Copyright: ©Author(s) 2026.
World J Diabetes. Apr 15, 2026; 17(4): 115437
Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115437
Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115437
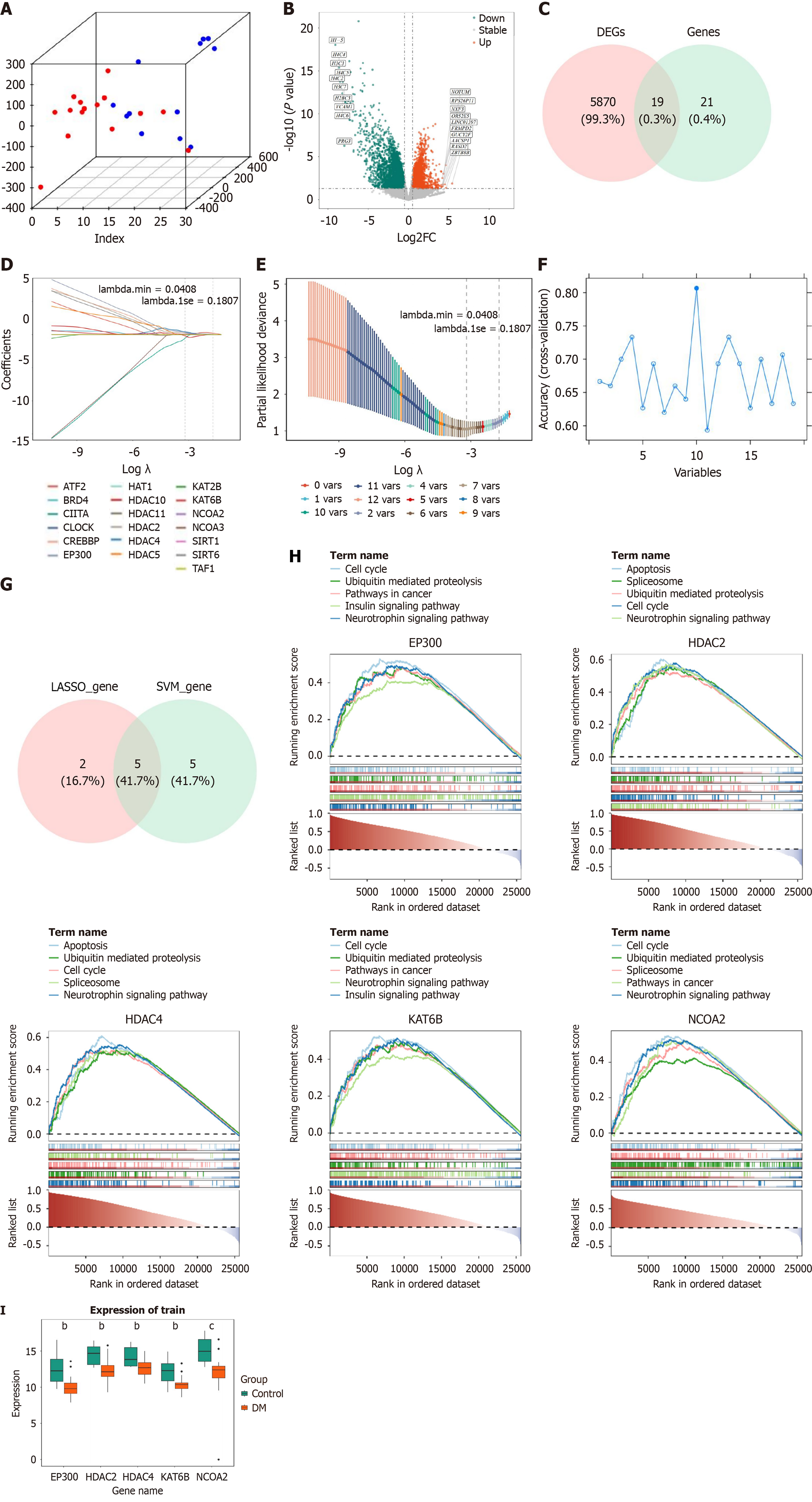
Figure 1 Transcriptome screening of candidate genes.
A: Three-dimensional principal component analysis ; B: Variance analysis volcano map; C: Venny plots of candidate genes; D: Penalty plot of least absolute shrinkage and selection operator logistic regression coefficients; E: Least absolute shrinkage and selection operator cross validation diagram; F: Potential biomarkers screened by machine learning-support vector machine-recursive feature elimination algorithm; G: Machine learning intersection genes; H: GSEA enrichment results for key genes; I: Box plot of key gene expression. bP < 0.01, cP < 0.001. DEGs: Differentially expressed genes; DN: Diabetic nephropathy.
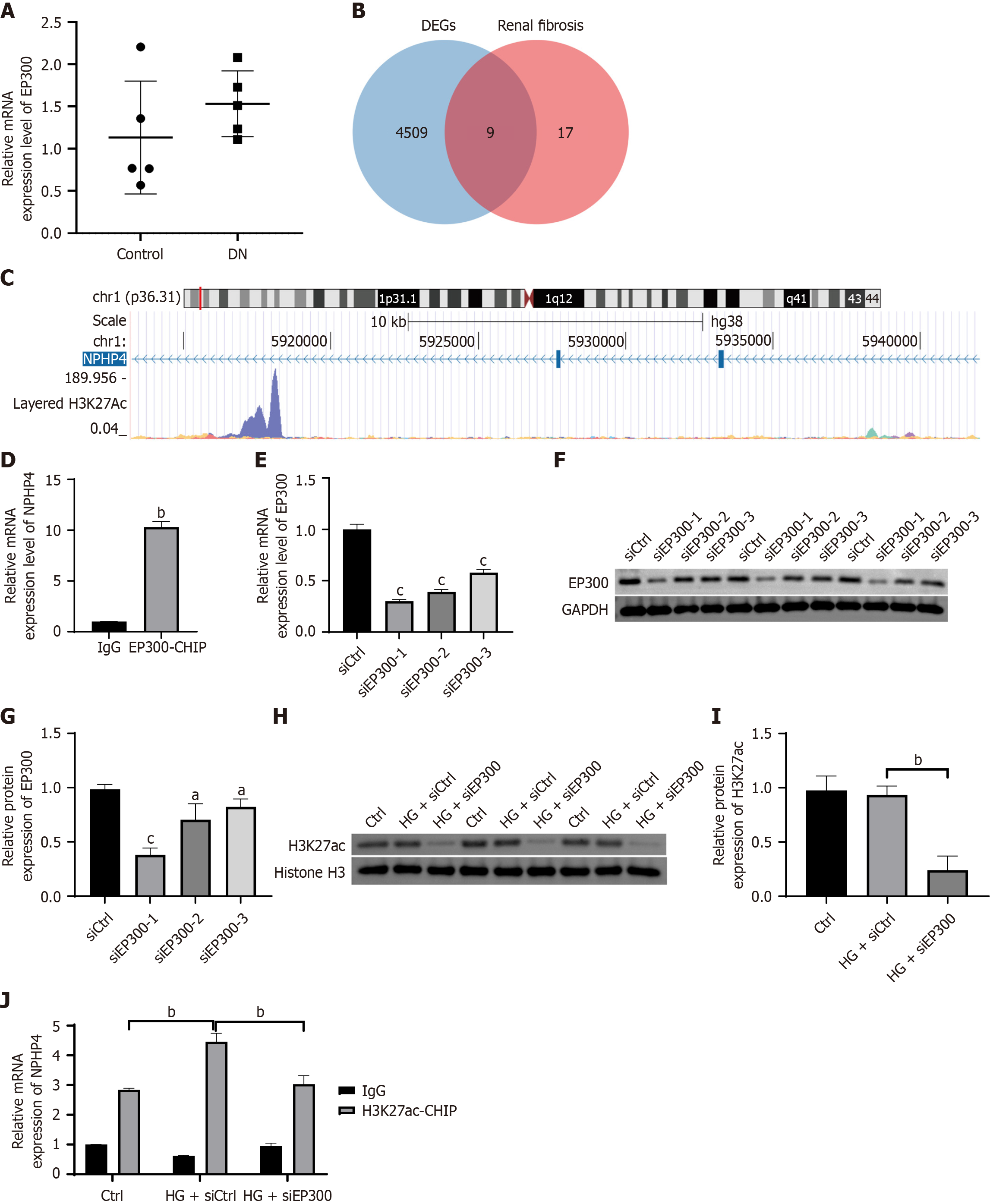
Figure 2 EP300 regulates histone acetylation levels in nephrocystin-4 chromatin fragments.
A: EP300 mRNA expression was measured using real-time quantitative polymerase chain reaction (RT-qPCR); B: Venny diagram of the intersection of EP300-regulated histone acetylation modification action targets with renal fibrosis-related targets; C: Chromatin immunoprecipitation (ChIP)-seq analysis to identify the key target nephrocystin-4 (NPHP4); D: NPHP4 gene abundance in ChIP products was measured using RT-qPCR; E: Measurement of EP300 interference efficiency using RT-qPCR; F and G: Interference efficiency of EP300 was measured using western blot; H and I: Representative western blot images and quantitative analysis of histone H3 lysine 27 protein levels; J: Enrichment of histone H3 lysine 27 at the NPHP4 locus was measured by ChIP-RT-qPCR in HK-2 cells transfected with siCtrl or small interfering RNA targeting EP300 under high glucose conditions. Data are presented as mean ± SD (n = 3). aP < 0.05, bP < 0.01, and cP < 0.001. ChIP: Chromatin immunoprecipitation; DN: Diabetic nephropathy; HG: High glucose; H3K27ac: Acetylated histone H3 lysine 27; IgG: Immunoglobulin G; siEP300: Small interfering RNA targeting EP300.
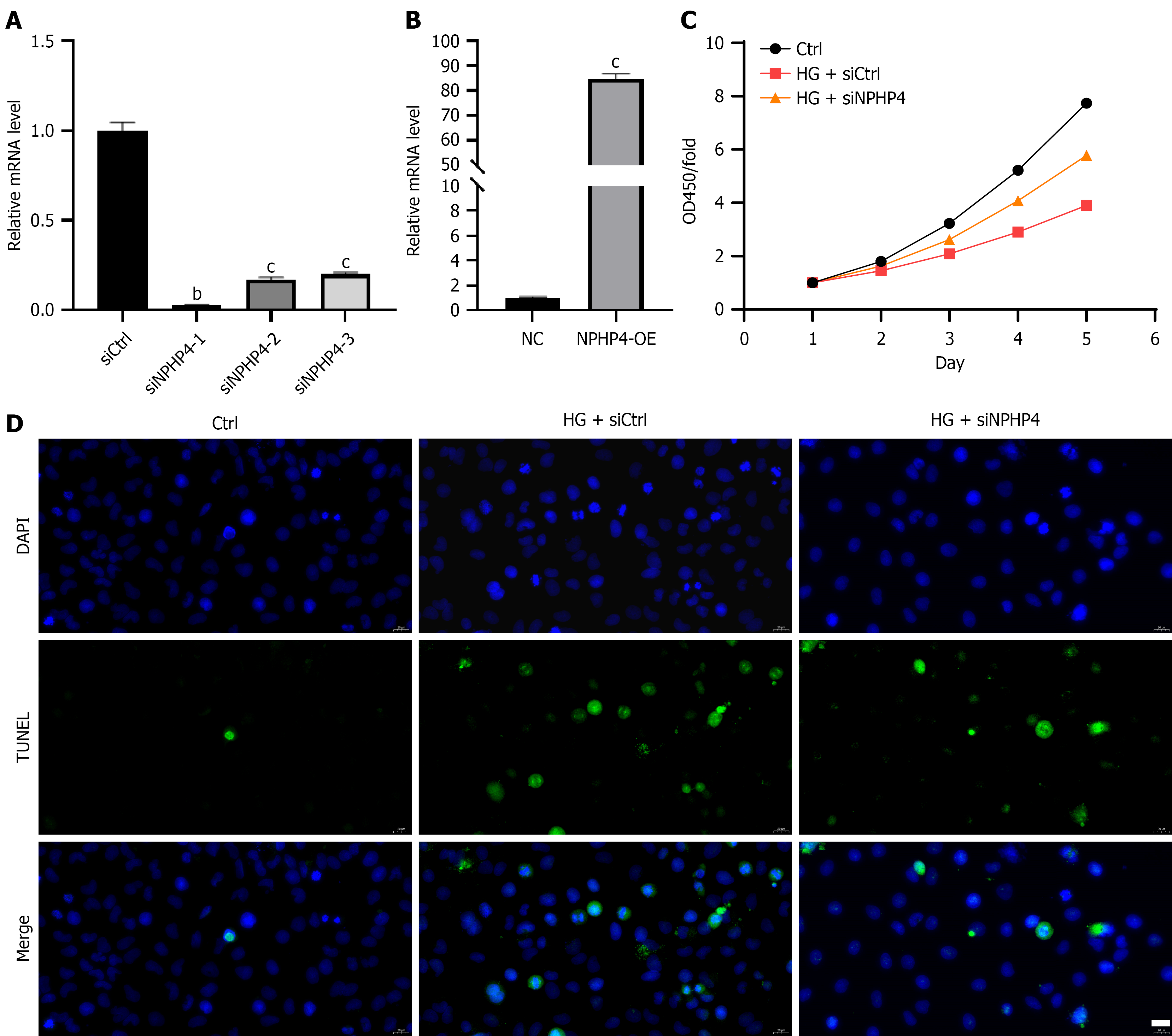
Figure 3 Effect of nephrocystin-4 on HK-2 cell proliferation and apoptosis.
A and B: Measurement of nephrocystin-4 knockdown and overexpression efficiency in HK-2 cells by real-time quantitative polymerase chain reaction; C: Measurement of HK-2 cell viability by cell counting kit-8 assay; D: Measurement of HK-2 cell apoptosis by TUNEL staining. Scale bar = 20 µm. Data are presented as mean ± SD (n = 3). bP < 0.01, cP < 0.001. HG: High glucose; NPHP4-OE: Overexpression of nephrocystin-4; siNPHP4: Small interfering RNA targeting nephrocystin-4.
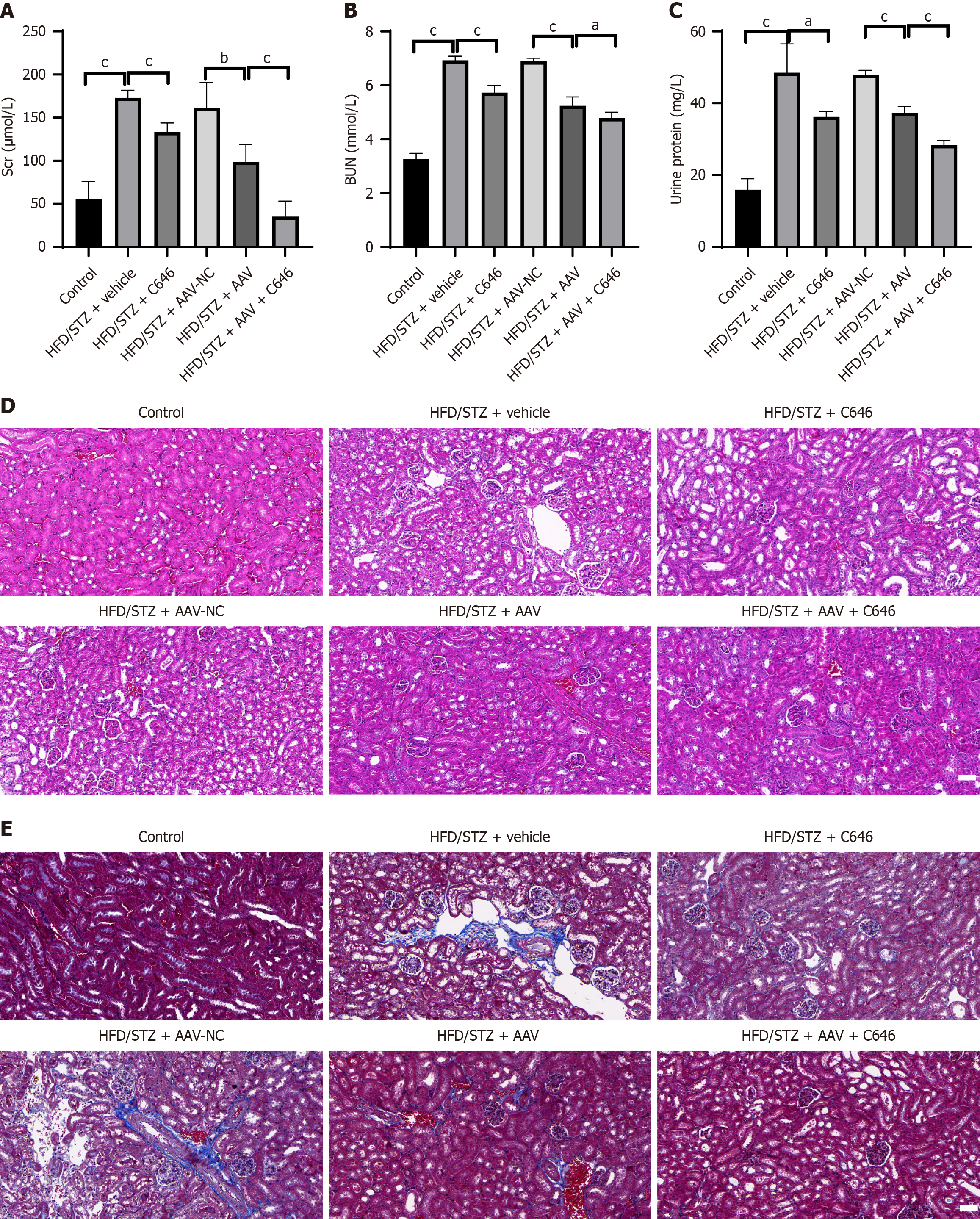
Figure 4 Inhibition of nephrocystin-4 improves renal function in diabetic nephropathy mice.
A: Serum creatinine content; B: Blood urea nitrogen content; C: Urinary protein content; D: Hematoxylin and eosin staining of renal tissue; E: Masson staining of renal tissue. Scale bar = 50 µm. Data are presented as mean ± SD (n = 3). aP < 0.05, bP < 0.01, and cP < 0.001. BUN: Blood urea nitrogen; HFD: High-fat diet; Scr: Serum creatinine; STZ: Streptozotocin.
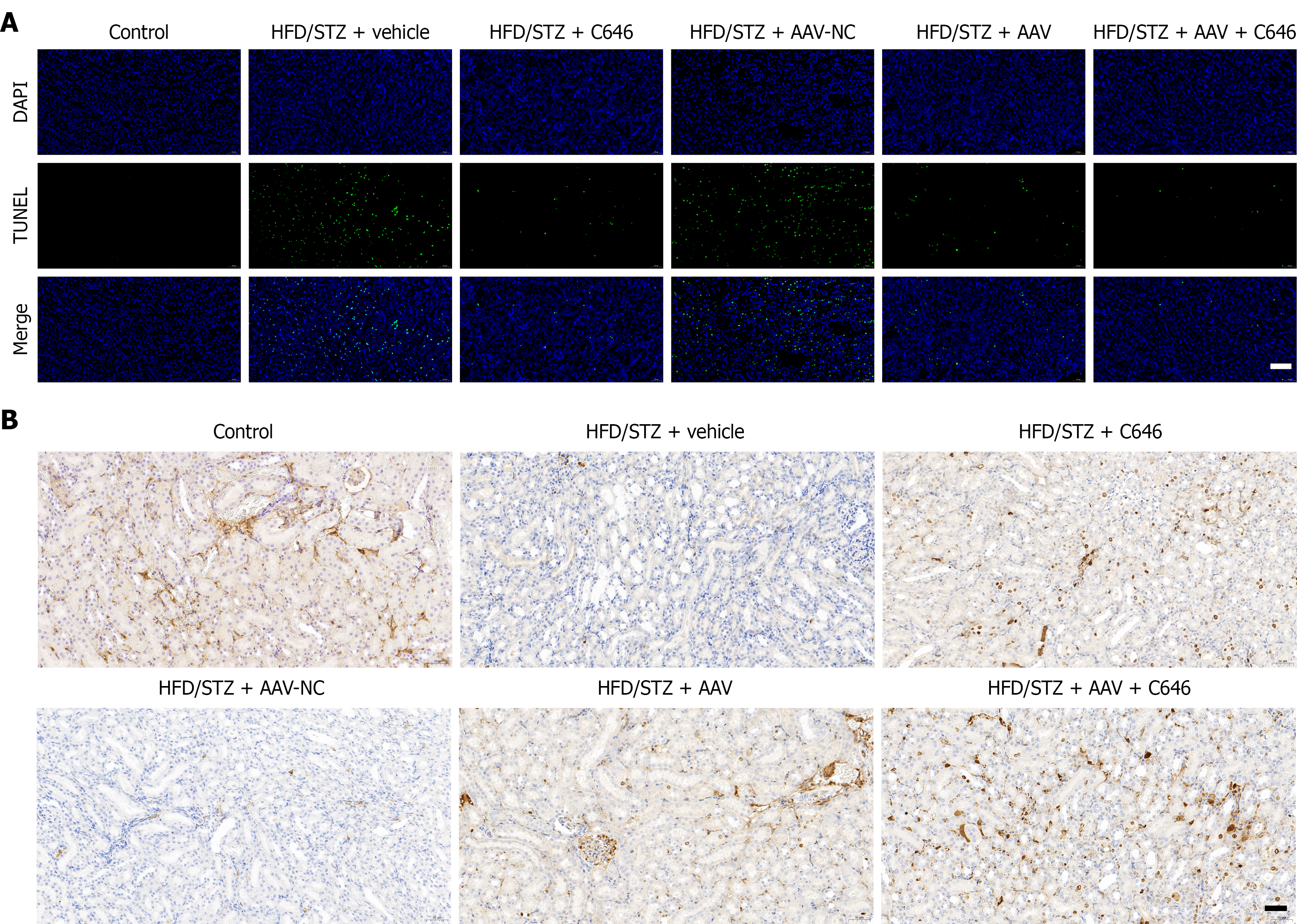
Figure 5 Inhibition of nephrocystin-4 affects apoptosis and proliferation in renal tissue of diabetic nephropathy mice.
A: TUNEL staining of renal tissue; B: Measurement of Ki67 protein levels by immunohistochemistry. Scale bar = 50 µm. HFD: High-fat diet; STZ: Streptozotocin.
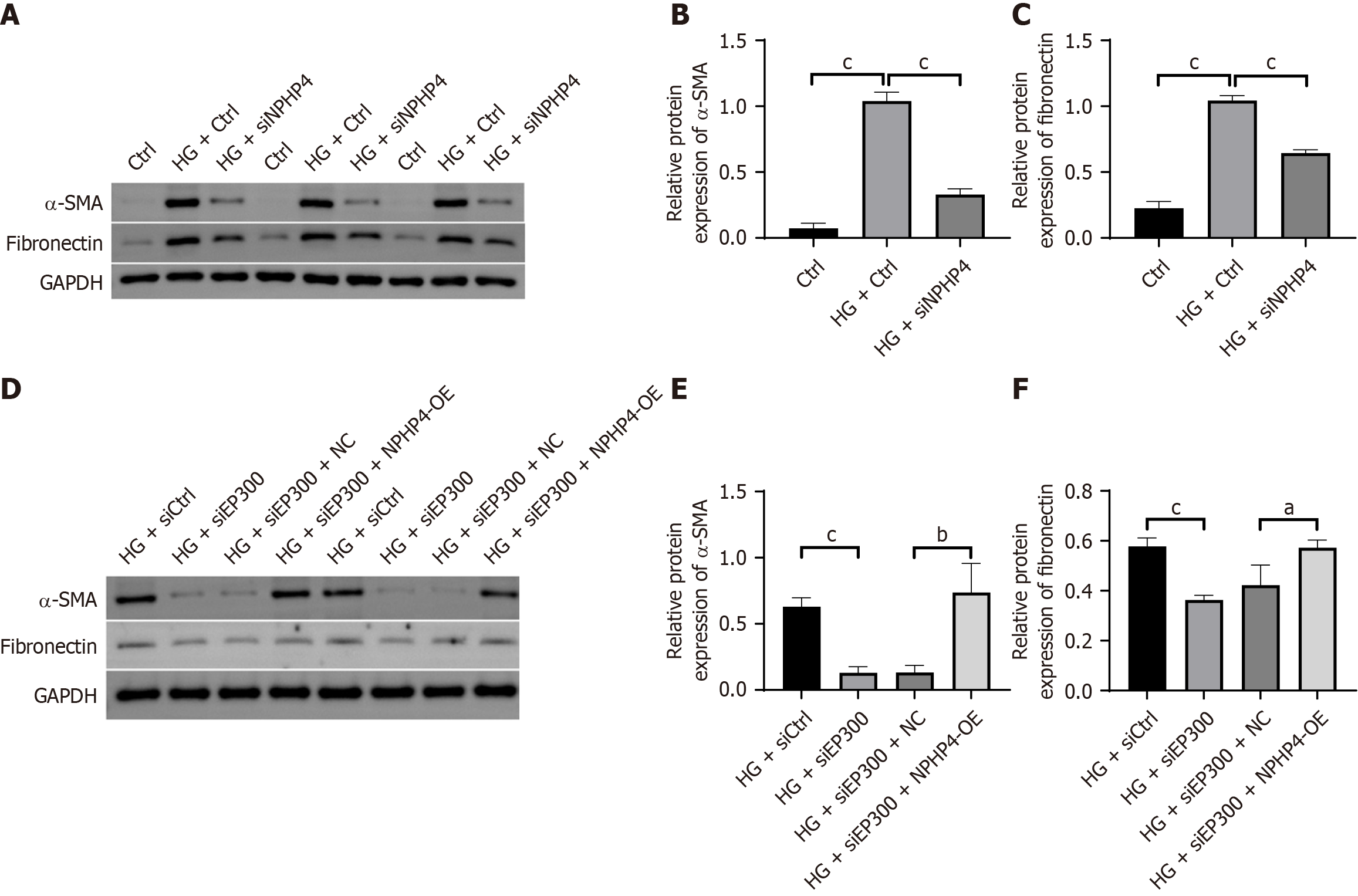
Figure 6 Inhibition of the EP300-nephrocystin-4 axis affects renal fibrosis markers in HK-2 cells.
A-C: Representative western blot images and quantitative analysis of alpha-smooth muscle actin and fibronectin protein levels in nephrocystin-4-knockdown cells; D-F: Representative western blot images and quantitative analysis of alpha-smooth muscle actin and fibronectin protein levels in nephrocystin-4-overexpressing cells. Data are presented as mean ± SD (n = 3). aP < 0.05, bP < 0.01, and cP < 0.001. α-SMA: Alpha-smooth muscle actin; HG: High glucose; NPHP4-OE: Overexpression of nephrocystin-4; siNPHP4: Small interfering RNA targeting nephrocystin-4.
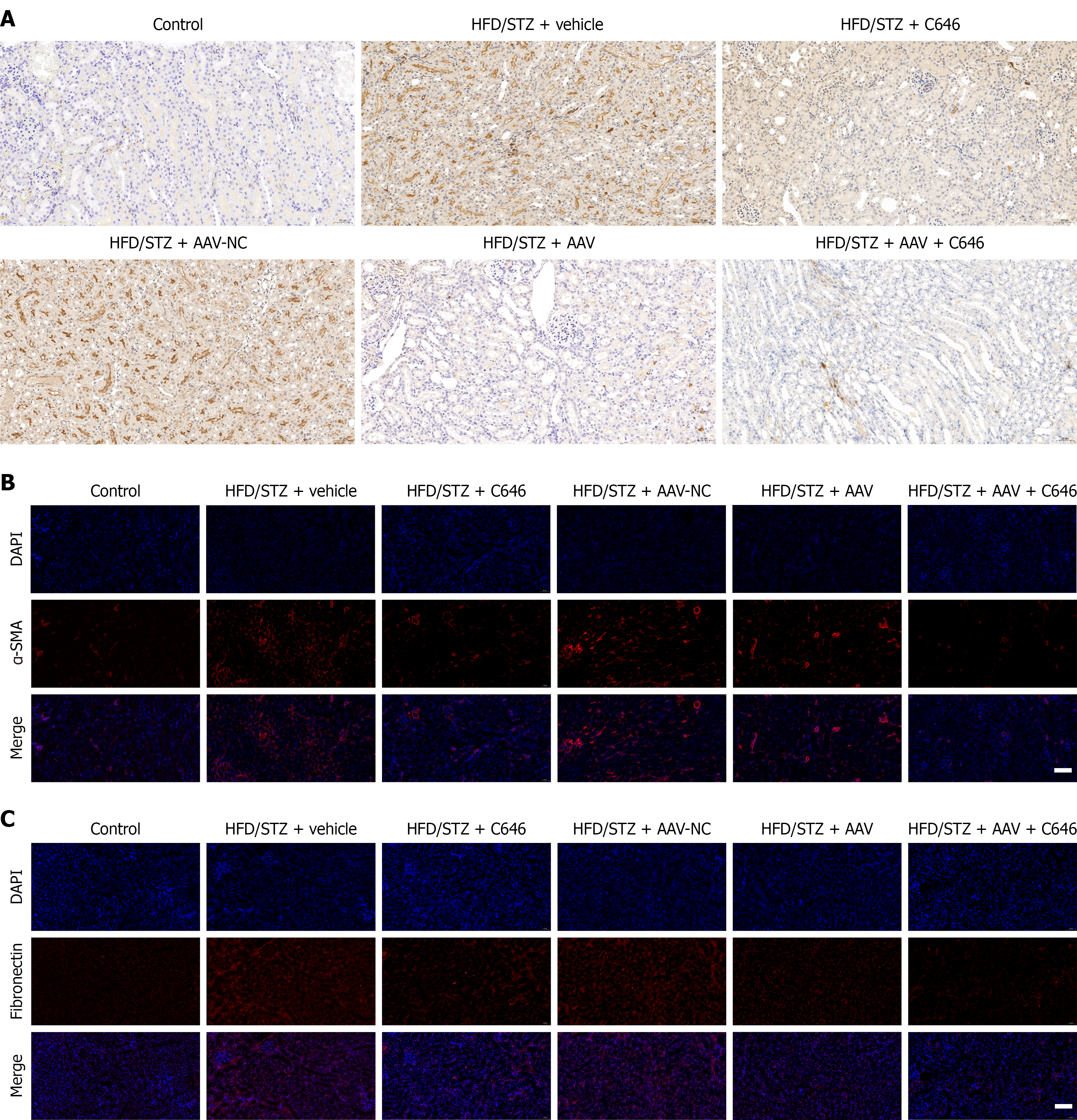
Figure 7 Inhibition of the EP300-nephrocystin-4 axis affects renal fibrosis markers in diabetic nephropathy mice.
A: Measurement of nephrocystin-4 protein levels by immunohistochemistry. Scale bar = 100 µm; B: Measurement of alpha-smooth muscle actin protein levels by immunofluorescence. Scale bar = 50 µm; C: Measurement of fibronectin protein levels by immunofluorescence. Scale bar = 50 µm. α-SMA: Alpha-smooth muscle actin; HFD: High-fat diet; STZ: Streptozotocin.
- Citation: Si W, Dai Y, Hu GP, Zhang Q, Lv F, Zhang Q. EP300 drives renal fibrosis in diabetic nephropathy via histone acetyltransferase-mediated nephrocystin-4 expression. World J Diabetes 2026; 17(4): 115437
- URL: https://www.wjgnet.com/1948-9358/full/v17/i4/115437.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i4.115437