Copyright: ©Author(s) 2026.
World J Diabetes. Apr 15, 2026; 17(4): 115275
Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115275
Published online Apr 15, 2026. doi: 10.4239/wjd.v17.i4.115275
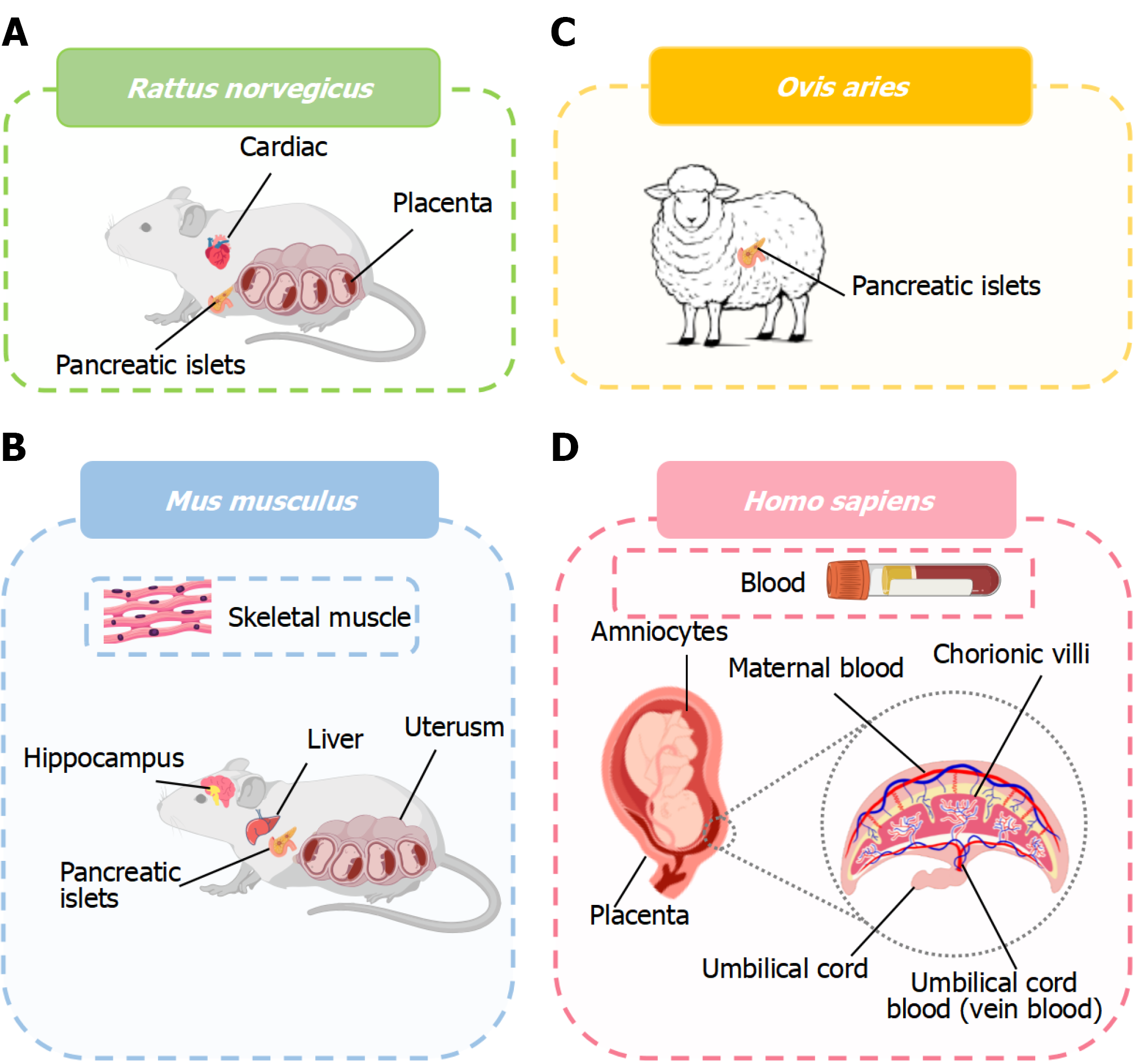
Figure 1 Species and tissues coverage in gestational diabetes mellitus transcriptomic studies.
A: Rattus norvegicus. Datasets most frequently sample pancreatic islets, cardiac tissue and placenta; B: Mus musculus. Commonly profiled tissues include hippocampus, pancreatic islets, liver, skeletal muscle and uterus; C: Ovis aries. Sheep studies primarily focus on pancreatic islets; D: Homo sapiens. Human datasets encompass placenta, maternal peripheral blood, chorionic villi, amniocytes and fetal umbilical cord blood (e.g., venous blood).
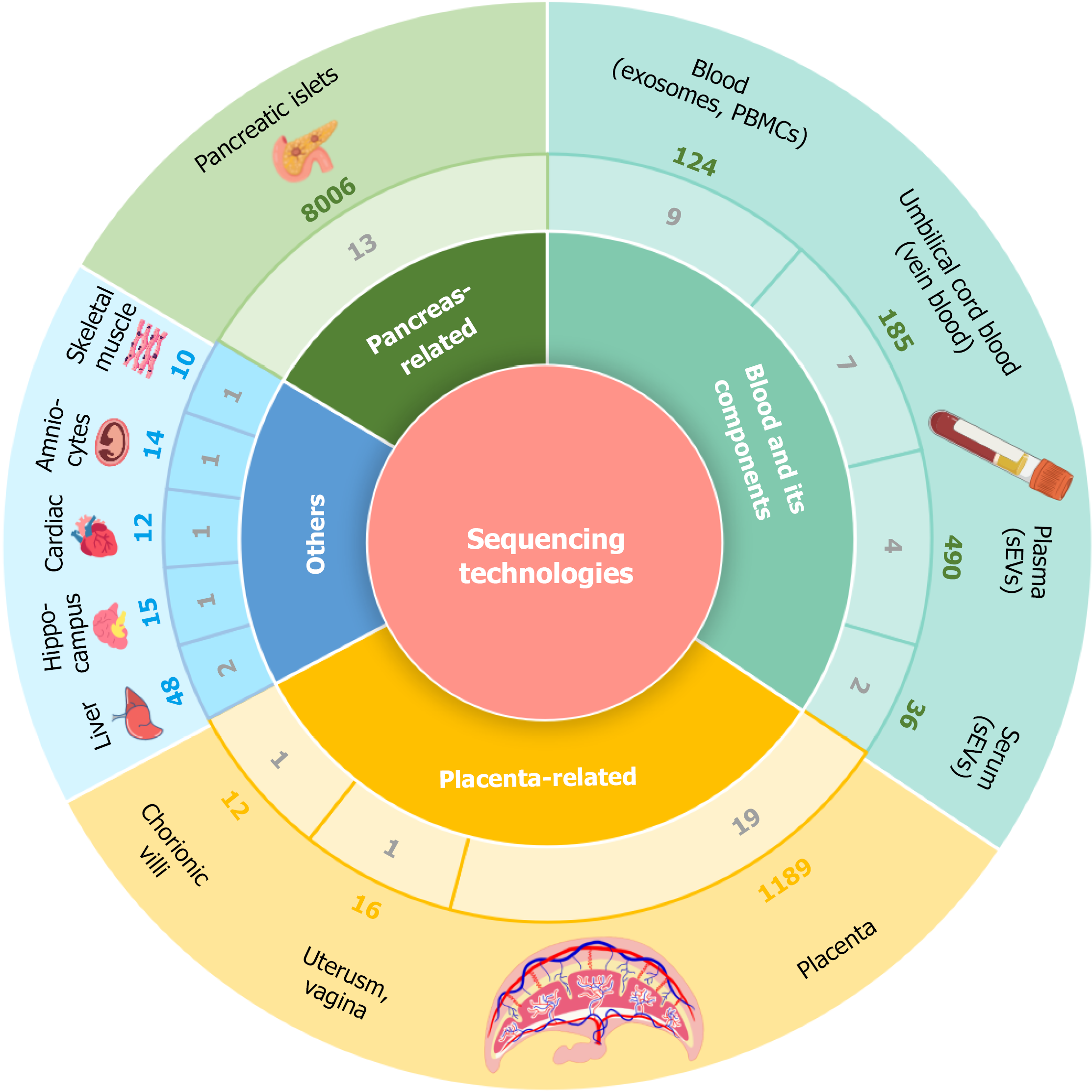
Figure 2 The dataset counts and sample sizes across four major categories in gestational diabetes mellitus transcriptomic studies.
From the innermost to the outermost ring, the concentric layers represent: (1) Broad tissue categories (placenta-related, blood and its components, pancreas/islet-related and others); (2) The number of datasets assigned to each subcategory; and (3) The cumulative sample size for each category. For example, within the placenta-related category, 19 datasets focusing on placenta were identified, together comprising 1189 samples. PBMCs: Peripheral blood mononuclear cells; sEVs: Secreted extracellular vesicles.
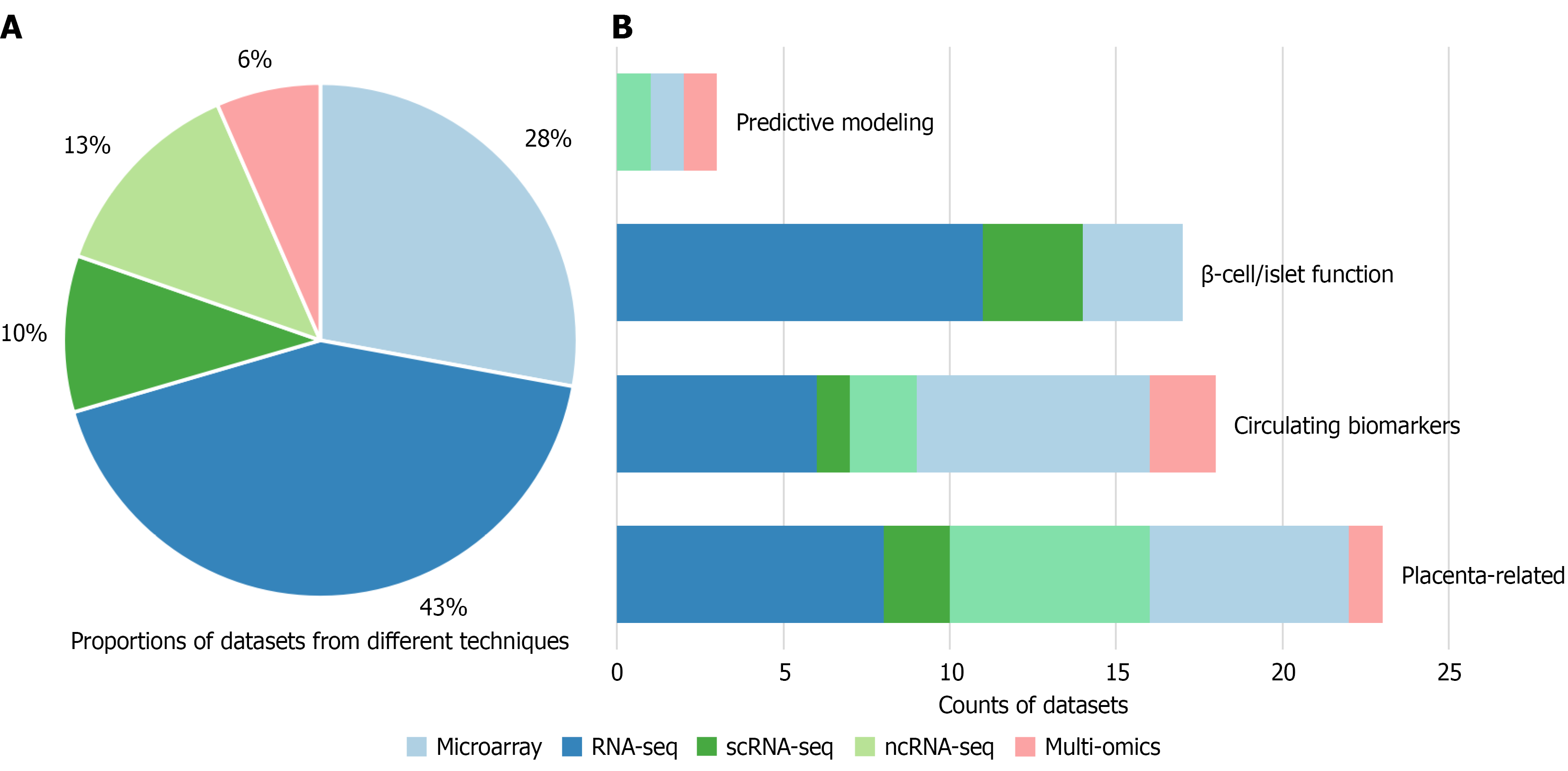
Figure 3 Various transcriptomic technologies in gestational diabetes mellitus research.
A: Overall proportions of transcriptomic technologies represented in gestational diabetes mellitus (GDM) datasets. RNA-sequencing (RNA-seq) remains the most prevalent (43%), followed by DNA microarrays (28%), non-coding RNA-seq (13%), multi-omics (6%) and single-cell RNA-seq (scRNA-seq) (10%). These distributions highlight both the persistence of classical platforms and the growing role of emerging modalities in GDM research; B: Distribution of dataset types across research themes in GDM. Placenta-related datasets dominate, followed by circulating biomarkers and β-cell/islet, while predictive modeling studies remain fewer. RNA-seq (blue) and microarray (light blue) are the most widely used platforms overall, whereas non-coding RNA-seq (light green) and scRNA-seq (dark green) contribute substantially to biomarker and islet-related studies. Multi-omics approaches (pink), though relatively limited, are increasingly adopted in biomarker discovery and predictive modeling. RNA-seq: RNA-sequencing; scRNA-seq: Single-cell RNA-sequencing; ncRNA-seq: Non-coding RNA-sequencing.
- Citation: Xie LL, Li SW, Zou Y, Qin D, Sun J, Xiao YX, Li T, Hao YJ, Li B. Mining transcriptomic data for gestational diabetes mellitus: What public datasets reveal. World J Diabetes 2026; 17(4): 115275
- URL: https://www.wjgnet.com/1948-9358/full/v17/i4/115275.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i4.115275