wong I, Moller A, Giachini FR, Lima VV, Toledo F, Stojanova J, Sobrevia L, San Martín S. Placental structure in gestational diabetes mellitus. Biochim Biophys Acta Mol Basis Dis. 2020;1866:165535. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 101] [Article Influence: 16.8] [Reference Citation Analysis (3)] |
| 66. | Li J, Wang A, Guo H, Zheng W, Chen R, Miao C, Zheng D, Peng J, Wang J, Chen Z. Exosomes: innovative biomarkers leading the charge in non-invasive cancer diagnostics. Theranostics. 2025;15:5277-5311. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 31] [Article Influence: 31.0] [Reference Citation Analysis (2)] |
| 67. | Strutz J, Cvitic S, Hackl H, Kashofer K, Appel HM, Thüringer A, Desoye G, Koolwijk P, Hiden U. Gestational diabetes alters microRNA signatures in human feto-placental endothelial cells depending on fetal sex. Clin Sci (Lond). 2018;132:2437-2449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 68. | Rung J, Brazma A. Reuse of public genome-wide gene expression data. Nat Rev Genet. 2013;14:89-99. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 321] [Cited by in RCA: 258] [Article Influence: 19.8] [Reference Citation Analysis (1)] |
| 69. | Arutyunyan A, Roberts K, Troulé K, Wong FCK, Sheridan MA, Kats I, Garcia-Alonso L, Velten B, Hoo R, Ruiz-Morales ER, Sancho-Serra C, Shilts J, Handfield LF, Marconato L, Tuck E, Gardner L, Mazzeo CI, Li Q, Kelava I, Wright GJ, Prigmore E, Teichmann SA, Bayraktar OA, Moffett A, Stegle O, Turco MY, Vento-Tormo R. Spatial multiomics map of trophoblast development in early pregnancy. Nature. 2023;616:143-151. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 247] [Cited by in RCA: 201] [Article Influence: 67.0] [Reference Citation Analysis (1)] |
| 70. | Mora-Ortiz M, Rivas-García L. Gestational Diabetes Mellitus: Unveiling Maternal Health Dynamics from Pregnancy Through Postpartum Perspectives. Open Res Eur. 2024;4:164. [PubMed] [DOI] [Full Text] |